Selenoprotein evolution: SECIS: Difference between revisions
Tomemerald (talk | contribs) |
Tomemerald (talk | contribs) |
||
| Line 590: | Line 590: | ||
== Reference sets of vertebrate SECIS elements == | == Reference sets of vertebrate SECIS elements == | ||
=== TXNRD1: 17 SECIS insertion sequences === | |||
<pre> | |||
>TXNRD1_homSap Homo sapiens (human) COVE score: 25.36 | |||
auuuggcagg gcaucga agggaugcaucc augaa gucaccagucuc aa gcccaugugg uaggcggugau ggaa caacug ucaaauc aguuuuagca | |||
>TXNRD1_panTro Pan troglodytes (chimp) COVE score: 25.36 | |||
auuuggcagg gcaucga agggaugcaucc augaa gucaccagucuc aa gcccaugugg uaggcggugau ggaa caacug ucaaauc aguuuuagca | |||
>TXNRD1_otoGar Otolemur garnettii (bushbaby) COVE score: 23.42 | |||
aucggcagug caucgac gggaugcgucc augaa gucaccagccuc aa gccugugugg ugggcagugau ggaa caacu guccaau caguuucuau | |||
>TXNRD1_tupBel Tupaia belangeri (tree_shrew) COVE score: 25.13 | |||
ucuuggcagc gcaucag agggaugcgucc augaa gucaccagccuc aa gcccgugcgg ugggcggugau ggaa cgacug ccagauc aguuucagca | |||
>TXNRD1_musMus Mus musculus (mouse) COVE score: 27.9 | |||
aucuggcaga gcaucac aggcaugcgucc augaa gucacuggccuc aa gcccaagug gugggcagugac agaa gagcu gccgggu cuguugagcu | |||
>TXNRD1_ratNor Rattus norvegicus (rat) COVE score: 28.03 | |||
uucggcagag caucacg gugcgucc augaa gucacuagccuc aa gcccaagugg ugggcagugac agaa agc ugucgau cuguuggguu | |||
>TXNRD1_cavPor Cavia porcellus (guinea_pig) COVE score: 23.88 | |||
gcggcggcac cguaggg ugcgucc augag gucaccagccuc aa gcccgaggg ugggcggugac ggau cgc gccgcg uggcucagcu | |||
>TXNRD1_eriEur Erinaceus europaeus (hedgehog) COVE score: 26.25 | |||
ugucagcaga gcaucaa agggaugcgucc augaa gucaccagccuc aa gcccgugcggg ugggcagugac ggaa cacug ucgaagc aguuucaaca | |||
>TXNRD1_canFam Canis familiaris (dog) COVE score: 24.54 | |||
auucggcaug caucggc guggugcgucc augaa gucacuggccuc aa gccaugcg gugggcagugau ggag caac ugucgag caguuuuagu | |||
>TXNRD1_felCat Felis catus (cat) COVE score: 24.14 | |||
acucggcagc gcaucgg agggcgcgucc augaa gucaccggcccc aa gcccccgcg gugggcggugau ggaa caagu gccgagc aguuuuagcg | |||
>TXNRD1_equCab Equus caballus (horse) COVE score: 21.76 | |||
acucggcagu gcaucga agggaugcgucc augaa gucacuggccuc aa agcccaugug gugggcggugau ggaa cagcug ucgaagc aguuuuagca | |||
>TXNRD1_loxAfr Loxodonta africana (elephant) COVE score: 22.55 | |||
auuggcagcg caucgag ggaugcaucc augaa gucacuggccuc aa gcccaugug gggggcggugau ggaa cagcu gucgaau cagcuuuggc | |||
>TXNRD1_echTel Echinops telfairi (tenrec) COVE score: 19.5 | |||
guggcagugc aucaaga gaugcguuc augaa aucgcuugcccc aa gcccga guggcgggcag cgau ggaaca ucugucu caucaguuuc | |||
>TXNRD1_monDom Monodelphis domestica (opossum) COVE score: 23.46 | |||
ggcucgcggu gcaucgg ugagaugcguuc augaa gucgcugccug aa gcccauaucccgugg uggguggugac cgaa agaaccg ccggcc uccguuuuau | |||
>TXNRD1_ornAna Ornithorhynchus anatinus (platypus) COVE score: 18.89 | |||
aggagugcac ccaaggg cugcauuu augaa gucagagccaa aa gccagcauuuugcgg uuggcugugau ggaa aaa cuccug ccacaguuuu | |||
>TXNRD1_anoCar Anolis carolinensis (lizard) COVE score: 32.53 | |||
uggcaaggca uuguuca agaugcuucc augaa gucacagucua aa accagugcuuucugg uaggcagugau ggaa aga uugcugg cacaacuuga | |||
>TXNRD1_galGal Gallus gallus (chicken) COVE score: 26.64 | |||
uagcagggca uuucaca caugcuuuc augaa aucacagccug aa gccugcacugucugg ugggcagugau ggaa gaacu gcugaca cagcugaaca | |||
</pre> | |||
=== TXNRD3: 9 SECIS insertion sequences (problematic) === | |||
<pre> | |||
>TXNRD3_homSap Homo sapiens (human) COVE score: 19.10 | |||
gacagcgaga agcagug ggacugcuucc uugac gccuuagcuu gg agccccguuaugaggu gagccaaggc ugac ucu cgcaagc caggacugag | |||
>TXNRD3_panTro Pan troglodytes (chimp) COVE score: 19.10 | |||
agcagugggc GACAGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUU GG AGCCCCGUUAUGAGGU GAGCCAAGGC UGAC UCU CGCAAGC CAGGACUGAG | |||
>TXNRD3_macMul Macaca mulatta (rhesus) COVE score: 17.86 | |||
GACuGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUU GG AGCCCuGUUgUGAGGU GAGCCAAGGC cGAC UCU CGCAAGC CAGGACUcAG | |||
>TXNRD3_calJac Callithrix jacchus (marmoset) COVE score: 25.26 | |||
GACuGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUc Ga AGCCCuGUUAcGAGGU GAGCCAAGGC UGAu UCU CGCAAGC gAGGACUGAG | |||
>TXNRD3_musMus Mus musculus (mouse) COVE score: 21.86 | |||
CUGACGGCAU GCAGCAG CCAGGCUGCUUCC UUGAC ACCUUGGCUC GG AACCUGCAGAGGU GAGCCAAGGC CGAC UUC UGCACGU CAGCCUCGAC | |||
>TXNRD3_ratNor Rattus norvegicus (rat) COVE score: 25.37 | |||
cugauggcgu gcagcag ccaggcugcaucc uugac gccuuggcuc gg aaccugcagagg ugagccaaggc cgac ucc ugcacgu cagccucgac | |||
>TXNRD3_canFam Canis familiaris (dog) COVE score: 26.93 | |||
GCUGGCUGGA GAGGCAG GCAGGCUGCCUCC UUGAC GCCUUAGCUC GG AACCGCUGUGAGG UGAGCUAAGGC CGAU GUC CUCCAU GCCAGGCCAG | |||
>TXNRD3_felCat Felis catus (cat) COVE score: 20.14 | |||
CUGGCUgGGA GAGGCAG GucGGCUGCCUCC UUGAC GuCUUAGCUC GG AgcccgaUGUGAGG UGAGCUAAGGC CGAU Ggu cuuccac gucagaaucg | |||
>TXNRD3_bosTau Bos taurus (cow) COVE score: 12.55 | |||
GGCUGACGGG AGGGCAG ACUCGCUGCCUCC UUGAC GUCUUCGCUC AG AGCCGCCAGGU GAGCCAAGAC CGAC CUC UGCCCA CCAGCUCCUC | |||
</pre> | |||
=== TXNRD2: 10 SECIS insertion sequences === | |||
<pre> | |||
>TXNRD2 homSap Homo sapiens (human) COVE score: 31.00 | |||
cacccccccc caggcuc cuggugccagaug augac gaccugggugg aa accuacccugugg gcacccauguc cgag ccccc uggcauu ucugcaaugc | |||
>TXNRD2 panTro Pan troglodytes (chimp) COVE score: 31.00 | |||
ggcacccccc caggcuc cuggugccagaug augac gaccugggugg aa accuacccugugg gcacccauguc cgag ccccc uggcauu ucugcauugc | |||
>TXNRD2 tupBel Tupaia belangeri (tree shrew) COVE score: 36.55 | |||
cucccucccc aggcccc cgaugccagaug augac ggccuggacag aa acccacccuguggg cugcccagguc ugaa cccuccc uggugucu uuggggugua | |||
>TXNRD2 musMus Mus musculus (mouse) COVE score: 36.94 | |||
gccagccucu gacacuc ccagcgucagaug augau ggccugggcag aa accccauguggg ccgcccagguu ugaa ccccu ggcauuu cuagagcacu | |||
>TXNRD2 ratNor Rattus norvegicus (rat) COVE score: 34.71 | |||
cagccuucac acacugc cagugucagaug augac ggccugugcag aa acccccacguggg cugcccagguu ugaa ccccug gcauuu cuggagugcu | |||
>TXNRD2 cavPor Cavia porcellus (guinea pig) COVE score: 34.41 | |||
ucuggccccc caggucc ccagugccaguug augau ggccugggcag aa acccacccuguggg caguccauguc ugaa cuccc uggcauu ucuggagugc | |||
>TXNRD2 eriEur Erinaceus europaeus (hedgehog) COVE score: 34.68 | |||
ccagccccac caggccc ccgaugccagaua augau gacuugugcag aa acccacccggg cugcccauguc ugag ccucug uggcauu cuggagugua | |||
>TXNRD2 canFam Canis familiaris (dog) COVE score: 34.36 | |||
accagccccg ccaggcc ccgaugccagaag augac gacgugugcag aa accccccuguggg cugcccgcguc cgag cccccuggc guuucugg aauguaaaua | |||
>TXNRD2 equCab Equus caballus (horse) COVE score: 31.37 | |||
ccagcccugc cagguuc ccgaugccagacg augac gaccugcgcgg aa acccacccuguggg cugcccacguc cgag ccccc uggcauu ucugaagugc | |||
>TXNRD2 echTel Echinops telfairi (tenrec) COVE score: 37.41 | |||
cccuugcccc caccuca gcgccagaug augaa gacaugugcag aa acccagcccguggg cugcccauguc ugag ccccc ugacgu uucuggagug | |||
</pre> | |||
=== SELS: 20 SECIS insertion sequences === | === SELS: 20 SECIS insertion sequences === | ||
Revision as of 16:02, 5 June 2008
Introduction to selenocysteine insertion into selenoproteins
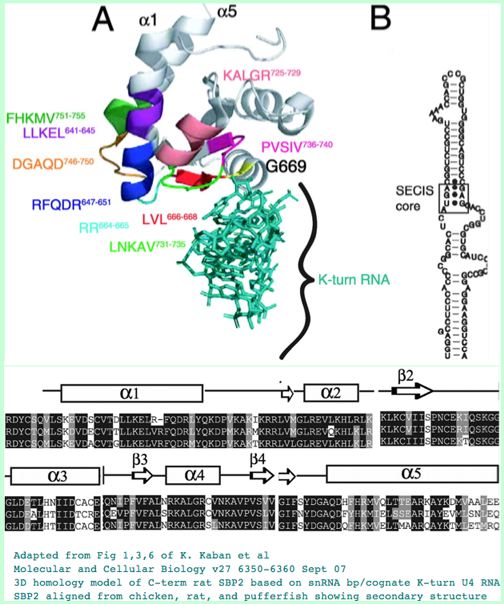
The SECIS element in 3' UTR of selenoproteins allows the insertion of selenocysteine for the appropriate TGA codon at time of translation. Auxillary gene products are needed for this to happen, including a selenocysteine-specific translation elongation factor (eEFSec), Sec-tRNA[Ser]Sec, ribosomal protein L30, and a binding protein (SBP2).
The SBP2 protein has three domains, including a conserved L7Ae RNA binding motif for SECIS RNA binding, a not completely overlapping ribosome binding site and a redox-sensitive cysteine-rich domain. SBP2 shuttles between the nucleus and the cytoplasm via nuclear localization motif SELEQNESSKKNKKKKEKSKSSYEVLPVQE and nuclear export signal motifs, the latter using the CRM1 pathway.
The kink-turn tertiary structure in the SECIS element is at the core of binding recognition. A mutation in that of SEPN or in SB2 lead to rigid spine muscular dystrophy or abnormal thyroid hormone function, the latter through inadequate production of DIO2 needed to activate thyroid hormone T4.
A strong paralog of BP2 was noticed and briely studied earlier. That protein, KIAA0256, also has 17 coding exons with near-identical intron locations and phases and full-length alignability, establishing KIAA0256 and BP2 as resulting from an ancestral gene duplication. The percent identity overall is only 42% but reaches 72% (human to human) in the 3 exons defining the L7Ae kink-turn binding motif.
Since both genes can be recovered from chondrichthyes, lamprey, and amphioxus, the duplication event preceded the emergence of chordates. The roots of the selenoproteome and the insertion mechanism lie much deeper in pre-metazoa so it follows that the parent gene must have assumed the key SECIS recognition roles (as did both descendent genes, at least initially).
We could ask if KIAA0256 still plays a role in SECIS in contemporary organisms. One possibility is that some of the 30-odd selenoproteins preferentially use KIAA0256 over BP2, correlating to the two observed structural classes of SECIS elements (which needs to be revisited in the genomics era). Another possibility is differential use during development or in various cell types. It's worth noting that KIAA0256 is considerably more conserved across its length than BP2, which has only a few exons conserved back to fish) suggesting the latter has a more limited role.
A literature review establishes that DIO, GPX, and SEPP1 were generally taken as proxies for all selenoprotein SECIS elements in terms of establishing a role for BP2. Yet the timing of expansion of these and specialization of the former to thyroid during the period of gene duplication may have tainted their representativeness. Possibly many other selenoproteins utilize KIAA0256 at least to some extent. That would account for the greater conservation of KIAA0256 and the difficulty finding SECIS elements in expected position for certain selenoproteins, ie their elements may be adapted to a different receptor and so have low Cove scores to the extent that tool is based on the BP2 fold paradigm.
Ribosomal protein L30 appears also to bind the SECIS kink-turn and play a role in selenocysteine insertion despite its meagre length of 114 amino acids. Likely many ribosomal proteins, L30 has remarkable sequence conservation, here still 94% at human-lamprey.
L30 has weak homology and secondary structure parallels to the pertinent region of BP2 and KIAA0256. These latter proteins might be viewed as domain composites that have accreted the kink-turn binding domain long ago from this undoubtedly ancient ribosomal protein. This opens a window on the origin of selenocysteine insertion in early eukaryotes.
