Opsin evolution: key critters (deuterostomes): Difference between revisions
Tomemerald (talk | contribs) |
Tomemerald (talk | contribs) |
||
| (83 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
'''See also:''' [[Opsin_evolution|Curated Sequences]] | [[Opsin_evolution:_key_critters_%28cnidaria%29|Cnidaria]] | [[Opsin_evolution:_key_critters_%28ecdysozoa%29|Ecdysozoa]] | [[Opsin_evolution:_key_critters_%28lophotrochozoa%29|Lophotrochozoa]] | [[Opsin_evolution:_update_blog|Update Blog]] | |||
=== Marsupials: Sarcophilus harrisii (tasmanian devil) .. 8 opsins === | |||
The 454-based genome of Sarcophilus harrisii (tasmanian devil) has recently become available (with better coverage than the new Macropus eugenii Sanger genome). Sarcophilus provides a fresh check on received wisdom on the mammalian opsin portfolio. Here the entire repertoire of opsin genes are collected for this species along with a summary of what is available for the marsupial clade and its current phylogenetic organization. As expected, Sarcophilus has but 8 of the 20 ancient vertebrate opsin genes. | |||
RHO1: 5 marsupials | |||
SWS2: 10 marsupials | |||
LWS: 10 marsupials | |||
ENCEPH: 3 marsupials | |||
TMT: 3 marsupials | |||
PER: 3 marsupials | |||
NEUR1: 3 marsupials | |||
MEL1: 4 marsupials | |||
The optimal wavelength for scotopic (dim light) vision of Sarcophilus is easily predictable provided key tuning residues are covered by the assembly. The 97% match to Sminthopsis and agreement at tuning residues suggests this aspect of vision will be nearly identical between the two species. | |||
Cone rhodopsin RHO2 has been lost in all mammals and no debris from this gene is expected in Sarcophilus. The short wavelength cone opsin SWS2, while still present in platypus, has also been lost in all theran opsin too long ago to leave detectable remnants in syntenic position. Cone opsin SWS1 has this turned around, being present in theran mammals but only as debris in platypus. A nearly full length gene, most similar to Sminthopsis, can be recovered from Sarcophilus read coverage. | |||
The basal long wavelength LWS imaging opsin is available from 97 vertebrates and has [[Opsin_evolution:_LWS_PhyloSNPs|already been analyzed]] for phyloSNPs and rare genomic events. The Didelphimorphia experienced a 3-4 residue insert in exon 1 that separates them from all other marsupials. Note this region has quite a complicated indel history. The extra residues have repeat character DVNE DDND suggesting replication slippage. The gene is present and intact in Sarcophilus though two exons are not currently available. LWS so far in tasmanian devil is identical to its Sminthopsis ortholog. | |||
LWS_loxAfr MAQQWGPHRLTGARLQDASE---DSTQASIFVYTNTNT elephant | |||
LWS_echTel MAQRWGAHRLTGGQLQDTYE---GSTRTSIFVYTNSTS tenrec | |||
LWS_monDom MTQAWDPAGFLARRRDVNE<font color="blue">DDND</font>ETTRSSLFVYTNSNN Didelphimorphia | |||
LWS_didAur MTQAWDPVGFLARRRDENE<font color="blue">DDHD</font>DTTRASLFVYTNSNN Didelphimorphia | |||
LWS_tarRos MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia | |||
LWS_macEug MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia | |||
LWS_smiCra MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Dasyuromorphia | |||
LWS_sacHar MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Dasyuromorphia | |||
LWS_setBra MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia | |||
LWS_cerCon MTQAWDPAGFLAWQEDENE----ETTRASLFVYTNSNN Diprotodontia | |||
LWS_myrFas MTQAWDPAGFLAWRREENE----ETTRASLFTYTNSNN Dasyuromorphia | |||
LWS_isoObe MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Peramelemorphia | |||
LWS_ornAna MTPAWNSGVYAARRRFEDEE---DTTRTSVFVYTNSNN platupus | |||
LWS_tacAcu MTQAWDPAGFLAWRRDENEE---TTRASLFVYTNSNNT echidna | |||
[[Image:MarsupTree.jpg|left]] | |||
Pinopsin, parapinopsin, parietopsin and VA opsin terminate in sauropods so are missing in all mammals including marsupials. Encephalopsin has a [[Opsin_evolution:_Encephalopsin_gene_loss|very peculiar history]] of gene loss in tetrapods, requiring some seven independent and asynchronous events including platypus. While this limits the phylogenetic utility of any gene loss within marsupials, the status of the gene within Sarcophilus is still informative. A full length gene can be recovered with 94% identity to opossum, strongly indicating that encephalopsin is fully functional within Sarcophilus. | |||
TMT is an ancient locus that is [[Opsin_evolution:_Encephalopsin_gene_loss#Post-marsupial_loss_of_TMT_encephalopsin|present in monotremes and marsupials]] but lost in all placentals. | |||
RGR has apparently been [[Opsin_evolution:_RGR_phyloSNPs|lost specifically in the marsupial clade]], though support for that is only provided by the Monodelphis and Macropus genome projects. It would be of considerable interest to find the gene or a fragment thereof in syntenic position in Sarcophilus. However nothing can be found with tblastn of current reads. | |||
Sarcophillus can be expected to have this gene and it does. The protein sequence substantiates the 4 [[Opsin_evolution:_Peropsin_phyloSNPs#PhyloSNPs_in_vertebrate_peropsins|previously defined phyloSNPs]] characteristic of the marsupial/placental transition. | |||
Here Sarcophilus can be [[Opsin_evolution:_Neuropsin_phyloSNPs|predicted]] to contain only NEUR1 because the ancient vertebrate genes NEUR2 and NEUR3 appear to terminate in sauropods and NEUR4 in platypus. | |||