KIAA0256: the forgotten, misannotated SECIS binding protein
>SECISBP2_homSap Homo sapiens (human) full length 0 MASEGPREPESE 0 0 GIKLSADVKPFVPRFAGLNVAWLESSEACVFPSSAATYYPFVQEPPVTE 2 1 QKIYTEDMAFGASTFPPQYLSSEITLHPYAYSPYTLDSTQNVYSVPGSQYLYNQPSCYRGFQTVKHRNENTCPLPQEMKALFK 0 0 KKTYDEKKTYDQQKFDSERADGTISSEIKSARGSHHLSIYAENSLKS 1 2 DGYHKRTDRKSRIIAKNVSTSKPEFEFTTLDFPELQGAENNMSEIQKQPKWGPVHSVSTDISLLREVVKPAAVLSK 0 0 GEIVVKNNPNESVTANAATNSPSCTR 1 2 ELSWTPMGYVVRQTLSTELSAAPKNVTSMINLKTIASSADPKNVSIPSSEALSSDPSYNKEKHIIHPTQK 0 0 SKASQGSDLEQNEASRKNKKKKEKSTSKYEVLTVQEPPRIE 0 0 DAEEFPNLAVASERRDRIETPKFQSKQQPQ 0 0 DNFKNNVKKSQLPVQLDLGGMLTALEKKQHSQHAKQSSKPVVVS 1 2 VGAVPVLSKECASGERGRRMSQMKTPHNPLDSSAPLMKKGKQREIPKAKKPTSLKK 0 0 IILKERQERKQRLQENAVSPAFTSDDTQDGESGGDDQFPEQAELS 1 2 GPEGMDELISTPSVEDKSEEPPGTELQRDTEASHLAPNHTTFPKIHSRRFRD 2 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTVAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALKEKEEPHY 1 2 IEIWKKHLEAYSGCTLELEESLEASTSQMMNLNL* 0 407–525 domain required for U insertion but not SECIS binding (399–516 in rat) 540 R540Q allele of SBP2 decreases GPX1 and DIO2 650–752 L7Ae motif kink-turn binding motif 676 invariant glycine (669 in rat) >KIAA0256_homSap Homo sapiens (human) length=1101 0 MDRAPTEQ 0 0 NVKLSAEVEPFIPQKKSPDTFMIPMALPNDNGSVSGVEPTPIPSYLITCYPFVQENQSNR 2 1 QFPLYNNDIRWQQPNPNPTGPYFAYPIISAQPPVSTEYTYYQLMPAPCAQVMGFYHPFPTPYSNTFQAANTVNAITTECTERPSQLGQVFPLSSHRSRNSNRGSVVPK 0 0 QQLLQQHIKSKRPLVKNVATQKETNAAGPDSRSKIVLLVDASQQT 1 2 DFPSDIANKSLSETTATMLWKSKGRRRRASHPTAESSSEQGASEADIDSDSGYCSPKHSNNQPAAGALRNPDSGTMN 0 0 HVESSMCA 1 2 GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ 0 0 VGFRCRGHSTSSERRQNLQKRPDNKHLSSSQSHRSDPNSESLYFE 0 0 DEDGFQELNENGNAKDENIQQKLSSKV 0 0 LDDLPENSPINIVQTPIPITTSVPKRAKSQKKKALAAALATAQEYSEISMEQKKLQ 0 0 EALSKAAGKKNKTPVQLDLGDMLAALEKQQQAMKARQITNTRPLSYT 1 2 VVTAASFHTKDSTNRKPLTKSQPCLTSFNSVDIASSKAKKGKEKEIAKLKRPTALKK 0 0 VILKEREEKKGRLTVDHNLLGSEEPTEMHLDFIDDLPQEIVSQE 1 2 DTGLSMPSDTSLSPASQNSPYCMTPVSQGSPASSGIGSPMASSTITKIHSKRFRE 2 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 2 ETNWRNMVETSDGLEASENEKEVSCKHSTSEKPSKLPFDTPPIGKQPSLVATGSTTSATSAGKSTASDKEEVKPDDLEWASQQSTETGSLDGSCRDLLNSSITSTTSTLVP GMLEEEEDEDEEEEEDYTHEPISVEVQLNSRIESWVSETQRTMETLQLGKTLNGSEEDNVEQSGEEEAEAPEVLEPGMDSEAWTADQQASPGQQKSSNCSSLNKEHSDSNYTTQTT* 0 phosphoserines predicted at SwissProt; no counterparts in SECISBP2 exon 8 skipped in RefSeq KIAA0256 exon 11 lacks counterpart in SECISBP2
Note the Irish proband with compound mutation K438stop/IVS8ds+29G/A (paternal allele inactive, maternal allele a splice donor mutation leading to early truncation) has been incorrectly described as a SECISBP2 knockout; in fact 48% production of wildtype maternal allele still occurs. Additionally, KIAA0256 may be able to partially compensate for reduction in level or loss of SECISBP2. Knockout mice for tRNA(Sec), unable to make any selenoproteins, die in utero.
In two earlier papers, the same authors in twice-published Figure 1B aligned full length SECISBP2 to an intermediate region of the KIAA0256 gene that happened to begin with a methionine, namely residues 422-849 of the 1056 residue protein (which includes the motif-bearing residues 632-829 of exons 14-16). Worse, the model KIAA0256 transcript was an exon-skipping minor splice variant and contains an additional exon 11 without counterpart in SECISBP2:
KIAA0256 originally arose in a GenBank submission package from a large-scale mRNA project at the Kaluza Institute. It skips over highly conserved exon 8 (which does not alter downstream reading frame since it resides in a series of consecutive phase 00 spliced exons). Modelling on this oddity, NCBI later confused the record by mixing in its own predicted genes (from genome assemblies without significant transcript programs) and labelling them mRNAs.
While baboon also has experimental transcripts skipping this exon (FC178616, FC178616), mammalian transcripts almost always retain the exon, for example human (AK307480), macaque (CJ457866), mouse (AK145135), rat (CK602552), dog (CO708934), horse (CX604216), cow (CK846448), sheep (EE864720), and even chicken (DR417186). SwissProt provides full length protein Q93073 without providing a supporting accession.
Transcription and its processing in mammals is a noisy process and artefacts abound. While exon-skipping in some cases may have physiological significance, lacking significant comparative genomics support, the null hypothesis is that they do not. Here the full length gene must contain exon 8 since it is quite conserved in amino acid sequence throughout tetrapods and assuredly ancestral. As all SECIS binding experiments to date used protein lacking exon 8; consequently we know nothing about the SECIS binding properties of intact protein.
Exon 8 comparative genomics: VGFRCRGHSTSSERRQNLQKRPDNKHLSSSQSHRSDPNSESLYFE Homo sapiens VGFRCRGHSTSSERRQNLQKRPDNKHLSSSQSHRSDPNSESLYFE Macaca fascicularis VGFRCRGHSTSSERRQNLQKRQDNKQLNPSQSHRSDSNSESLYFE Tupaia belangeri VGFRCRGHSTSSERRQNLQKRQDNKHLNSTQSHRSDPNSESLYFE Mus musculus VGFRCRGHSTSSERRQNLQKRQDNKHLNSTQSHRSDPNSESLYFE Rattus norvegicus VGFRCRGHSTSSERRQNLPKRQDNNKQLNASQSHRGDSNSESLYFE Canis familiaris VGFRCRGHSTSSERRQNLQKRQDNKQLNPSQSHRGNPNSESLYFE Equus caballus VGFKCRGHSTSSERRQNLQKRQDNKQLNPNQSHRSDPNSESLYFE Myotis lucifugus VGFRCRGHSTSSERRQNLQKRQDNKQLNPSQSHRGDPNSESLYFE Bos taurus VGFRCRGHSTSSERRQNLQKRQDNKQLNPSQSHRGDPNSESLYFE Ovis aries VGFRCRGHSTSSERRQNLQKKQDNKQLNSSQSHRGDPNSESLYFE Dasypus novemcinctus VGFRCRGHSTSSERRQNLQKRQDNKQLNPIQSQRGDPNSESLYFE Loxodonta africana VGFRCRGHSTSSERRQSLQKRQDNKPL-GNHSHRVETSSDPLYFE Monodelphis domestica SGFRCRGHSTSSERRQNLQKRHE-KPLTTSQSSRAEQSPEPLYFE Gallus gallus PAFRCRGHSTSSERRQNLQKKPE-KPVSSSQSSKREQSPGSLYFE Anolis carolinensis LGYRLRGQSTSSERRHNLQRKQDNKTGTPASSNKSGQSPDHLYFE Xenopus tropicali
KIAA0256 and SECISBP2 actually align moderately well over their entire lengths and have 17 near perfectly comparable exons (trillions:one odds for coincidence), meaning they reflect a segmental gene duplication. It is imperative to enforce exon boundaries to achieve true homological alignment of two proteins this diverged and so gappy N-terminally; structure-based alignment has different rules (allowing convergent evolution) and different goals.
The teleost fish Pimephales promelas has sufficient transcript coverage to allow recover of an accurate nearly full length KIAA0256 gene with a respectable 62% identity to human. (No fish has sufficient transcripts to recover full length SECISBP2.) That gene is shown below as homologically gapped exon-by-exon to human. Some early exons are quite well conserved over many billions of years of branch length, strong evidence that they retain an unknown function under fairly strong selection. However the gaps in other early exons are incompatible with retention of tertiary protein structure. No early pfam domain can be found.
>KIAA0256_pimPro Pimephales promelas (minnow) based on transcript tiling; exons by homology 0 MD-AGERK 0 0 DVKLSAEVEPFIPQKKGVEASLLPMSLCGEGGA----EPTQIPSYLITCYPFVQENQSNSR 2 1 QLPMYNGGDQRWQQLNPSPGGPYLAYPILSSPQPPVTSDYATYYHAIMPTPCPPVMGFYQPFPGPFAGPVPAG-VLNPVS-DCSDRPT---------SQRGRGVPRTPVLHK 0 0 QPMAQP-MRAKRPVMRSVAVQKEVCATGPDGRTKTVLLVDAAQQT 1 2 DFPGEASGSGAVRCVSDXASPQLWSNKARRXRTSQQ--ESSSEQGVSEADIDSDSGYCSPKH--NQGANNTSTNQHTPAA 0 0 A-VDAGVMT 1 2 A-VSWGNVSSQAVQK-PWPDRNTPFFRGSRTPERSYTQDFQ 0 0 MSFGCRA---AGPRRSTPPETP-NTHLT--------P--EPLYFQ 0 0 DEDEFPDLATGGAAQRNKPDPVQPKLPKTL 0 0 LDNLPENSPISIVQTPIPITSSVPKRAKSQRKKALAAALATAQEYSEISMEQKKLQ 0 0 EALSKAAGKKSRTPVQLDLGDMLAALEKQQQAMRARQLNNTKPLSYT 1 2 VGTVSALHSKDCGSRVTGLKNTHT-PPHNILDSSAPRIKRGKEREIPKVKKTTAMKK 0 0 IILQEREVKKGKSSADQGVSGADEQRDS-LSFTDTLTQE---QD 1 2 ENGLSMPSDASLSPASQNSPYSITPVSQGSPASSGIGSPMAASAITKIHSRRFRE 2 1 YCNQVLSKDIDESVTLLLQELVRFQERVYQNEPSKAKAKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 ALFNTLVSLTEEARRAYKEMVSALEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 2 ETNWRTMVENADAPEPPDSEP-ISRGNNRDQREVVSP---PP---QPT--ANQSLTP--SPGVARAPD--ESRTDDRLEWASL-STETGSLDGSGRDRLNSSHHSTTSTLVPGMLEEE...* 0
There is no reason to believe an odd fragment studied on a few SECIS sequences could accurately represent binding properties of full length protein in regards to the 25-odd orthology classes of SECIS elements. However these results became accepted folklore within the selenocysteine research community and the properties of full length KIAA0256 were never experimentally determined.
We have to wonder how sea urchin, which has a full length apparent ortholog of KIAA0256 on Scaffold18963:101,648-115,302 but nothing clustering to SECISBP2, can insert selenocysteine into its numerous selenoproteins (SEPHS1, SELU1, SELU2, SELM, SELO, SELW, SELN1, GPX3, GPX2, GPX4, GPX7,...). Unless a second copy has been lost, all SECIS interaction at the ribosome at sea urchin divergence appears to have been handled by KIAA056.
Using the kink-turn binding motifs of the two human proteins in turn as blastp query against the both collections of deuterostome KIAA0256 and SECISBP2 sequences, establishes KIAA0256 as the slower evolving protein by a wide margin. This fits KIAA0256 retaining ancestral function and its gene duplicate SECISBP2 specializing to a neofunctionalization. It's difficult to extend the table into invertebrates because of high divergence even in the L7ae region. The table shows Blastp score ratios KIAA0256/SECISBP2 relative to human query:
galGal 1.41 72% identity anoCar 1.35 xenTro 1.41 68% identity danRer 1.44 tetNig 1.59 takRub 1.45 64% identity gasAcu 1.60 oryLat 1.52 calMil 1.43 65% identity
It has not been previously noted that both proteins bristle with potential NxT/S x not P glycosylation sites, 13 for KIAA0256 and 6 for SECISBP2, with implications for cellular localization. These do not lie in homologous positions, unsurprisingly in view of the deep divergence of these genes and volatility of glycosylation sites as seen in other gene families, eg the 17 mammalian sulfatases. Even within orthologs of one gene here, they are conserved only to moderate depth (and that could be for reasons unrelated to glycosylation). Hence these site do not provide reliable anchors in region of poor sequence conservation.