Similarly Sarcophilus can be [[Opsin_evolution:_Melanopsin_gene_loss|expected]] to have the main melanopsin MEL1 but not the paralog MEL2 which terminates in sauropods. | |||
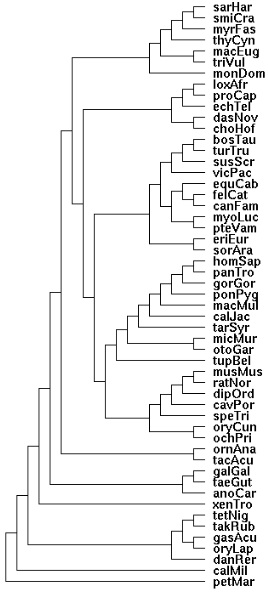
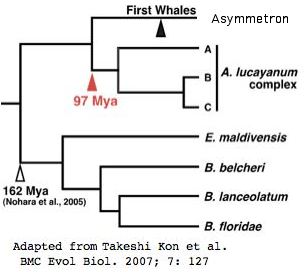
Marsupial phylogenetic relationships have been reviewed in a [http://genome.cshlp.org/content/19/2/213.full 2009 paper] that established the mitochondrial genome sequences of the Tasmanian tiger (Thylacinus cynocephalus) and numbat (Myrmecobius fasciatus). It emerges from this that thylacine is basal among Dasyuromorpha with numbat diverging next: (thyCyn,(myrFas,(smiCra, sarHar))). Dunnart and tasmanian devil are very similar at the protein level, in some cases 100% identical. | |||
<pre>Newick tree that generates the marsupial-centric vertebrate phylogenetic tree: | |||
((((((((((((sarHar,smiCra),myrFas),thyCyn),(macEug,triVul)),monDom), | |||
((((loxAfr,proCap),echTel),(dasNov,choHof)), | |||
((((((bosTau,turTru),susScr),vicPac),((equCab,(felCat,canFam)),(myoLuc,pteVam))),(eriEur,sorAra)), | |||
(((((((((homSap,panTro),gorGor),ponPyg),macMul),calJac),tarSyr),(micMur,otoGar)),tupBel), | |||
(((((musMus,ratNor),dipOrd),cavPor),speTri),(oryCun,ochPri)))))), | |||
(ornAna,tacAcu)), | |||
((galGal,taeGut),anoCar)), | |||
xenTro), | |||
(((tetNig,takRub),(gasAcu,oryLap)),danRer)), | |||
calMil), | |||
petMar); | |||
</pre> | |||
<br clear="all"> | |||
<pre> | |||
>RHO1_sacHar Sarcophilus harrisii (tasmanian_devil) 97% identity Sminthopsis crassicaudata | |||
0 MNGTEGPNFYVPHSNKTGVVRSPYEEPQYYLAEPWMFSCLAAYMFMLIVLGFPINFLTLYVTIQHKKLRTPLNYILLNLAVADLFMVICGFTTTLVTSLNGYFVFGTTGCQIEGFFATTG 1 | |||
2 GEVALWALVVLAIERYIVVCKPMSNFRFGENHAIMGVVFTWIMALACSVPPLFGWSR 2 | |||
1 YIPEGMQCSCGIDYYTLNPEFNNESFVIYMFVVHFTIPLTVIFFCYGQLVFTVKE 0 | |||
0 AAAQQQESATTQKAEKEVTRMVIIMVIAFLICWVPYASVAFYIFTHQGSDFGPIFMTLPAFFAKSSSIYNPVIYIMMNKQ 0 | |||
0 FRTCMITTLCCGKNPLGDDEASATVSKTETSQVAPA* 0 | |||
>SWS1_sacHar Sarcophilus harrisii (tasmanian_devil) part of last exon missing 96% identity Sminthopsis crassicaudata | |||
0 MSGDEEFYLFKNISPVGPWDGPQYHIAPAWAFHLQTAFMGFVFFAGTPLNGVVLIATLRYKKLRQPLNYILVNISLAGFIFCVFSVFTVFVSSSQGYFVFGRHVCAMEGFLGSVA 1 | |||
2 SGLVTGWSLAFLAFERFIVICKPFGNFRFHSKHATMVVLATWVIGIGVSIPPFFGWSR 2 | |||
1 YIPEGLQCSCGPDWYTVGTKYRSEYYTWFLFIFCFIVPLSLICFSYSQLLGALRAVS 0 | |||
0 VAAQQQESATTQKAEREVSRMVVVMVGSFCLCYVPYAALAMYMVNNRNHGLDLRLVTIPAFFSKSACVYNPIIYCFMNQ 0 | |||
0 KPMTDDSETTSSQKTEVSTVSSSQVGPS* 0 | |||
>LWS_sacHar Sarcophilus harrisii (tasmanian_devil) half of exon 2, all of exon 4 missing frag 100% identical to Sminthopsis | |||
0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTK 1 | |||