Potential for phosphoserine conservation in exon 5 of KIAA0256: DFPSDIANKSLSETTATMLWKSKGRRRRASHPTAESSSEQGASEADIDSDSGYCSPKHSNNQPAAGALRNPDSGTMN homSap .FPSDIANKSLSESTATMLWKAKGRRRRASHPAVESSSEQGASEADIDSDSGYCSPKH-NNQSAPGALRDPASGTMN musMus DFPSDIANKSLSESSATMLWKSKGRRRRASHPTAESSSEQGASEADIDSDSGYCSPKHSNNQPAAGALRNPDSSTMN canFam DFPSDIANKSLSESSSTMLWKSKGRRRRSSHPTAESSSEQGASEADIDSDSGYCSPKHSNNQATAMTSRNTDSGSIN monDom DFPLDIANKSLSESAATVLWKSKGRRRRASHPAAESSSEQGASEADIDSDSGYCSPKHGNNQAAGPAARSADSGPAN ornAna G insertion DFPSDIANKSLSESASTMLWKSKGRRRRASHPAAESSSEQGASEADIDSDSGYCSPKHGNNQAAAVTSRNADSCAMN galGal DFPSEIASKSLSESMSTMHWKPKTRRRRSSHP-AESSSEQGASEADIDSDSGYCSPKHS-NQAAAVTSRSVESAAGN anoCar DFPNEIANKTICESVGATPWKSKVRRRRLSHPAAESSSEQGASEADIDSDSGYCSPKHC--QAAAMCTRHADCGAV. xenTro DFPGEASGGVRCVSDQVSPQQWKNKPRRRRTSQQESSSEQGASEADIDSDSGYCSPKH--NQGAA............ danRer DFPGEVSGRCAAERASPQLWKNKTKRRRASHP-AENYSEQGASEADIDSDSGYCSPKH--NQAAGVTQR........ gasAcu DFPGEAAVRCVSDQASPQLWSNKARRRRTSQ--QESSSEQGVSEADIDSDSGYCSPKHSTNQPAAAV----DAGVM pimPro SGSG NQGANNT HT insertions DFPDDIADKSLRDKPSPLLRKSKARRLASRRPQDPSSTDSEEDEGGIDSDSGYSSPKHGRNQSA..............braFlo DFPEAIANKPLSDKTSNLTSRSKAKTRKKSQGNASSSSDSEVENTPHDSDSGYYSPLHAQQ................ strPur QTGRD insertion Comparative genomics of 4 glycosylation sites in exon 7 of KIAA0256: GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ Homo sapiens (human) GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ Macaca mulatta (rhesus) GGVNWPKVTCQATQKRPWMEKNQAFSRGGRQTEQRNNLQ Mus musculus (mouse) GGVNWPKVTCQATQKRPWMEKNQAFSRGGRQTEQRNNSQ Rattus norvegicus (rat) GSVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ Canis familiaris (dog) GGVNWSNVTSQATQKKPWMEKNQTFSRGGRQAEQRNNSQ Sus scrofa (pig) GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ Equus caballus (horse) GGVNWSNVTCQGTQKKPWLEKNQTFSKGGRQMEQRNNSQ Dasypus novemcinctus (armadillo) GHVNWSNVTCQATQKKPWMEKHQTFSRGGRQTEQRNNAQ Loxodonta africana (elephant) GGASWSNVTSQATQKKPWMEKSQPFSRGGRQTEQRNNSQ Monodelphis domestica (opossum) .GVSWTNVNSQATQKKPWIEKTQTFIRGGRQAEQRNSSQ Gallus gallus (chicken) AGATWANVSSQATQKKPWMERTPAFSRGGRQAEQHNSSQ Anolis carolinensis (lizard)
Reference set of 27 vertebrate SECIS BP2 L7Ae motif exons
>SECISBP2_homSap Homo sapiens (human) full length 0 MASEGPREPESE 0 0 GIKLSADVKPFVPRFAGLNVAWLESSEACVFPSSAATYYPFVQEPPVTE 2 1 QKIYTEDMAFGASTFPPQYLSSEITLHPYAYSPYTLDSTQNVYSVPGSQYLYNQPSCYRGFQTVKHRNENTCPLPQEMKALFK 0 0 KKTYDEKKTYDQQKFDSERADGTISSEIKSARGSHHLSIYAENSLKS 1 2 DGYHKRTDRKSRIIAKNVSTSKPEFEFTTLDFPELQGAENNMSEIQKQPKWGPVHSVSTDISLLREVVKPAAVLSK 0 0 GEIVVKNNPNESVTANAATNSPSCTR 1 2 ELSWTPMGYVVRQTLSTELSAAPKNVTSMINLKTIASSADPKNVSIPSSEALSSDPSYNKEKHIIHPTQK 0 0 SKASQGSDLEQNEASRKNKKKKEKSTSKYEVLTVQEPPRIE 0 0 DAEEFPNLAVASERRDRIETPKFQSKQQPQ 0 0 DNFKNNVKKSQLPVQLDLGGMLTALEKKQHSQHAKQSSKPVVVS 1 2 VGAVPVLSKECASGERGRRMSQMKTPHNPLDSSAPLMKKGKQREIPKAKKPTSLKK 0 0 IILKERQERKQRLQENAVSPAFTSDDTQDGESGGDDQFPEQAELS 1 2 GPEGMDELISTPSVEDKSEEPPGTELQRDTEASHLAPNHTTFPKIHSRRFRD 2 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTVAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALKEKEEPHY 1 2 IEIWKKHLEAYSGCTLELEESLEASTSQMMNLNL* 0 >SECISBP2_panTro Pan troglodytes (chimp) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQQELVGEPRPQAPPSLPTQGPSCPAEDGPPALTEKEEPHY 1 >SECISBP2_macMul Macaca mulatta (rhesus) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQQELAGEPRPQAPPSPPTQGPSCPAEDGPPALTEKEEPHY 1 >SECISBP2_otoGar Otolemur garnettii (bushbaby) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKERRLVLGLREVLKHLKLKKLICVISPNCERQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVQRELAGEPGPQVPSSLPMEGPSCSVEDSPPAPTEKEEPHY 1 >SECISBP2_tupBel Tupaia belangeri (tree_shrew) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRVVLGLREVLKHLKLKKLKCVIISPIZEKIQSK 1 2 GGLDDTLHTIIAYACAQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMEARQAYRSMLESARQELAGEPGLQAPPQPPVQGPRASSEGSAPAPTGRQEPHC 1 >SECISBP2_musMus Mus musculus (mouse) 1 YCSQMLSKEVDACVTGLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKTQSK 1 2 GGLDDTLHTIIDCACEQNIPFVFALNRKALGRSLNKAVPVSIVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLETMRQEQAGEPGPQSPPSPPMQDPIPSTEEGTLPSTGEEPHY 1 >SECISBP2_ratNor Rattus norvegicus (rat) exons 1416 1 YCSQMLSKEVDACVTGLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKTQSK 1 2 GGLDDTLHTIIDCACEQNIPFVFALNRKALGRSLNKAVPVSIVGIFSYDGAQDQ 0 0 FHKMVELTMAARQAYKTMLETMRQEQAGEPGPQTPPSPPMQDPIQSTDEGTLASTGEEPHY 1 >SECISBP2_cavPor Cavia porcellus (guinea_pig) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCIIISP 1 2 GLDDTLHTIIDYACAQNIPFVFALNRKALGRSLNKTVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENVRQELAGEPRPQMPPDPPSEGPSSSLEDTAPDPSAEEPHY 1 >SECISBP2_oryCun Oryctolagus cuniculus (rabbit) 1 YCSQMLSKEVDACVTDLFKELVRFHDLMYQDPVKATTKCQFELRVGKALDHLRLKKLKCIIVFPKHKKQS 1 2 TIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKTMLENMRHELAGEPGPPTPQPVQGPSCSAEDGPPAPTEGEVPHY 1 >SECISBP2_canFam Canis familiaris (dog) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHRMVELTMAARQAYKTMLENVRQELAGEPGTPALANPPMQGLGCSTQDSPPAPTEKEEPHY 1 >SECISBP2_felCat Felis catus (cat) 1 YCSQMLSKEVDACVTDLLRELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIGYACEQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHRMVELTMAARQAYKTMLENARQELAGEPGPPAPGSPPPQPPAPAGRDEPRY >SECISBP2_equCab Equus caballus (horse) 1 YCSQILSKEVDACVTELLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPCVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTKAARQAYKAMLENVHQELAGEPGPQAPASPPAQGPSCSTEGAPPAPTGKEEPHY 1 >SECISBP2_bosTau Bos taurus (cow) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKAKRRLVLGLREVLKHLKLRKLKCIIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACDQNIPFVFALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYRTMLENARQELPGELGPCAPVGPPSQGPGCPVEDSPLAPTEKEEPHY 1 >SECISBP2_eriEur Erinaceus europaeus (hedgehog) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCIIISPNCEKIQSK 1 2 GGLDETLHTIIDCACEQNIPFVFALNRKALGRSLNKGVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKALLENMRQELAEESGSPAPSSPPVQSPSEDGPPAPAEKEEPHY 1 >SECISBP2_dasNov Dasypus novemcinctus (armadillo) 1 YCSQVLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGELDDTLHTIIDYAASRHSICVALNRKALGRSLNKAVPVSVVGIFSYDGAQ 0 0 DQFHKMVELTMAARQAYKAMLENVRKELAGEPGPRSPPSPPALGPHSSAGDVHPTSAGKEEPHY 1 >SECISBP2_loxAfr Loxodonta africana (elephant) 1 YCSQMLSKEVDACVTDLLKELVRFQDRMYQKDPVKAKTKRRLVLGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDDTLHTIIDYACEQNIPFVFALHRKALGRSLNKPVPVSVVGIFSYDRAQ 0 0 DQFHKMVELTMAARQEYKTMLESVRQELAEEPRAGSPPSPPTQGPGCSAEVPRPAPTEKEEPRY 1 >SECISBP2_monDom Monodelphis domestica (opossum) 1 YCSQMLSKEVDDCVMDLLKELVRFQDRMYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKSKSK 1 2 GGLDETLHTIIDYACEQNVPFVFALNRKALGRSVNKVVPVSVVGIFSYDGAQ 0 0 DQFHKMIALTMEARQAYKIMLSTLKEEPALETENPPSPSLPRPSESCPSELGQTDPTQEEEPNY 1 >SECISBP2_triVul Trichosurus vulpecula (possum) 1 YCSQMLSKEVDDCVMDLLKELVRFQDRMYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKSKSK 1 2 GGLDETLHTIIDYACEQNVPFVFALNRKALGRSVNKVVPVSVVGIFSYDGAQ 0 0 DQFRKMIELTMEARQAYKVMLATLKEGAEALQTENPLPTSLTPQGQGCSSELSKTTDPTKEEEPNY 1 >SECISBP2_galGal Gallus gallus (chicken) 1 YCSQVLSKEVDSCVTDLLKELVRFQDRLYQKDPVKAKIKRRLVMGLREVLKHLRLKKLKCVIISPNCEKIQSK 1 2 GGLDETLHNIIDCACEQNIPFVFALNRKALGRCVNKAVPVSVVGIFSYDGAQ 0 0 DHFHRMVQLTTEARKAYKDMVAALEEELKELSKPLNZKSCLSETGKTSSTKEDIPNY 1 >SECISBP2_anoCar Anolis carolinensis (lizard) 1 YCTQVLSKEVDSCVTDLLKELVRFQDRLYQKDPVKAKTKRRLVMGLREVLKHLKLKKLKCVIISPNCEKIQSK 1 2 GGLDETLHLIIDSACEQNIPFVFALNRKALGRCLNKAVPVSVVGIFSYDGAQ 0 0 DYFHKMVELTMEARQAYKDMISALERELKKKTVRKKPLQSRPLDTVEASSTEEDVPDY 1 >SECISBP2_xenTro Xenopus tropicalis (frog) NM_001097262 1 YCSQVLSKDVDNCVMELLKELVRFQDRLFLKEPAKAKSKRRLVMGLREVLKHLKLQKLKCIIISPNCEKIQSK 1 2 GGLDDTLQTIISHACEQNVPFVFALNRKALGRCLNKAVPVSVVGVFSYDGAQ 0 0 DHFHKLCELTVQARQAYKDMIAAAQEQQSETEAGKNEEDPVAVNGQNKSDDMREESKAEEPDEPNY 1 >SECISBP2_danRer Danio rerio (zebrafish) 1 YCNQVLSKDVDECVSNLLKELVRFQDRLYQKDPMKARMKRRLVMGLREVLKHLKLKKVKCVIISPNCERIQSK 1 2 GGLDEALHNIIDTCRDQSVPFVFALSRKALGRCVNKAVPVSLVGIFNYDGAQ 0 0 DFYHKMIELSSEARTAYEVMLLNLEQTDAEEAQQTSPLAEKVETSSGDPQPEEPEY 1 >SECISBP2_tetNig Tetraodon nigroviridis (pufferfish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DFYHKMIELSSEARIAYEVMLSNLEQTSAEEEPQTCTLAEKINTSSEDAQPEEPEY 1 >SECISBP2_takRub Takifugu rubripes (fugu) 1 YCTQMLSKDVDECVTTLLKELVRFQDRLYQKDPIKARMKRRIVMGLREVQKHLKLRKLKCVIISPNCERIQSK 1 2 GGLDEALHTIIDTCREQAVPFVFALSRRALGRCVNKAVPVSLVGIFNYDGAQ 0 0 DFYHKMIELSSEARTAYEVMLLNLEQTDAEEAQQTSPLAEKVETSSGDPQPEEPEY 1 >SECISBP2_gasAcu Gasterosteus aculeatus (stickleback) 1 YCNQVLSKEIDESVTMLLQELVRFQERIYQKDPTKAKTKRRLVMGLREVTKHMKLNKIKCVLISPNCEKIQAK 1 2 GGLDEALYNVIAMARDQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DFYHKMIELSSEARRAYEVMVSSLEQTGQADPESVEEKLQISSAAEEAELGRDITPPEEPEY 1 >SECISBP2_oryLap Oryzias latipes (medaka) 1 YCSQMLRKDVDECVTVLLKELVRFQDRLYHKDPIKARMKRRLVMGLREVLKHLKLRKVKCVIISPNCEQIQSK 1 2 GGLDEALHTIIQTCREQAVPFVFALSRKALGHCVNKAVPVSLVGIFNYDGAQ 0 0 DHYHKMIELSAEARKAYEVLVSSLERDQQEESHPDRGTCFGSVTAEPEKPHY 1 >SECISBP2_calMil Callorhinchus milii (elephantfish) AAVX01044988 1 YCSQVLSKDVDSCVTDLLKELVRFQDRLYQKDPIKAKKKRRIVMGLREVLKHLKLKRLKCIIISPNCEKIQSR 1 2 GGLDDALHNIISIACEQEIPFVFALNRKALGQCVNKPVPVSVLGIFSYDGAE 0 0 NQFHQMVEITEEARKAYQEMLDALQQELEADEEKGDSEEQPLISSESSTIHFNNVTSQPFSEADEPEY 1 >SECISBP2_braFlo Branchiostoma floridae (amphioxus) extra exon 1 YCNQVLDKEIDATVTMLLQDLVRFQDRQYHK 00 DPIKAKAKRRIVMGLREVTKHLKLRKLKCIIIAPNLEKIQSK 1 2 GGLDDAIETILNLCMEQDVPFVFALGRKALGRAVNKLVPVSVVGVFNYDGAE 0 0 1
Reference set of 23 deuterostome SECIS KIAA0256 L7Ae motif exons
>KIAA0256_homSap Homo sapiens (human) length=1101 13 glycosylation sites 0 MDRAPTEQ 0 0 NVKLSAEVEPFIPQKKSPDTFMIPMALPNDNGSVSGVEPTPIPSYLITCYPFVQENQSNR 2 1 QFPLYNNDIRWQQPNPNPTGPYFAYPIISAQPPVSTEYTYYQLMPAPCAQVMGFYHPFPTPYSNTFQAANTVNAITTECTERPSQLGQVFPLSSHRSRNSNRGSVVPK 0 0 QQLLQQHIKSKRPLVKNVATQKETNAAGPDSRSKIVLLVDASQQT 1 2 DFPSDIANKSLSETTATMLWKSKGRRRRASHPTAESSSEQGASEADIDSDSGYCSPKHSNNQPAAGALRNPDSGTMN 0 0 HVESSMCA 1 2 GGVNWSNVTCQATQKKPWMEKNQTFSRGGRQTEQRNNSQ 0 0 VGFRCRGHSTSSERRQNLQKRPDNKHLSSSQSHRSDPNSESLYFE 0 0 DEDGFQELNENGNAKDENIQQKLSSKV 0 0 LDDLPENSPINIVQTPIPITTSVPKRAKSQKKKALAAALATAQEYSEISMEQKKLQ 0 0 EALSKAAGKKNKTPVQLDLGDMLAALEKQQQAMKARQITNTRPLSYT 1 2 VVTAASFHTKDSTNRKPLTKSQPCLTSFNSVDIASSKAKKGKEKEIAKLKRPTALKK 0 0 VILKEREEKKGRLTVDHNLLGSEEPTEMHLDFIDDLPQEIVSQE 1 2 DTGLSMPSDTSLSPASQNSPYCMTPVSQGSPASSGIGSPMASSTITKIHSKRFRE 2 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 2 ETNWRNMVETSDGLEASENEKEVSCKHSTSEKPSKLPFDTPPIGKQPSLVATGSTTSATSAGKSTASDKEEVKPDDLEWASQQSTETGSLDGSCRDLLNSSITSTTSTLVP GMLEEEEDEDEEEEEDYTHEPISVEVQLNSRIESWVSETQRTMETLQLGKTLNGSEEDNVEQSGEEEAEAPEVLEPGMDSEAWTADQQASPGQQKSSNCSSLNKEHSDSNYTTQTT* 0 >KIAA0256_panTro Pan troglodytes (chimp) 2 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_macMul Macaca mulatta (rhesus) 2 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_tupBel Tupaia belangeri (tree_shrew) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_musMus Mus musculus (mouse) 0 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 2 SLFNRLVELTEEARKAYKDMVAATEQEQAEEALRSVKTVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_ratNor Rattus norvegicus (rat) 2 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 0 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 1 SLFNRLVELTEEARKAYKDMVAATEQEQAEEALRSVKAVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_canFam Canis familiaris (dog) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_equCab Equus caballus (horse) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVALTEEARRAYKDMVAALEQEQAEEASKNVKKGPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_dasNov Dasypus novemcinctus (armadillo) 1 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSKG 1 2 GLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_monDom Monodelphis domestica (opossum) 0 YCNQVLCKEIDECVTLLLQELVSFQERIYQKDPVKAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYCGAE 0 2 SLFNKLVELTEEARKAYKDMVAAMEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_galGal Gallus gallus (chicken) 1 YCNQVLSKEIDECVTLLLQELVSFQERIYQKDPMRAKARRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 DLFNKLVSLTEEARKAYRDMVAAMEQEQAEEALKNVKKAPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_anoCar Anolis carolinensis (lizard) 1 YCNQVLSKEIDECVTLLLQELVSFQEQIYQKDPMRAKAKRRLVMGLREVTKHMKLSKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 NLFNKLVSLTEEARKAYRDMVAAMEQEQEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_xenTro Xenopus tropicalis (frog) 1 YCNQVLSKEIDECVTVLLQELVSFQERVYQKDPVKAKSKRRLVMGLREVTKHMKLNKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFSYSGAE 0 0 SLFHNLVSLTEEARKAYKDMVSSMEQEQAEEALKNIKKVHMGHSRNPSAASAISFCSVISEPISEVNEKDY 1 >KIAA0256_danRer Danio rerio (zebrafish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKEPSKAKAKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 2 GGLDEALHNIIDTCRDQSVPFVFALSRKALGRCVNKAVPVSLVGIFNYDGAQ 0 0 GLFNKLVSLTEEARRAYKEMVSALEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 >KIAA0256_tetNig Tetraodon nigroviridis (pufferfish) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 SLFNQLVSLTEEARKAYKDMVSALEQEQTEEALKNEKKVPHQMGHYRNHSAASAVSFCSIFSEPISEVNEKEY 1 >KIAA0256_takRub Takifugu rubripes (fugu) 1 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPTKAKSKRRLVMGLREVTKHMKLQTIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNFSGAE 0 0 SLFNQLVSLTEEARKAYKDMVSALEQEQTEEALKNEKKVPHQMGHYRNHSAASAVSFCSIFSEPISEVNEKEY 1 >KIAA0256_gasAcu Gasterosteus aculeatus (stickleback) 1 YCNQVLSKEIDESVTMLLQELVRFQERIYQKDPTKAKTKRRLVMGLREVTKHMKLNKIKCVLISPNCEKIQAK 1 2 GGLDEALYNVIAMARDQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 GLFNRLVSLTEEARKAYKDMVSALEQEQAEEAQKNDKKLPHHMGHSRNHSAASAISFCSIFSEPISEVNEKEY 1 >KIAA0256_oryLap Oryzias latipes (medaka) 0 YCNQVLSKEIDESVTLLLQELVRFQERVYQKDPSKAKSKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 1 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 2 GLFNQLVSLTEEARKAYKEMVSALEQEQAEEALKHDKKVPHHMGHSRNHSAASAISFCSILSEPISEVNEKEY 1 >KIAA0256_pimPro Pimephales promelas (minnow) based on transcript tiling; exons by homology; 62% identity 0 MDAGERK 0 0 DVKLSAEVEPFIPQKKGVEASLLPMSLCGEGGAEPTQIPSYLITCYPFVQENQSNSR 2 1 QLPMYNGGDQRWQQLNPSPGGPYLAYPILSSPQPPVTSDYATYYHAIMPTPCPPVMGFYQPFPGPFAGPVPAGVLNPVSDCSDRPTSQRGRGVPRTPVLHK 0 0 QPMAQPMRAKRPVMRSVAVQKEVCATGPDGRTKTVLLVDAAQQT 1 2 DFPGEASGSGAVRCVSDXASPQLWSNKARRXRTSQQESSSEQGVSEADIDSDSGYCSPKHNQGANNTSTNQHTPAA 0 0 AVDAGVMT 1 2 AVSWGNVSSQAVQKPWPDRNTPFFRGSRTPERSYTQDFQ 0 0 MSFGCRAAGPRRSTPPETPNTHLTPEPLYFQ 0 0 DEDEFPDLATGGAAQRNKPDPVQPKLPKTL 0 0 LDNLPENSPISIVQTPIPITSSVPKRAKSQRKKALAAALATAQEYSEISMEQKKLQ 0 0 EALSKAAGKKSRTPVQLDLGDMLAALEKQQQAMRARQLNNTKPLSYT 1 2 VGTVSALHSKDCGSRVTGLKNTHTPPHNILDSSAPRIKRGKEREIPKVKKTTAMKK 0 0 IILQEREVKKGKSSADQGVSGADEQRDSLSFTDTLTQEQD 1 2 ENGLSMPSDASLSPASQNSPYSITPVSQGSPASSGIGSPMAASAITKIHSRRFRE 2 1 YCNQVLSKDIDESVTLLLQELVRFQERVYQNEPSKAKAKRRLVMGLREVTKHMKLHKIKCVIISPNCEKIQAK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYSGAE 0 0 ALFNTLVSLTEEARRAYKEMVSALEQEQAEEALKNVKKVPHHMGHSRNPSAASAISFCSVISEPISEVNEKEY 1 2 ETNWRTMVENADAPEPPDSEPISRGNNRDQREVVSPPPQPTANQSLTPSPGVARAPDESRTDDRLEWASLSTETGSLDGSGRDRLNSSHHSTTSTLVPGMLEEE * 0 >KIAA0256_calMil Callorhinchus milii (elephantfish) AAVX01105236 1 YCNQVLSKDIDECVTLLLQELVRFQERVYQKDPIKAKMKRRLVMGLREVTKHMKLRKIKCVIISPNCEKIQSK 1 2 GGLDEALYNVIAMAREQEIPFVFALGRKALGRCVNKLVPVSVVGIFNYFGAE 0 0 1 >KIAA0256_petMar Petromyzon marinus (lamprey) 1 LHKLRALIISPNCEKIQAK 1 2 GGLDEALQTVIALASEQSVPFVFALNRKALGHCLNKKVPVSVVGVFHYGGAE 0 0 THFQRLVALTEEARSAYRNMVSSLQRQEAAATSEPTGHTEDPLEASASPPSVPAHDPTALLHLLRPQQGPREDDPAEASGRSPGRNA 1 >KIAA0256_cioInt Ciona intestinalis (tunicate) 1 YCCQVLDKRVDEMSNQMLQRLVYFQDR 21 RLYKTDPAKAKRKRRVVLGFREVTKHLKMKKLRCVIISPNLEKIESK 1 2 GGLDDVLHEILDLCKEQNIPYVFALGKKALGRAVSKTVPVSIVGVFDYSGAE 0 0 1 >KIAA0256_strPur Strongylocentrotus purpuratus (sea_urchin) 1 YCNQVLDKDIDGCCTTLLQTLVKFQDRQYHKDPAK 00 AKMKRRLVMGLREVTKHLKLKKIKCVVVSPNLERIQSK 1 2 GGLDEAMDRISSLASEQNVPLIFALGRKALGRAVNKVVPVSVVGIFNYDGAE 0 0 DTYKQLLDLSTRARNAYADMVRKFQQELEAANAASAARMAKHRHHMGHNRNLFKG 1
Reference set of 10 deuterostome ribosomal L30 L7Ae motifs
>L30_homSap Homo sapiens (human) 4 exons numerous pseudogenes 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_tupBel Tupaia belangeri (tree_shrew) 0 MVAAKKT 0 0 KKSLESINSQLQLAMKDGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRACTLAIMDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_ratNor Rattus norvegicus (rat) Sep15 Gpx4 Gpx1 Dio1 quite weak homology 35% with BP2 exons 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQTGEK* 0 >L30_myoLuc Myotis lucifugus (microbat) 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYLLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 ISEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSD-IRSMPEQTGEK* 0 >L30_echTel Echinops telfairi (tenrec) 0 MVAAKKT 0 0 KNSLESINSRLQLVMKSGKYMLGYKQMLKMIRQGKAKLVVLANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGHNIELGTACGKSCRVCTLAITDP 1 2 GDADIIRSMPEQTGEK* 0 >L30_anoCar Anolis carolinensis (lizard) 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIQQGKAKLVILANNCPALG 2 1 KSEIEYYAMLAKTGVHHYSGNNIEMGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMQEQTAEK* 0 >L30_danRer Danio rerio (zebrafish) 94% 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQSQKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPDQQQGGEK* 0 >L30_squAca Squalus acanthias (spiny dogfish) 97% 0 MVAAKKT 0 0 KKSLESINSRLQLVMKSGKYVLGYKQTLKMIRQGKAKLVILANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQISEK* 0 >L30_petMar Petromyzon marinus (lamprey) 94% 0 MSAKKT 0 0 KKAIESINSRLQLVMKSGKYCLGYRQTLKMIRQGKAKLVLLANNCPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIEMGTACGKYYRVCTLAIIDP 1 2 GDSDIIRSMPEQQQPQPGDK* 0 >L30_braFlo Branchiostoma floridae (amphioxus) 84% to homSap 0 MKQK 0 0 RKTMESINSRLQLVMKSGKYVLGLKETLKVLRQGKAKLIIIANNTPALR 2 1 KSEIEYYAMLAKTGVHHYSGNNIELGTACGKYFRVCTLAITDP 1 2 GDSDIIRSMPAEDKGESK* 0
Positioning of SECIS elements in 3' UTR
Reference sets of vertebrate SECIS elements
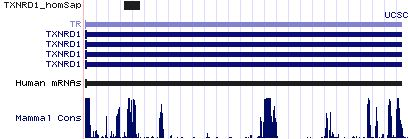
TXNRD1: 17 SECIS insertion sequences
>TXNRD1_homSap Homo sapiens (human) COVE score: 25.36 auuuggcagg gcaucga agggaugcaucc augaa gucaccagucuc aa gcccaugugg uaggcggugau ggaa caacug ucaaauc aguuuuagca >TXNRD1_panTro Pan troglodytes (chimp) COVE score: 25.36 auuuggcagg gcaucga agggaugcaucc augaa gucaccagucuc aa gcccaugugg uaggcggugau ggaa caacug ucaaauc aguuuuagca >TXNRD1_otoGar Otolemur garnettii (bushbaby) COVE score: 23.42 aucggcagug caucgac gggaugcgucc augaa gucaccagccuc aa gccugugugg ugggcagugau ggaa caacu guccaau caguuucuau >TXNRD1_tupBel Tupaia belangeri (tree_shrew) COVE score: 25.13 ucuuggcagc gcaucag agggaugcgucc augaa gucaccagccuc aa gcccgugcgg ugggcggugau ggaa cgacug ccagauc aguuucagca >TXNRD1_musMus Mus musculus (mouse) COVE score: 27.9 aucuggcaga gcaucac aggcaugcgucc augaa gucacuggccuc aa gcccaagug gugggcagugac agaa gagcu gccgggu cuguugagcu >TXNRD1_ratNor Rattus norvegicus (rat) COVE score: 28.03 uucggcagag caucacg gugcgucc augaa gucacuagccuc aa gcccaagugg ugggcagugac agaa agc ugucgau cuguuggguu >TXNRD1_cavPor Cavia porcellus (guinea_pig) COVE score: 23.88 gcggcggcac cguaggg ugcgucc augag gucaccagccuc aa gcccgaggg ugggcggugac ggau cgc gccgcg uggcucagcu >TXNRD1_eriEur Erinaceus europaeus (hedgehog) COVE score: 26.25 ugucagcaga gcaucaa agggaugcgucc augaa gucaccagccuc aa gcccgugcggg ugggcagugac ggaa cacug ucgaagc aguuucaaca >TXNRD1_canFam Canis familiaris (dog) COVE score: 24.54 auucggcaug caucggc guggugcgucc augaa gucacuggccuc aa gccaugcg gugggcagugau ggag caac ugucgag caguuuuagu >TXNRD1_felCat Felis catus (cat) COVE score: 24.14 acucggcagc gcaucgg agggcgcgucc augaa gucaccggcccc aa gcccccgcg gugggcggugau ggaa caagu gccgagc aguuuuagcg >TXNRD1_equCab Equus caballus (horse) COVE score: 21.76 acucggcagu gcaucga agggaugcgucc augaa gucacuggccuc aa agcccaugug gugggcggugau ggaa cagcug ucgaagc aguuuuagca >TXNRD1_loxAfr Loxodonta africana (elephant) COVE score: 22.55 auuggcagcg caucgag ggaugcaucc augaa gucacuggccuc aa gcccaugug gggggcggugau ggaa cagcu gucgaau cagcuuuggc >TXNRD1_echTel Echinops telfairi (tenrec) COVE score: 19.5 guggcagugc aucaaga gaugcguuc augaa aucgcuugcccc aa gcccga guggcgggcag cgau ggaaca ucugucu caucaguuuc >TXNRD1_monDom Monodelphis domestica (opossum) COVE score: 23.46 ggcucgcggu gcaucgg ugagaugcguuc augaa gucgcugccug aa gcccauaucccgugg uggguggugac cgaa agaaccg ccggcc uccguuuuau >TXNRD1_ornAna Ornithorhynchus anatinus (platypus) COVE score: 18.89 aggagugcac ccaaggg cugcauuu augaa gucagagccaa aa gccagcauuuugcgg uuggcugugau ggaa aaa cuccug ccacaguuuu >TXNRD1_anoCar Anolis carolinensis (lizard) COVE score: 32.53 uggcaaggca uuguuca agaugcuucc augaa gucacagucua aa accagugcuuucugg uaggcagugau ggaa aga uugcugg cacaacuuga >TXNRD1_galGal Gallus gallus (chicken) COVE score: 26.64 uagcagggca uuucaca caugcuuuc augaa aucacagccug aa gccugcacugucugg ugggcagugau ggaa gaacu gcugaca cagcugaaca
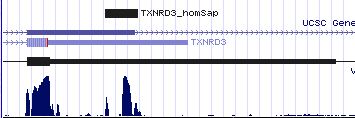
TXNRD3: 9 SECIS insertion sequences (problematic)
>TXNRD3_homSap Homo sapiens (human) COVE score: 19.10 gacagcgaga agcagug ggacugcuucc uugac gccuuagcuu gg agccccguuaugaggu gagccaaggc ugac ucu cgcaagc caggacugag >TXNRD3_panTro Pan troglodytes (chimp) COVE score: 19.10 agcagugggc GACAGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUU GG AGCCCCGUUAUGAGGU GAGCCAAGGC UGAC UCU CGCAAGC CAGGACUGAG >TXNRD3_macMul Macaca mulatta (rhesus) COVE score: 17.86 GACuGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUU GG AGCCCuGUUgUGAGGU GAGCCAAGGC cGAC UCU CGCAAGC CAGGACUcAG >TXNRD3_calJac Callithrix jacchus (marmoset) COVE score: 25.26 GACuGCGAGA AGCAGUG GGACUGCUUCC UUGAC GCCUUAGCUc Ga AGCCCuGUUAcGAGGU GAGCCAAGGC UGAu UCU CGCAAGC gAGGACUGAG >TXNRD3_musMus Mus musculus (mouse) COVE score: 21.86 CUGACGGCAU GCAGCAG CCAGGCUGCUUCC UUGAC ACCUUGGCUC GG AACCUGCAGAGGU GAGCCAAGGC CGAC UUC UGCACGU CAGCCUCGAC >TXNRD3_ratNor Rattus norvegicus (rat) COVE score: 25.37 cugauggcgu gcagcag ccaggcugcaucc uugac gccuuggcuc gg aaccugcagagg ugagccaaggc cgac ucc ugcacgu cagccucgac >TXNRD3_canFam Canis familiaris (dog) COVE score: 26.93 GCUGGCUGGA GAGGCAG GCAGGCUGCCUCC UUGAC GCCUUAGCUC GG AACCGCUGUGAGG UGAGCUAAGGC CGAU GUC CUCCAU GCCAGGCCAG >TXNRD3_felCat Felis catus (cat) COVE score: 20.14 CUGGCUgGGA GAGGCAG GucGGCUGCCUCC UUGAC GuCUUAGCUC GG AgcccgaUGUGAGG UGAGCUAAGGC CGAU Ggu cuuccac gucagaaucg >TXNRD3_bosTau Bos taurus (cow) COVE score: 12.55 GGCUGACGGG AGGGCAG ACUCGCUGCCUCC UUGAC GUCUUCGCUC AG AGCCGCCAGGU GAGCCAAGAC CGAC CUC UGCCCA CCAGCUCCUC
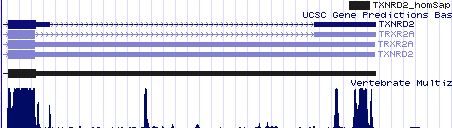
TXNRD2: 10 SECIS insertion sequences
>TXNRD2 homSap Homo sapiens (human) COVE score: 31.00 cacccccccc caggcuc cuggugccagaug augac gaccugggugg aa accuacccugugg gcacccauguc cgag ccccc uggcauu ucugcaaugc >TXNRD2 panTro Pan troglodytes (chimp) COVE score: 31.00 ggcacccccc caggcuc cuggugccagaug augac gaccugggugg aa accuacccugugg gcacccauguc cgag ccccc uggcauu ucugcauugc >TXNRD2 tupBel Tupaia belangeri (tree shrew) COVE score: 36.55 cucccucccc aggcccc cgaugccagaug augac ggccuggacag aa acccacccuguggg cugcccagguc ugaa cccuccc uggugucu uuggggugua >TXNRD2 musMus Mus musculus (mouse) COVE score: 36.94 gccagccucu gacacuc ccagcgucagaug augau ggccugggcag aa accccauguggg ccgcccagguu ugaa ccccu ggcauuu cuagagcacu >TXNRD2 ratNor Rattus norvegicus (rat) COVE score: 34.71 cagccuucac acacugc cagugucagaug augac ggccugugcag aa acccccacguggg cugcccagguu ugaa ccccug gcauuu cuggagugcu >TXNRD2 cavPor Cavia porcellus (guinea pig) COVE score: 34.41 ucuggccccc caggucc ccagugccaguug augau ggccugggcag aa acccacccuguggg caguccauguc ugaa cuccc uggcauu ucuggagugc >TXNRD2 eriEur Erinaceus europaeus (hedgehog) COVE score: 34.68 ccagccccac caggccc ccgaugccagaua augau gacuugugcag aa acccacccggg cugcccauguc ugag ccucug uggcauu cuggagugua >TXNRD2 canFam Canis familiaris (dog) COVE score: 34.36 accagccccg ccaggcc ccgaugccagaag augac gacgugugcag aa accccccuguggg cugcccgcguc cgag cccccuggc guuucugg aauguaaaua >TXNRD2 equCab Equus caballus (horse) COVE score: 31.37 ccagcccugc cagguuc ccgaugccagacg augac gaccugcgcgg aa acccacccuguggg cugcccacguc cgag ccccc uggcauu ucugaagugc >TXNRD2 echTel Echinops telfairi (tenrec) COVE score: 37.41 cccuugcccc caccuca gcgccagaug augaa gacaugugcag aa acccagcccguggg cugcccauguc ugag ccccc ugacgu uucuggagug
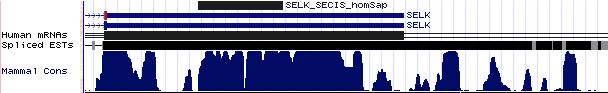
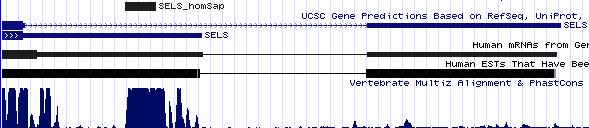
SELS: 20 SECIS insertion sequences
>SELS_homSap Homo sapiens (human) COVE score 36.61 uaggacaguc ucuguga cagguugcguuga augau gucuuccuuauc aa uggugagcccacca gugaggauuac ugau gugga caguuga ugggguuugu >SELS_panTro Pan troglodytes (chimp) COVE score 36.61 uaggacaguc ucuguga cagguugcguuga augau gucuuccuuauc aa uggugagcccacca gugaggauuac ugau gugga caguuga ugggguuugu >SELS_macMul Macaca mulatta (rhesus) COVE score 36.61 uaggacaguc ucuguga cagguugcguuga augau gucuuccuuauc aa uggugagcccacca gugaggauuac ugau gugga caguuga ugggguuugu >SELS_otoGar Otolemur garnettii (bushbaby) COVE score 38.23 uaggacaguc ucuguga cagguugcguuga augau gucuuccuuau aa auggugagcccacca gugaggauuac ugau acaga caguuga ugggguuugu >SELS_tupBel Tupaia belangeri (tree_shrew) COVE score 40.14 gagaacaguc ucuguga cagggugcgucga augau gucuuccuuau aa auggugagcccacca cugaggaguac ugau gcaga caguuga caggguuugu >SELS_musMus Mus musculus (mouse) COVE score 41.68 caggaugguc ucuguga cgggaugcguuga augau gucuuccuuau aa auggugaacccacca gugaggauuac ugau guu cacagu ugacgggguu >SELS_ratNor Rattus norvegicus (rat) COVE score 40.42 cagggugguc ucuguga caggaugcguuga augau gucuuccuuau aa auggugagcccacca gugaggauuac ugau gua cacagu ugaugggguu >SELS_cavPor Cavia porcellus (guinea_pig) COVE score 40.42 aaggacaguc ucuguga cgagcugcguuga augau gucuuccuuau aa auggugagcccacca gugaggauuac ugau gcaga caguuga ugggguuuau >SELS_canFam Canis familiaris (dog) COVE score 39.26 uaggacaguc ucuguga cagguugcguuga augau gucuuccuugu aa acggugagcccacca gcgaggauuac ugau gcaga caguuga ugggguuguu >SELS_felCat Felis catus (cat) COVE score 41.01 uaggacaguc ucuguga cagguugcguuga augau gucuuccuugu aa auggugagcccacca gcgaggauuac ugau gcaga caguuga ugggguuguu >SELS_equCab Equus caballus (horse) COVE score 41.01 uaggacaguc ucuguga cagguugcguuga augau gucuuccuugu aa auggugagcccacca gcgaggauuac ugau gcaga caguuga uggguuguuu >SELS_bosTau Bos taurus (cow) COVE score 41.15 uaggacaguc ucuguga cagcuugcguuga augau gucuuccuugu aa auggugagcccacca gcaaggauuac ugau gcaga caguuga ugggguuguu >SELS_eriEur Erinaceus europaeus (hedgehog) COVE score 36.87 cucucuguga uggguug cguugg augau gucuuccuuguc aa uggugaacccacca gcgaggaucac ugau gcag acaguug augggguugu >SELS_sorAra Sorex araneus (shrew) COVE score 35.70 ugccgggccc gucucug ugauaggcgacga augac gucguccucgg aa auggugugccacca gcgaggaccac cgau gaagacacu ccugggc gccccccccc >SELS_dasNov Dasypus novemcinctus (armadillo) COVE score 35.89 uagaacaguc ucuguga cagguugcguuga augau gucuuccuuau aa aauggugaacccacca gugaggauuac ugau aauaga caguuga ugggguuugu >SELS_echTel Echinops telfairi (tenrec) COVE score 38.43 uagaacaguc ucuguga cagguugcguuga augaa gucuuccuuau aa auggugaacucacca gugaggauuac ugau aaaga caguuga ugagguuugu >SELS_monDom Monodelphis domestica (opossum) COVE score 40.64 uagaugaguc ucuguga caggcugcgcaga augau gucuuccuuau aa auggugagcucacca gugaggauuac ugau gauagauag uuguugg uguuuuuuuc >SELS_ornAna Ornithorhynchus anatinus (platypus) COVE score 33.47 ucaauaaguc ucuguga caggcggcauuga augau gucuuccuugu aa auggugaaacccacca gugaggauuac ugau aua gacagg agaugguauu >SELS_anoCar Anolis carolinensis (lizard) COVE score 25.44 gaaacucccu guaacaa gcagcaacaa augac gugguccuuau aa augguggacacauca cugaggaccuc cgaa gauaagacag cugauug gggacaguuc >SELS_galGal Gallus gallus (chicken) COVE score 35.60 cgggagaguc ucuguga caagcugcucugu augau guuuuccuuau aa augguaaacaaacca augaggauuac ugau gcua gacagca gauggggguu
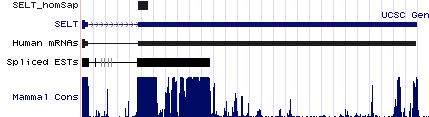
SELT: 21 SECIS insertion sequences
>SELT_homSap Homo sapiens (human) COVE score: 39.15 gaucauugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_panTro Pan troglodytes (chimp) COVE score: 39.15 gaucauuuca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_macMul Macaca mulatta (rhesus) COVE score: 39.15 gaucauugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_otoGar Otolemur garnettii (bushbaby) COVE score: 39.15 gauuuuugca agagcag cgugacugacagu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_musMus Mus musculus (mouse) COVE score: 39.15 ggauuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau guuucuugg caggcuc guuguaccuc >SELT_ratNor Rattus norvegicus (rat) COVE score: 39.15 ggauuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau guuucuugg caggcuc guuguaccuc >SELT_cavPor Cavia porcellus (guinea_pig) COVE score: 39.15 aauuuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uuuuug gcaggaucgu >SELT_oryCun Oryctolagus cuniculus (rabbit) COVE score: 39.15 ggauuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uuuuug gcaggcucgu >SELT_canFam Canis familiaris (dog) COVE score: 39.15 auuuuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_felCat Felis catus (cat) COVE score: 39.15 auuuuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_equCab Equus caballus (horse) COVE score: 38.49 gacuuuugca agagcag cguggcugacagu augaa ggccuguacug aa gacagcaagcugu uaguacagacc cgau gcu uucuug gcaggcucgu >SELT_bosTau Bos taurus (cow) COVE score: 39.15 gauuuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_eriEur Erinaceus europaeus (hedgehog) COVE score: 39.15 gauuuuugca agagcag cgugacugacauu augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau gcu uucuug gcaggcucgu >SELT_sorAra Sorex araneus (shrew) COVE score: 35.80 ggauucucca agagcag cgcggcugcccuc augaa ggccugcacug aa gacagcagcugu uggugcaggcu ggau cuuccugg caggcuc guuguaccuc >SELT_loxAfr Loxodonta africana (elephant) COVE score: 36.15 gauuuuuuca agagcag cgugacugacauu augaa ggccuguacug aa gacagcgagcugu uaguacagacc agau gcu uuuuug gcaggcgcgu >SELT_monDom Monodelphis domestica (opossum) COVE score: 36.95 auuuuugcga gagcagc gugguuggcacca augaa ggccuguacug aa gacagcaagcugu uaguacagacc agau acuu acuuugc agccucguug >SELT_ornAna Ornithorhynchus anatinus (platypus) COVE score: 36.17 gaucuuucua agggcaa cgugauugaaacu augaa ggccuguacug aa ggcagcaaacugu uaguacagacc agau guuccu uugcagu cucguuguac >SELT_anoCar Anolis carolinensis (lizard) COVE score: 34.03 acaguuuuga aaggcaa caugaacgaaaua augaa ggucuguacug aa gacagcaugcugu uggugcagacu ggau acuucuc cucgccuu cauguuguug >SELT_galGal Gallus gallus (chicken) COVE score: 38.05 gugauuucca aggccgc gugucucggacca augaa ggccuguacug aa gacagcuugcugu ugguacagacu ggau gcuucucuu gcaguc acguugugcc >SELT_danRer Danio rerio (zebrafish) COVE score: 34.86 ccggucagcg ggacugu ggacuguggcaua augaa ggccugcgcugc aa acagcacacug uuggcacaggcu ggau gcu cagccac cacacacacu >SELT_tetNig Tetraodon nigroviridis (pufferfish) COVE score: 33.32 ggugggucuc ccugugg gcugugagguu augaa ggucugugcug aa ggcaggacgacugc uagcucagacu ggau gucccca ucacug uucaccgugc
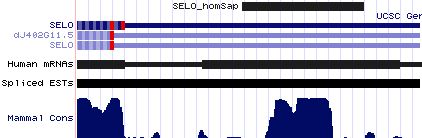
SELO: 5 SECIS insertion sequences
>SELO_homSap Homo sapiens (human) COVE score: 25.77 cugcccuggc ccaugca cacccgucuuucc augau ggcagagacau cc agucaggaccuga cccgucucuguc ugag gccggcuc agcagug cagccugguc >SELO_panTro Pan troglodytes (chimp) COVE score: 25.77 cugcccuggc ccaugca cacccgucuuucc augau ggcagagacau cc agucaggaccuga cccgucucuguc ugag gccggcuc agcagug cagcuugguc >SELO_macMul Macaca mulatta (rhesus) COVE score: 25.77 cugccugggc ccaugca caccugucuuucc augau ggcagagacau cc agucaggaccuga cccgucucuguc ugag gccagcuc agcagug cagccugguc >SELO_monDom Monodelphis domestica (opossum) COVE score: 24.43 gucuggggcc uccccga uggcugcuucucc augaa ggcagagaugu cc agucggcuccga cccgucucuguc cgac gcugac ucggca gcccgagcuc >SELO_takRub Takifugu rubripes (fugu) COVE score: 20.04 ccugcuugcc ccgaguu cuaccugcgaaua augac gacucagacaa ca agucugccagugac cugucugacuc ugaa cugugu ggugucg ccucagcaaa
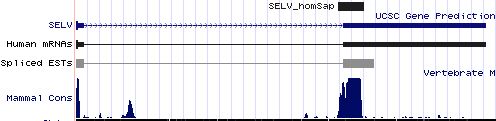
SELV: 10 SECIS insertion sequences
>SELV_homSap Homo sapiens (human) COVE score: 34.00 uuucucuccc aucuuag gagucucagcugg augau gagaagggcug aa auguugccaagu cagguccuuuu cugau ggug gcugggg cuggggugag >SELV_panTro Pan troglodytes (chimp) COVE score: 34.00 uuucucuccc aucuuag gagucucagcggg augau gagaagggcug aa auguugccaagu cagguccuuuu cugau ggug gcugggg cuggggugag >SELV_otoGar Otolemur garnettii (bushbaby) COVE score: 27.28 cuucucuccc aucucag gagccucgguagg augac gagaagggcug aa auauugccaugu cagguccuuuu cugau ggug gcuggga cugggguacg >SELV_musMus Mus musculus (mouse) COVE score: 33.37 cucucucccc aucucag gagccucagcagg augau gagaagggcug aa augcugccaaac cagguccuuuu cugau ggug gcugggg cuuggguggg >SELV_ratNor Rattus norvegicus (rat) COVE score: 33.37 uucucucccu gucucag gaaccucagcggg augau gagaagggcug aa augcugccaaac cagguccuuuu cugau ggug gcuggggc uugguuaggg >SELV_cavPor Cavia porcellus (guinea_pig) COVE score: 34.34 uuuucucccc auguuag gagcugcagcagg augau gagaagggcug aa auguuguugagu cagguccuuuu cugau ggugg cugggcu gaguggagcu >SELV_equCab Equus caballus (horse) COVE score: 32.67 uucucucucc aucucag gagccucggugag augac gagaagggcug aa auguugccaagu cagguccuuuu cugau ggcg gcugggg cugagguggg >SELV_sorAra Sorex araneus (shrew) COVE score: 26.16 cuucucuccc caucuca ggaaccucagaag augaa gggaagggcug aa auguugg cagguccuuuu cugac agu ggcugug gcuaugaugg >SELV_loxAfr Loxodonta africana (elephant) COVE score: 34.00 uucucucccc aucucag gagccucagcaag augau gagaagggcug aa auguugccaagu cagguccuuuu cugau ggug gcugggg cugggguggg >SELV_echTel Echinops telfairi (tenrec) COVE score: 26.31 uucucucucc accccag gagccgcagcaag augac gagaagggcug aa augucgu caggcccuuuu cugag ggug gcugggg cuaggcucgg
SELH: 18 SECIS insertion sequences
>SELH_homSap Homo sapiens (human) COVE score: 23.94 UUUGUGUCCC UGGUGAU GUUGGAACAUUA AUGAU GGAACAUGGCC AA ACUUC AGUCAUGAUCC UGAA GCC AUGGUU UCUUCCCUGC >SELH_panTro Pan troglodytes (chimp) COVE score: 23.94 uuuguguccc uggugau guuggaacauua augau ggaacauggcc aa acuuc agucaugaucc ugaa gcc augguu ucuucccugc >SELH_macMul Macaca mulatta (rhesus) COVE score: 26.97 cuuugugucc cugguga ugcuggaacauua augau ggaacauggcc aa acuuc agucaugugcc ugaa gccaugg uuucua ccccgccaga >SELH_otoGar Otolemur garnettii (bushbaby) COVE score: 22.59 cuuucugugc uggugau guuggagcaugu augac gggacauggcc aa acuua agucaugugcc ugaa gcu acaguuu cuuccccucc >SELH_tupBel Tupaia belangeri (tree_shrew) COVE score: 26.55 gcuuuuccug gugaugu ugaagggacauuu augau gggacauggcc aa acuuc agucaugugcc ugaa gcc aagguuu cuuccccucc >SELH_musMus Mus musculus (mouse) COVE score: 24.55 cuuucagucc cuggaga uguugaagcauuu augau ggugcauggcc aa acuua agcuaugcacc ugaa gccauag uuucuu ccucaccaga >SELH_ratNor Rattus norvegicus (rat) COVE score: 23.97 cuuucagucc cuggaga uguugaagcauuu augau ggugcauggcc aa acuua agcuauguacc ugaa gccauag uuucuu ccucaccaga >SELH_cavPor Cavia porcellus (guinea_pig) COVE score: 19.25 uuucuguucc ugguccu guuggggcgucu gugau gggacauggc ca aacuaaag ccauggccc ugaa gcc augguc ucuagcaucc >SELH_canFam Canis familiaris (dog) COVE score: 23.64 agcauauccc uggugau guuggagcauuu augac ggaacauggcc aa acuuc agucauguacc ugaa gcc augguu ucuucccucc >SELH_felCat Felis catus (cat) COVE score: 24.80 cuucucuccc uggugac guuuggagcauuu augac gggacauggcc aa auuuc agucaugugcc ugaa gcccug guuucu uccccuccag >SELH_equCab Equus caballus (horse) COVE score: 25.73 ccuucguccc ugguguu guuggagcauuu augac gguacauggcc aa acauc agucaugugcc ugaa gcu gugguuu cuuccccucc >SELH_bosTau Bos taurus (cow) COVE score: 28.39 ccucuguccc uggugau guuggagcauuu augac gggacauggcc aa acuuc agucauguccc ugaa gcu gugguu uccuccccuc >SELH_sorAra Sorex araneus (shrew) COVE score: 24.10 cuuucuguuc uggugau guuggagcauug augau ggaacaugacc aa acguc agucaugugcc ugaa gccaa guuucu uucucuccag >SELH_dasNov Dasypus novemcinctus (armadillo) COVE score: 25.97 cuucuauccu uggugau guuggagcauuu augau gggacauggcc aa acuuc agucauguacc ugaa gcca guuucu cccccuccag >SELH_loxAfr Loxodonta africana (elephant) COVE score: 25.83 cuucuguccc uggugau guuggagcauuu augac ggaacauggcc aa accuc agucaugugcc ugaa gcc acgguuu cuuccccucc >SELH_echTel Echinops telfairi (tenrec) COVE score: 26.97 agucccgucc cugguga uguuggagcauuu augau ggaacauggcc aa acuuc agucaugugcc ugaa gcccguc uccuca cuuccagaaa >SELH_monDom Monodelphis domestica (opossum) COVE score: 26.74 ugccuguccc cagugag uuuggagcaucc augau ggaaccugacc aa aucuccca gucacguccc ugaa gcuuggg cuccuu cuccugggaa >SELH_anoCar Anolis carolinensis (lizard) COVE score: 27.10 auacauucuc ugugagu uggagcauuc augau gggauaugauc aa guggagcaaaucc agucacaucuc ugaa gcu guugcc cuccuccaca
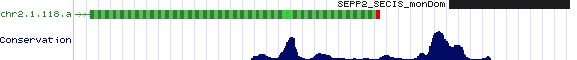
SEPP2: 5 SECIS sequences
>SEPP2_SECIS_monDom Monodelphis domestica (opossum) chr2:9455773-9456962 COVE: 30 55 bp after stop codon ACGTGCCGCTGCCCCTCCCTCCCTCCAAGAATGACGCCCACAGTGAAACCCAGAGAACTGGTCCCTGTGGGCTGATGCCCCAGAGGGGAGGAGAGGC >SEPP2_SECIS_macEug Macropus eugenii (wallaby) SECIS COVE score: 26.38 trace ti|975557399 contains terminal exon CCTCATTCCTCTTCTCCATGGCATCAGCCCACCATGGCCGGTTCCCCGGGTTTCACTGTGGGCATCATTAATTGAGGGAGGGAAGGGCAATGGGCAAG >SEPP2_SECIS_ornAna Ornithorhynchus anatinus (platypus)Ultra131:321101-348720 COVE: 34.28 no match to chicken TCCCTCGACTCCCGCTCCCGCCTCGCACTCATGACGTCCACGGTGTCAACCGGCCCGCCGGGCACCGTGGACTGACGCCGGTCGAGGCGGAGGGGT >SEPP2_SECIS_ornAna Ornithorhynchus anatinus (platypus) Ultra131:321101-348720 COVE: 22.82 no match to chicken GGAATCAGGAACCCAGTAACATGAGGTCATCTTCGGAAGCCTGTGCCTAGAGGACCAAGATAATGGAAAAAGTGACGGACAAGGGTGTGTAGCTGG >SEPP2_SECIS_tetNig Tetraodon nigroviridis (pufferfish) COVE: 17 GCTGGACCCAGGCTGCTGGTGGTCCCGTTGATGACGTCTGCGCTGGTAAACCTGCCTGCAGGAGCCTGTGGACCGACGTGTGTGGACCCACCGGCAG
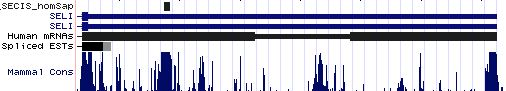
SELI: 17 SECIS sequences
>SELI_SECIS_homSap Homo sapiens (human) COVE score: 21.61 UUUCACUGAA UGAAGUU UGUGCUUGA AUGAA GAGUGUAUCUUA AA CCCCCUUUUUUUGGA CAGGCUGCACUU GGAU AAAAUA GGCACCA CUGUGUUGAU >SELI_SECIS_panTro Pan troglodytes (chimp) COVE score: 21.61 uuucacugaa ugaaguu ugugcuuga augaa gaguguaucuua aa cccccuuuuuuugga caggcugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_macMul Macaca mulatta (rhesus) COVE score: 21.25 uuucacugaa ugaaguu ugugcuuga augaa gaguguaucuua aa ccuccuuuuuuugga caggcugcacuu ggau aacaua ggcacca cuguguugau >SELI_SECIS_otoGar Otolemur garnettii (bushbaby) COVE score: 21.25 uuucacuaag ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuuuuuuggac agguugcacuu ggau aaaaua ggcacca uuguguugau >SELI_SECIS_musMus Mus musculus (mouse) COVE score: 24.62 uuccacugaa ugaaguu ugugcuuaa augaa gagugugucuu aa acccuuuuuuuuggac agguugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_ratNor Rattus norvegicus (rat) COVE score: 24.62 uuccacugaa ugaaguu ugugcuuaa augaa gagugugucuu aa acccuuuuuuuuggac agguugcacuu ggau aacaua ggcaccg cuguguugau >SELI_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 17.42 uuccagugaa ugaaguu ugugcuuga augaa gaguguaucuua aa cccuuuuuuuuuugga cagguugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_oryCun Oryctolagus cuniculus (rabbit) COVE score: 19.47 ucuacugaau gaaguuu gugcuuga augaa gaguguaucuua aa cccuuuuuuuuugga cagguugcacuu ggau aaaau aggcac cacuguugau >SELI_SECIS_sorAra Sorex araneus (shrew) COVE score: COVE score: 8.13 loose uuccacugaa ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuauuuuguuuuuuugga cggauuacacuu ggau auaaua ggcacca uucuguugau >SELI_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 14.43 loose cacggaauga aguaugu gcuuga augaa gaguguaucuu aa acccuuuuuuuuuuuugga cagguugcacuu ggau aua auaggc accacucugu >SELI_SECIS_canFam Canis familiaris (dog) COVE score: 14.04 uuccacugaa ugaaguu ugugcucga augaa gaguguaucuua aa cccuuuuuuuuuugga uggauugcacuu ggau aaaaua ggcacca cuguguugau >SELI_SECIS_equCab Equus caballus (horse) COVE score: 19.57 uuucacugag ugaaguu ugugcucga augaa gaguguauucuu aa acccuuguuuuugga uggauugcacuu ggau aaaaua agcacca cuguguugau >SELI_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 17.76 loose cuacugaauu aaguuug ugcuuga augaa gaguguaucuu aa acccuuuguuuuuuugga cagguugcacuu ggau aaaa uaggca ccacuauguu >SELI_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 25.81 uuccacugaa ugaaguu ugugcuuga augaa gaguguaucuu aa acccuuuuuuuuggac agguugcacuu ggau aaagua ggcacca cuguguugau >SELI_SECIS_monDom Monodelphis domestica (opossum) COVE score: 26.46 uucuacugaa ugcaauu ugugcuuga augaa gagugugucuu aa auccuuuauauggac aggcugcacuu ggag aga auaagca caaccauguu >SELI_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 19.25 ucccagugaa ugaagcu ugugcuuga augaa gagugcaucuua aa cccauuuuuuuugga aaagcugcacuu ggag agaaag ggcacg acuguguuua >SELI_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 22.02 uccccuuugu guguguc cuuugugcgugu augaa gagugcggccuc aa cccaggcgucuugga agggccgcaccc ggaa gaa acggagc acagcaaaga >SELI_SECIS_galGal Gallus gallus (chicken) COVE score: 24.62 uuuuauugag uuuauuu gugcuuaa augaa gagugcgcuuc aa acccagaccaggag agggcgcacuu ggag ugagc gagucaa accuugcucc
SELK: 19 SECIS sequences
>SELK_SECIS_homSap Homo sapiens (human) COVE score: 34.53 acaaggacug cucugug uccucacagauga augag gucaugcuggg aa uucccucugcaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_panTro Pan troglodytes (chimp) COVE score: 34.53 acaaggacug cucugug uccucacagauga augag gucaugcuggg aa uucccucugcaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_macMul Macaca mulatta (rhesus) COVE score: 36.14 acaaggacug cucugug uccucacagauga augag gucaugcuagg aa uucccucuacaggga acuggccugac ugac augcaguuc cauaaa ugcagauguu >SELK_SECIS_otoGar Otolemur garnettii (bushbaby) COVE score: 31.99 caaggauugc ucugugu cuucacagauga augag gucaggcuggg aa uucucucuucaggga acuggccugac ugac augcagu ucuauaa acgcacuuuu >SELK_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 36.65 acaaggauug cucugug uccccacagauga augag guuaugccggg aa uucccuccacaggga ucuggccugac ugau acgcaguuc uauaaa ugcacauguu >SELK_SECIS_musMus Mus musculus (mouse) COVE score: 31.73 acaaggauug cucugug uccccacagauga augag gucaugcuggg aa uucccucugcagga ucuagccugac ugau acgcaguuc uauaaa uguacauguu >SELK_SECIS_ratNor Rattus norvegicus (rat) COVE score: 33.02 acaaggacug cucugug uccucacagaaga augag gucaugcuggg aa cucccucugcagga ucuggccugac ugau gugcaguuc uauaaa uguacaugug >SELK_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 33.72 cucuguguuc ucacaga uaa augag gucacgccagg aa uucucucagcaggga ucuggcuugac ugau acgcagu ucucua aaugcauaug >SELK_SECIS_oryCun Oryctolagus cuniculus (rabbit) COVE score: 38.80 accaggauug cucugug uccccacagauga augau gucaggcuggg aa uucccuccacaggga ucuggccugau ugag augcaguuc uauaaa ugcguauguu >SELK_SECIS_sorAra Sorex araneus (shrew) COVE score: 40.91 auaaggacug uucugug uccacacagauga augag gucaugcuggg aa uucccucuacgggga ucuggcaugac ugau augcag uucgaua aaugcacaug >SELK_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 37.30 acaaggauug cucugug uucucacagguga augag guuaugcuggg aa uucccuccaugggga ucuggcaugac ugau augcaguuc uauaaa ugcacauguu >SELK_SECIS_canFam Canis familiaris (dog) COVE score: 36.30 acaaggauug cucugug ccuucacagacgg augag guugugcuagg aa uucccuccccaggga ucuggcaugac ugac augcaguuc uauaaa ugcacauguu >SELK_SECIS_felCat Felis catus (cat) COVE score: 37.35 acgagaauug cucugug uccucacagacag augag gucgugcuggg aa uucccuccccaggga ucuggcaugac ugac augcaguuc uauaaa ugcacauguu >SELK_SECIS_bosTau Bos taurus (cow) COVE score: 39.16 cucugugucc ucacaga cga augag gucaugcuggg aa uucccuccgcaggga ucuggcaugac ugac augcagu ucuaua aaugcacguu >SELK_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 29.00 acaaggauug cucugug uccucacagauga augag gucauguuuggg aa uucccucugcaggg aucuggcaugac ugac uugcaguuc cauaaa ugcacauguu >SELK_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 34.85 acaaagacug cucugug uccccacagacgg augag gcugugcuggg aa uucccucugcaggga ucuggcauggc ugac augcaguuc cauaaa ugcacauguu >SELK_SECIS_monDom Monodelphis domestica (opossum) COVE score: 21.75 aagaauugcu cugucua cacagauua augau guugugcuggg aa cucccaucuuacagga uccaguguaac ugau ugcaa uuguaua aaugcacaug >SELK_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 33.68 aauaauugug cugugaa caagcagauua augau guuuugcuggg aa uuccuucaggga uccaguauaac ugau aaagc aauuaua uaaaggcaca >SELK_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 33.87 aauucccugc ucugcca auuggcgggacc augau guuguccuggg aa uuccuuauucuggga uccagggcaac ugaa aagcaguuc uguuaa auuaaaugca >SELK_SECIS_galGal Gallus gallus (chicken) COVE score: 30.90 ugagaacugu ucugcaa uauaagcagauga augaa guuguacuggg aa cuccuucaagga uccaguguaac ugaa gugcagug uuauuaa auacauguuu
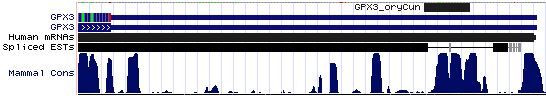
GPX3: 15 SECIS sequences
>GPX3_SECIS_homSap Homo sapiens (human) COVE score: 27.80 ACGGACCCCA UGGCAGG GGUGGCGUCUUC AUGAG GGAGGGGCCCA AA GCCCUUGUGGGC GGACCUCCCC UGAG CCUG UCUGAG GGGCCAGCCC >GPX3_SECIS_panTro Pan troglodytes (chimp) COVE score: 27.80 cacguacccc augucgg ggguggugucuuc augag ggaggggccca aa gcccuugugggc ggaccucccc ugag ccugu cugagg ggccagcccu >GPX3_SECIS_macMul Macaca mulatta (rhesus) COVE score: 27.80 cacguacccc augucag ggguggcgucuuc augag ggaggggccca aa gcccuugugggc ggaccucccc ugag ccugu cugagg ggccagcucu >GPX3_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 29.38 ccacaucccu gugucag ggguggcaucucc augag ggaggggcccg aa gcccuuguggg cggaccucccc ugag ccugu cugagg ggccggcccu >GPX3_SECIS_musMus Mus musculus (mouse) COVE score: 33.37 cguguacccc aggucag ggguggugucucu augaa ggaggggcccg aa gcccuuguggg cgggccucccc ugag cccgu cuguggu gccagcccuu >GPX3_SECIS_ratNor Rattus norvegicus (rat) COVE score: 33.37 caugugcucc aagucag ggguggugucucc augaa ggaggggcccg aa gcccuuguggg cgggccucccc ugag cccgu cuguggu gccagcccuu >GPX3_SECIS_canFam Canis familiaris (dog) COVE score: 29.96 cauguucccc gugucag gaauggcaucucc augaa ggaggggccc ga agcccucaugggc ggaccucccc ugag ccugu cugaag ggccggcccu >GPX3_SECIS_felCat Felis catus (cat) COVE score: 30.08 gccacguccc gugucag ggauggcgucucc augaa ggaggggccc ga agcccuugugggc ggaccucccc ugag ccugu cugaag ggccagcccu >GPX3_SECIS_equCab Equus caballus (horse) COVE score: 26.20 uacauucccc augucag ggguggcaucucc augau ggaggggcccg aa gcccugguggg cggaccucccc agag ccugu cugaag ggccagcccu >GPX3_SECIS_bosTau Bos taurus (cow) COVE score: 30.87 acguuccccg ugucagg gggcggcaucgcc augaa ggaggggcccg aa gcccgcguggg cgggccucccu ugag ccu gucugag gggccagccu >GPX3_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 16.32 cuacgucccc gugucaa gagcggcaucucc augau gguggggcccg aa gccccugugg cggaccucccc agag ccugucc caugggc cagcccuuag >GPX3_SECIS_echTel Echinops telfairi (tenrec) COVE score: 12.78 ggcugugucc cgugcag agggagcaucucc augag ggugaggcccg aa gcccccgugg cggaccucgac ccgag ucug cucugg gccugccuuc >GPX3_SECIS_monDom Monodelphis domestica (opossum) COVE score: 22.94 ugaacauaag ggauggc aucucu augau ggugggaucca aa gccucuucaggg cggguuccauc agag ccugcaaaa gguguc aggacccuua >GPX3_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 31.33 ugucccagaa uggggag guggcaucauc augac agcggggucug aa agccccuccugga uggaccccgcc cgaa ccug cucggcg guggcaugac >GPX3_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 28.20 GAGAUUUUGG GCCAAGG AAGUUGCAUCACU AUGAG GGUUAGGUCUG AA AGCUCCCAAAAAGAG CGGACCUAGCC UGAG GCUGCAAA GCUCUGGU GUAGCCCUUU >GPX3_SECIS_galGal Gallus gallus (chicken) COVE score: 30.65 GCCCUGGGGA GCAGAGG AUGACAUCUCC AUGAA GGCCUGGCCUG AA AGCCCCCACCAUGGGG UGGGCUCGGCC CGAU CCCG CCCAGGC GCGGUGCAGC
GPX5: 0 SECIS sequences (cysteine all species)
GPX6: 10 SECIS sequences (most species have selenocysteine
>GPX6_SECISa_homSap Homo sapiens (human) chr6:28,578,901-28,580,158 Cove score: 18.10 cccaccucac augaagg gaagggcaucucc augau gguggauccc aa aaccccucuggguc gcacccugcc agag ccu uccuug gugccugucc >GPX6_SECIS_panTro Pan troglodytes (chimp) COVE score: 23.45 cccaccucac augaagg gaggggcaucucc augac gguggguccc aa aaccucucgggguc ggacccugcc agag ccu uccuug gugccugucc >GPX6_SECIS_macMul Macaca mulatta (rhesus) COVE score: 21.81 accucauuca cagaagg gaggggcaucuuu augau ggugggucuc aa aaccucucuggguc agacccuacc agag ccu uccuug gugccugucc >GPX6_SECIS_cavPor Cavia porcellus (guinea_pig) AAKN02051715 Cove score: 19.91 AGAAGCCUCC CUAGAAG GAAGGGUGUCUCC AUGAU GGUGGGUCCC AA AAGCCCUGGAUC GGACCCUACC AGAA CCU UCCCUGG UGCCUGUCCU >GPX6_SECIS_canFam Canis familiaris (dog) Cove score: 24.33 but not found in 28way CCUGGCCUCA UAUGAAA GGAGGGCAUCUCC AUGAU GGUGGGUUCC CA AGCCCCUGCGGUC GGACCCUACC AGAG CCU UCUUGG GUGCCUGUCC >GPX6_SECIS_equCab Equus caballus (horse) COVE score: 25.00 uucaucucac augagcu gaggggcaucucc augau gguggguccc aa agccccugugggc aggacccaac cagaa cucug ugccugu cccuuagugc >GPX6_SECIS_bosTau Bos taurus (cow) COVE score: 23.33 cucacuucac augagga aaagggcaucucc augau gguggguccc aa agccucucuggguc ggaccccacc agag ccu uccuugg ugccuguccc >GPX6_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 24.33 ccuggccuca uaugaaa ggagggcaucucc augau gguggguucc ca agccccugcgguc ggacccuacc agag ccu uccuugg ugucuguccu >GPX6_SECIS_sorAra Sorex araneus (shrew) frag COVE score: 22.95 uuucauauga aaggagg gggcaucucc augau ggugggucuc aa agccccucuggguc ggacccuacc agag cccugcugg guguccu gucccuuugu >GPX6_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 19.82 cucaccuuac augaagg gagggacauaucc augau gguggguccu aa agcccuucuggguc agacccuacc agaa ccu ucucug gugccugucc >GPX6_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 19.28 cucaccucac augaaug gaggggcauaucc augau gguggguccc aa agccucucuagguc ggacccuauc agaa ucuucccugg ugccugu cccuuagugc >GPX6_SECIS_echTel Echinops telfairi (tenrec) frag COVE score: 31.33 ugugucccag aaugggg agguggcaucauc augac agcggggucu ga aagccccuccuggau ggaccccgcc cgaa ccu ucccug gugccugucc
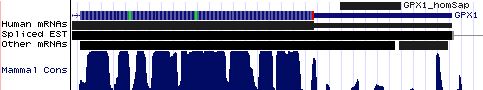
GPX1: 17 SECIS insertion sequences
>GPX1_homSap Homo sapiens (human) COVE score: 20.45 UGCUGUCUCG GGGGGGU UUUCAUCU AUGAG GGUGUUUCCUCU AA ACCUACGA GGGAGGAACACC UGAU CUUACAGAAA AUACCAC CUCGAGAUGG >GPX1_panTro Pan troglodytes (chimp) COVE score: 20.45 gugcugcucu guggggg gguuuucaucu augag gguguuuccucu aa accuacga gggaggaacacc ugau cuuacagaaa auacccc cucgagaugg >GPX1_macMul Macaca mulatta (rhesus) COVE score: 26.45 gcagcgcugc ucucugg gggguuuucaucu augag gguguuuccucu aa accuaca aggaggaacacc ugau aau acagaa aauacccccu >GPX1_otoGar Otolemur garnettii (bushbaby) COVE score: 25.30 auagcgcugc ucucugg ggggguuucauuc augau aguguuaccucu aa acuugcau gggggaacacc ugau gcc ccagaaa auccccugag >GPX1_tupBel Tupaia belangeri (tree_shrew) COVE score: 21.14 cagcgcugcg ucugggg gguuucaucc augac ggugucuccucu aa accccga aggaggaacgcc ugau guccggaaaa cccccca ggugggcgcc >GPX1_musMus Mus musculus (mouse) COVE score: 29.78 ggcugcacuc ugggggg cgguucuucc augau gguguuuccucu aa auuugca cggagaaacacc ugau uuccaggaa aaucccc ucagaugggc >GPX1_ratNor Rattus norvegicus (rat) COVE score: 25.72 ggcugcccuc cgggggg agguuuuucc augac gguguuuccucu aa auuuaca uggagaaacacc ugau uuccagaaa aaucccc ucagaugggc >GPX1_cavPor Cavia porcellus (guinea_pig) COVE score: 15.34 uugcccagau gcucucc ugaggguucuucc augaa gguguuuccucuc aa ccugua uagaggaacauc cgau uccca ggaauu ucccagagag >GPX1_oryCun Oryctolagus cuniculus (rabbit) COVE score: 24.38 guggccugcu gcucucu ggggguuucaucc augag ggcguucccccg aa aacaaa uggaggaacgcc ugau gucc gggaaac ccccaggugg >GPX1_eriEur Erinaceus europaeus (hedgehog) COVE score: 19.31 auagccgcga gccgcug ggcaucucaucc augac ggcgccgccuuc aa accugcgagcag gaaggagcgcc cgau agc cgcgaga gcccccagcg >GPX1_canFam Canis familiaris (dog) COVE score: 22.40 ucacagcgcu gcccucu ggggauuucaucc augau gguguuuccuug aa aucugca uggaggaacgcc ugau uucca ggaaagu ccccugagcu >GPX1_felCat Felis catus (cat) COVE score: 27.75 cacagcacug cccucug gggauuucaucc augau ggcguuuccucg aa auuugca uagaggaacgcc ugau uuc cagaag aauccccuga >GPX1_equCab Equus caballus (horse) COVE score: 27.08 ucacagugcu uuucucu ggggauuucaucc augau ggcguuuccucu aa acaugc augaggaacgcc ugau guu aaggaga aucccccgag >GPX1_bosTau Bos taurus (cow) COVE score: 29.83 agucagcgcu gcucucc agggauuuugccc augaa gguguucccucu aa accuacg uggaggaaugcc ugau gucca ggaaaau ccccugaggu >GPX1_dasNov Dasypus novemcinctus (armadillo) COVE score: 28.92 uguccuccac acggggu uuucaucc augac gguguuuccucu aa accugca gggaggaacacc ugaa guccggcaaa aucccc cgagaugggu >GPX1_loxAfr Loxodonta africana (elephant) COVE score: 25.48 acuaggcggc uuuccgu ggggguuucauua augag gguguuuucucu aa accugaa uggaggaacacc ugau guc ugggaa gauacccccc >GPX1_monDom Monodelphis domestica (opossum) COVE score: 26.38 gugcuaaggg uccguga ggguuuuaucu augau gguguuguuuu aa accauuaagga gaaagaacacu ugau aaugc uuguaa aaucccauga >GPX1_ornAna Ornithorhynchus anatinus (platypus) COVE score: 25.55 ugcuauuugc ccauggg ggagaucucauuu augau ggugcuccuucu aa accuuua cggaagagcacu ugau gga uccugag gaacuccaug
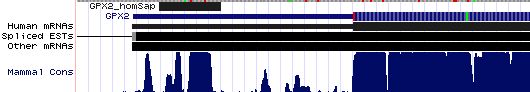
GPX2: 17 SECIS insertion sequences
>GPX2_homSap Homo sapiens (human) COVE score: 29.54 AGACUUGGGU AAGCUCU GGGCCUUCACAGA AUGAU GGCACCUUCCU AA ACCCUCA UGGGUGGUGUC UGAG AGGCGUGA AGGGCCU GGAGCCACUC >GPX2_panTro Pan troglodytes (chimp) COVE score: 29.54 agacuugggu aagcucu gggccuucacaga augau ggcaccuuccu aa acccuca ugggugguguc ugag aggcguga agggccu ggagccacuc >GPX2_macMul Macaca mulatta (rhesus) COVE score: 29.54 agacuugggu aagcucu gggccuucacaga augau ggcaccuuccu aa acccuca ugggugguguc ugag aggcguga agggccu gaagccaccc >GPX2_otoGar Otolemur garnettii (bushbaby) COVE score: 27.86 agacuugggu gggcucu gggccuucacaga augau ggcaccauccu aa acgccuc ugggugguguc ugag aagagcg gaaggcc uggagccacc >GPX2_tupBel Tupaia belangeri (tree_shrew) COVE score: 28.69 agacuuaggu gggcucu gagccuucacaga augau agcaccuuccu aa acccccc cgggagguguc ugag aagugug acaggcc cggagccagc >GPX2_musMus Mus musculus (mouse) COVE score: 33.65 agucuggggu agguucu gggccuucacaga augau ggcaucuuccu aa acccuuc ugggagauguc ugag aaguugug aaggguc cagagccagu >GPX2_ratNor Rattus norvegicus (rat) COVE score: 30.80 cugggguagg ugcuagg ccuucuucacaga augau ggcaucuuccu aa acccuuc ugggggauguc ugag acg uugugaa gggcccagag >GPX2_cavPor Cavia porcellus (guinea_pig) COVE score: 20.08 cuaggacagg uggcucu gggccuucacaga augac ggcaccguccu aa acgcua ugggugguauc ugag aagugug aauggcug gagccagccu >GPX2_sorAra Sorex araneus (shrew) COVE score: 29.00 aagcuggggc aaacucu gggccuucgcaga augau ggcaccucccu aa auccau ggggugguguc ugag gcgugcg agggccu ggaaacagcc >GPX2_eriEur Erinaceus europaeus (hedgehog) COVE score: 25.06 aaagcugggu gguuucu ggaccuucacaga augau agcaccuuccu aa accuaua gggaugguguc ugag aaaugu gaagggc cugaaguaaa >GPX2_canFam Canis familiaris (dog) COVE score: 22.57 aagacuuggg uggcucu gggccuucacaga augau ggcaccuuccu aa auagua ugggcgguguc ugag aagugug aagggcu cagagccagc >GPX2_felCat Felis catus (cat) COVE score: 22.45 aagacuuggg uggaucu gggccuucacaga augau ggcaucuuccu aa acugua ugggcgguguc ugag gaguguga agggcu cggagccagc >GPX2_equCab Equus caballus (horse) COVE score: 28.45 agacuucggu aggcucu gggccuucacaga augau ggcaccuuccu aa accugua uggacgguguc ugaa aagcgug aagggcc ccgagucagc >GPX2_bosTau Bos taurus (cow) COVE score: 26.14 agacaugggu gggcucu gugucuucacaga augau ggcaccuuccu aa aucugua ugggcgguguu ugag aagagu gaaggcc uggagccagc >GPX2_loxAfr Loxodonta africana (elephant) COVE score: 27.08 agacuugggu aggcucu ggaccuucgcaga augau ggcaccuuccu aa acucag ugggugguguc ugag aaaugug aagggccu agggccagcc >GPX2_echTel Echinops telfairi (tenrec) COVE score: 29.95 agugcuggug uggcucu gggccuucacaga augau ggcaccuuccu aa acccuc cggaagguguc ugag aaaugug aagggcc uggggccggc >GPX2_monDom Monodelphis domestica (opossum) COVE score: 26.38 gugcuaaggg uccguga ggguuuuaucu augau gguguuguuuu aa accauuaagga gaaagaacacu ugau aaugc uuguaa aaucccauga >GPX2_ornAna Ornithorhynchus anatinus (platypus) COVE score: 27.55 gggcucgagc agccucc agaccuucacaga augac ggugucuccuu aa acccuaac cgggaggcacc cgag agccggu gaagggc cuggugacug
GPX4: 14 SECIS sequences
>GPX4_SECIS_homSap Homo sapiens (human) chr19:1,057,512-1,057,793 COVE score: 29.19 CCACGCCCUU GGAGCCU UCCACCGGCACUC AUGAC GGCCUGCCUG CA AACCUGCUGGU GGGGCAGACC CGAA AAUCC AGCGUGC ACCCCGCCGG >GPX4_SECIS_panTro Pan troglodytes (chimp) COVE score: 29.19 CCACGCCCUC GGAGCCU UCCACCAGCACCC AUGAc ggccugccug ca aaccugcuggu ggggcagacc cgaa aaucc agcgugc acccugccgg >GPX4_SECIS_macMul Macaca mulatta (rhesus) COVE score: 33.09 CCACGCCCUC GGAGCCU UCCACCGGCACCC AUGAc ggccugcuugc aa accagcuccuggu gaggcagacc cgaa aaucc agcgugc accccgcugg >GPX4_SECIS_tupBel Tupaia belangeri (tree_shrew) COVE score: 31.05 CCGCGCCCCG GAACCUU CCACCCGGCACCU UUGAc ggucugccuau aa accugccacug gugaggcagacc cgag aaccu ggcgugc acccugccgg >GPX4_SECIS_musMus Mus musculus (mouse) COVE score: 35.62 UGACCCCUGG AGCCUUC CACCCCGGCACUC AUGAa ggucugccug aa aaccagccugcuggu ggggcagucc ugag gaccu ggcgugc acccugccgg >GPX4_SECIS_ratNor Rattus norvegicus (rat) COVE score: 34.33 UGACCCCUGG AGCCUUC CACCCCGGCACUC AUGAc ggucugccug aa aaccagcccgcuggu ggggcagucc cgag gaccu ggcgugc accccgccgg >GPX4_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 34.46 UGGCCGCCCC GAGUCUC CUACCCGGGUGCC AUGAc ggccugccug ca aaccagcgug cugguggggc agac ccg aggaugc gugcacugcc >GPX4_SECIS_canFam Canis familiaris (dog) COVE score: 31.12 AUGCCCCUCG GAGCCUU CCACCCGGCACCC AUGAc agucugucuaa aa accagcccgcugg uggggcagacc cgag aaccc ggcgugc acccugccgg >GPX4_SECIS_sorAra Sorex araneus (shrew) COVE score: 36.85 CUGCCCCUCG GAGCCUU CCACCCGGCGCCC AUGAc ggucugccugc aa accagccc gcugguggggc agac ccgagaaccc ggcgugc accuugccgg >GPX4_SECIS_bosTau Bos taurus (cow) COVE score: ? tgttccgagaAGCCTTCCCCCGGtgCctATGACGGTCTGtCTGAAAACCAGCCCCTGGTGGGGCAGaCCtGAGaACCTGGCGTGCACtccttccgg >GPX4_SECIS_echTel Echinops telfairi (tenrec) COVE score: 40.88 UUGAUCCCCA GAGUCUU CCACCCGGCACCA AUGAc ggucugccuuc aa accaggccucugg ugaggcagacc cgau gaccc ggcgugc acucagccgg >GPX4_SECIS_monDom Monodelphis domestica (opossum) COVE score: 40.88 UUGAUCCCCA GAGUCUU CCACCCGGCACCA AUGAC GGUCUGCCUUC AA ACCAGGCCUCUGG UGAGGCAGACC CGAU GACCC GGCGUGC ACCUCAGCCG >GPX4_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 33.89 ACCUCCGCUU UCCCGGG ACGCUCUGCCUCC AUGAc ggccggccuu ca agccaaaaccaguugg uggggccggcc ugaa caa accggca cgggucccgg >GPX4_SECIS_anoCar Anolis carolinensis (lizard) COVE score: 38.77 UGCUCCCCGU GGGCCCC CUCCUCCAGCACC AUGAc ggccugccuug aa gccagcuugcugg ugaggcagacc cgaa gauuc ggcgugc acugcuggag >GPX4_SECIS_galGal Gallus gallus (chicken) COVE score: 38.77 UGCUCCCCGU GGGCCCC CUCCUCCAGCACC AUGAc ggccugccuug aa gccagcuugcugg ugaggcagacc cgaa gauuc ggcgugc acugcuggag
GPX7: 0 SECIS sequences (all species have cysteine)
GPX8: 0 SECIS sequences (all species have cysteine or serine)
SEPW: 13 SECIS sequences
>SEPW_SECIS_homSap Homo sapiens (human) COVE score: 36.01 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_panTro Pan troglodytes (chimp) COVE score: 36.01 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUuGUGG CUGCUGUUGG >SEPW_SECIS_ponPyg Pongo pygmaeus (orang_sumatran)C OVE score: 30.55 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAAUCUUGUGGACACC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_macMul Macaca mulatta (rhesus) COVE score: 33.55 GACCCAGCCC CUCUCAG CAGACGCUUC AUGAU AGGAAGGACUG AA AAGUCUUGUGGACgCC UGGUCUUUCCC UGAU GUU CUCGUGG CUGCUGUUGG >SEPW_SECIS_musMus Mus musculus (mouse) COVE score: 34.04 UUGGCCCAGC CCCUCGU GGCAGACGCUUC AUGAU GGGAAGAACUG AA AUGUCUCGUGGACGC CUGGUCUUUCC CUGAU GUCCCU GCGACUG CCACGUAGGG >SEPW_SECIS_ ratNor Rattus norvegicus (rat) COVE score: 26.55 UGGCCGGCCU UUCUUGG CAGCCGCUUC AUGAC AGGAAGGACUG AA AUGUCUCAAAGACCUG UGGUCUUUCUU CGAU GUU CCUGCGG CCACCAAGUC >SEPW_SECIS_oryCun Oryctolagus cuniculus (rabbit) no SECIS element found AGTAACCTTGACCCAGCCCCTTTCATGCCTCAGCCTCGTCTCCATAGGCTAAGACTGGAGAAATGAGTCCCCTGAAGAACTGAAACTGGGGGTAGAGGGTTGGTGTTTTAAGATGTGGATGAGCTGGTCTTTAC >SEPW_SECIS_canFam Canis familiaris (dog) COVE score: 34.04 UUGGCCCAGC CCCUCGU GGCAGACGCUUC AUGAU GGGAAGAACU GA AAUGUCUCGUGGACGCC UGGUCUUUCC CUGAU GUCCCU GCGACUG CCACGUAGGG >SEPW_SECIS_susScr Sus scrofa (pig) no SECIS element found AGTAACCTTGACCCAGCCCCTTTCATGCCTCAGCCTCGTCTCCATAGGCTAAGACTGGAGAAGTCTTGTGGACGCCTGGTCTTTCCCTGATGTTCTCGTGGCTGCTGTTGGGGGCAGAGATGGATGAGCTGGTCTTTAC >SEPW_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 33.54 GGCCCAGCCC CUCUCAG CAGAUGCUUC AUGAC AGGAAGGACUG AA AUGUCUUGUGGACGCC UGGUCUUUCCC UGAU GUU CUUGUGG CUGCUGGUUG >SEPW_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: 28.71 CCCAGCUGCC CUUGGCA GACGCUUC AUGAG GGGAAGGACCU AA AUGCGUCGUGGAUGCC UGGUCUUUCCC UGAU GCUCCUUCAC CUGCCAG AUGGGGCAGA >SEPW_SECIS_loxAfr Loxodonta africana (elephant) no SECIS element found GGGACCTTGGCCCAGCCCCTTTCAGCAGACACTTCATGACAGGAGGACTGAAATGTCTCCCAGACGCCTGGCTCTTTCCCTGAATCTGTCGGCTGCAGGACAGGGCAGCGGTTGACTCTCTCGTTTTTTGCAT >SEPW_SECIS_echTel Echinops telfairi (tenrec) no SECIS element found AGGCCAGAGACCTTGGCCCAGTCCCTCCATGACAGGCAGAACTGAAATGTCCTCTGGACAAGTGGTCTTTTCCAGAAACCCCAGGGCTGCTGGGCCGGAGCCGAGGCTGACAACCCTGGTCTTTGCCT
DIO1: 29 SECIS sequences
>DIO1_SECIS_homSap Homo sapiens (human) COVE score: 29 ttttaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_ponPyg Pongo pygmaeus (orang_sumatran) COVE score: 29 ttttaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_macMul Macaca mulatta (rhesus) COVE score: 29 tttcaactctgtgtctttacatatttgtttatgatggccacagcctaaagtacacacggctgtgacttgattcaaaagaaaatgttataag >DIO1_SECIS_cavPor Cavia porcellus (guinea_pig) COVE score: 24 tgttaactctgcttcttttcatatttgttcatgacggtcacagtctaaagtacacacagctgtgacctgatttgaaagaaaatgttttaag >DIO1_SECIS_canFam Canis familiaris (dog) COVE score: - ttttaactctgcttcttttcatgtttgtctatgacggccacagcctaaagcacacacagctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_felCat Felis catus (cat) COVE score: - ttttaactctgcttcttttcacgtttgtctatgacggccacagtctaaagtgcacacagctgtgacttgacttgaacgaaaatgttttaag >DIO1_SECIS_bosTau Bos taurus (cow) COVE score: 28 ttttaactctgcctcttttcatatttgttcatgacggccacagcctaaagtacacacggctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_sorAra Sorex araneus (shrew) COVE score: 24 cggaaactcagcttctcttcatatttgtttatgacagccccagctgaaagtacacacagctgtggcttgattggaaagaaaatgttttaag >DIO1_SECIS_eriEur Erinaceus europaeus (hedgehog) COVE score: 26 tttaactctgctttcttctcatatttgcttatgatggtcacagcttaaagtatacacagctgtgacttgattggaaagaaaatattttaag >DIO1_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 29 ttttaactctgcttcttttcatatttgttcatgatggccacagcctaaagtacacacggctgtgacttgatttgaaagaaaatgttttaag >DIO1_SECIS_dasNov Dasypus novemcinctus (armadillo) COVE score: - ttttaactctgcttcttttcatatttgtttatgatggccacagtttaaagtacatacagctgtgacttgatatgaaaaagaaatattttaag >DIO1_SECIS_loxAfr Loxodonta africana (elephant) COVE score: - ttttaactctgcttcttttttcatgtatttatgatgggccacagcctaaagtgcacaacagctgtgacttgatttgaaaaacatctttaag >DIO1_SECIS_monDom Monodelphis domestica (opossum) COVE score: - tttccatcctgcttctacaaatatttatttatgacaatcacagcctaaagctcagggcagctgggattcgacgggagaaaaagtttgtaag >DIO1_SECIS_ornAna Ornithorhynchus anatinus (platypus) COVE score: 24 ccccggatccggttccgtgaatattggtttatgagggtcacagtgtaaagcgcatgcagctgtgacttgatctgagaaaatatttctgcggc >DIO2_SECIS_homSap Homo sapiens (human) 5,630 bp utr COVE score: 30 cagagatgtgcagagttgaccagtgtgcggatgataactactgacgaaagagtcatcgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_macMul Macaca mulatta (rhesus) COVE score: 30 cagagatgtgcagagttgaccagtgtgcggatgataactactgacgaaagagtcatcgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_musMus Mus musculus (mouse) COVE score: 29 cggagatgttcagagctcactggtgtgcgaatgataactactgacgaaagagctgtctgctcagtctgtggttggatgtagtcacacgagtctgcctttctgca >DIO2_SECIS_ratNor Rattus norvegicus (rat) COVE score: 28 ccgagatgttcggagctcactggtgtgcgaatgataactactgacgaaagagtcatctgctcagtctgtggttggatgtagtcacacgagtctgcctctccatc >DIO2_SECIS_canFam Canis familiaris (dog) COVE score: 27 ctgggatgtgcagaggtgaccagtgtgcgaatgataactactgatgaaagagtcactgactcagttagtggttggatacagtcacattagttttcctct >DIO2_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 30 ctgggatgtgcagaggtgaccagtgtgcaaatgataactactgatgaaagagtcattgactcagttagtggttggatgtagtcacattagtttgcctctc >DIO2_SECIS_dasNov Dasypus novemcinctus (armadillo) iodothyronine deiodinase type II ctgggaagttcagaggctaccagtgtgccaatgataactactgacgaaagaggcatcgactcagttagtggttggatgtagccacattagtttgcctctc >DIO3_SECIS_homSap Homo sapiens (human) COVE score: 31 ttgggtgcacaggagccccactgctgatgacgaactatctctaactggtcttgaccacgagctagttctgaattgcaggggcctcaaagcagca >DIO3_SECIS_macMul Macaca mulatta (rhesus) COVE score: 30 ttgggtgcacaggagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctagttctgaattgcaggggcctcaaaacagca >DIO3_SECIS_musMus Mus musculus (mouse) COVE score: 26 ttgggtgcgctggagccctggctgctgatgacgaaccgcctctaactgggcttgaccacgggtcggctctgaattgcagagaggctcgaaacagc >DIO3_SECIS_ratNor Rattus norvegicus (rat) COVE score: 26 ttgggtgcgctggagccctggctgctgatgacgaaccgcctctaactgggcttgaccacgggtcggctctgaattgcagagaggctcgaaacagc >DIO3_SECIS_canFam Canis familiaris (dog) COVE score: 26 ttgggtgctggcgagccccactgctgatgacgagccgcctctaactggtcttgaccacgagctggttctgagttgcaggggggcttgcagcggc >DIO3_SECIS_bosTau Bos taurus (cow) COVE score: 30 ttgggtgctcacgagccccactgctgatgaagagctgtctctaactggcctcgaccacgagctggttctgatttgcaggaggctcgcagcagc >DIO3_SECIS_borAnc Boreoeuthere ancestralis (ancestral) COVE score: 27 ttgggtgctcaggagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctggttctgaattgcagggggctcgcagcagca >DIO3_SECIS_loxAfr Loxodonta africana (elephant) COVE score: 22 ttcggtgcgctagagccccactgctgatgacgaactgtctctaactggtcttgaccacgagctgattccgaattgcagggaactcgcagcagc >DIO3_SECIS_echTel Echinops telfairi (tenrec) COVE score: - ttcggtgctctgcagccccactgctgatgacgaactgcctctcactggtcttgaccacgagctgcttctgaaatgcaggggactcgcagccgca