2 FKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 | |||
2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 | |||
1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMSTCCILPLSIIILCYIQVWLAIRA 0 | |||
0 0 | |||
0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 | |||
>ENCEPH_sacHar Sarcophilus harrisii (tasmanian_devil) 94% identity monDom | |||
0 MYSGNSSDDAGGGYWGSGGTGGAGGTGVAGEPAPEGSPRPAPLFSPGTYELLALLIATIGLLGLCNNLLVLVLYYKFQRLRTPTHLFLVNISFSDLLVSLFGVTFTFVSCLRSGWVWDSVGCAWDGFSNTLF 1 | |||
2 GIVSIMTLTVLAYERYNRIVHAKVINFSWAWRAITYIWLYSLIWTGAPLLGWNRYTLEIHGLGCSVDWKSKDPNDSSFVLFLFLGCLVLPVGVMAYCYGHILYAIRM 0 | |||
0 FRCVEELQTLQVIKILRYEKKVAKMCFLMIATFLFCWMPHAVICFLVANGYGSLVTPTVAIIPSLFAKSSTAYNPIIYIFMSRK 0 | |||
0 FRRCLLQLLcFRQLKFQQPKKDRAIIRTEKQIRPIVMSQKVGDRPKKKVTFSSSSIIFIITSDETQMIDDNDKNSETKVNVIQVRPL* 0 | |||
>TMT_sacHar Sarcophilus harrisii (tasmanian_devil) FP5MBH101BETOZ needed to finish | |||
0 MSINLTTNLSFGPLLIDSEEKPRSGLSRTGHTVVAVFLGIILILGFINNFIVLILFCKFKVLRNPVNMLLLNISISDMLVCLSGTTLSFASSIRGRWIGGYHGCRWYGFANSCF 1 | |||
2 GIVSLISLAILSYERYRTLTLCPRRGADYQKALLAVAGSWLYSLIWTVPPLIGWSSYGTEGAGTSCSVHWTSKSVQSVSYIMCLFIFCLVIPILIMIYFYGRLLYTVKQ 0 | |||
0 VGKIRKTAARKREYHVLFMVVTAVICYLICWVPYGLIALVATFGPPGVVSPVANIVPSILAKSSTVCNPIIYILMNKQ 0 | |||
>PER_sarHar Sarcophilus harrisii (tasmanian_devil) 5.5 of 7 exons | |||
0 MFKNDSFRSLEPEKEGHSVFSPAEHNIVAAYLITA 1 | |||
2 SILSNVIVLGIFVKFKELRTATNAIIINLA 0 | |||
0 1 | |||
2 GRRMTSFNYTIMILTAWVNGFFWALMPIVGWASYAPDPTGA 2 | |||
1 SFVSYTVTVIAINFVMPLVVMIYCYYNVSQKIKQYTPSNCPEYINRDWSNEVAVTK 0 | |||
0 MSVIMILMFLLAWSPYSVVCLWASFGDPKEIPPAMAIIAPLFAKSSTFYNPCIYVAANKK 2 | |||
1 FRrAISAMIQCQTHQSMSVSKALPMN* 0 | |||
>NEUR1_sarHar Sarcophilus harrisii (tasmanian_devil) 4 of 6 exons | |||
0 1 | |||
2 GVLSTLGNGYVIYMSSKRKKKLRPAEIMTVNLAVCDLGIS 1 | |||
2 VVGKPFTIISCFSHRWVFGWVGCRWYGWAGFFFGCGSLITMTAVSLDRYLKICHLSY 1 | |||
2 GTWLKRHHAYICLVIIWAYATFWATMPLAGLGNYAPEPFGTSCTLDWWLAQASVTGQTFILNILFFCLLLPTAVIVFSYVKIIAKVKSSTKEVAHFDSRIQNSHSHVLEMKLTK 0 | |||
0 VAMLICAGFLIAWIPYAVVSVWSAFGQPDSIPVQFSVVPTLLAKSAAMYNPIIYQVIDCKFACCQSGGQKAAKKESLRDYR 2 | |||
1 * 0 | |||
>MEL1_sarHar Sarcophilus harrisii (tasmanian_devil)96% identity smiCra last exon missing FKUJDAX01C1KMN needed | |||
0 MNPSPMLRHLSCSAQDTNCTKIMASISEWNNTEVDAYHLVDLPPITPT 0 | |||
0 AVVLPPYSQNVFPTADVPDYAHYTIGATILVVGFTGVLGNLLVIYTFCR 2 | |||
1 SRSLRTPANMFIINLAISDFFMSFTQAPVFFASSLYKRWIFGEK 2 | |||
1 GCEFYAFCGALFGITSMITLMVIALDRYFVITRPLASIGMISKKKTGLILLGVWLYSLAWSLPPFFGW 1 | |||
2 sAYVPEGLLTSCSWDYTTFTPSVRAYTILLFCFVFFIPLIVIIYCYIFIFRAIKDTNK 2 | |||
1 AVQNIGSRASTPSPRHFQRMKNEWKMAKIALVVILLFVLSWAPYSTVALVAFAG 2 | |||
1 YSHVLTPYMNSVPAIIAKASAIHNPIIYAISHPKYR 2 | |||
1 MAIAQNFPCLRAVLGIRHPRTQSFSSYRFTHRSTTASQASDISWQSRGRRQLSLGSESEA 0 | |||
0 GWNNIEAGIEGLTLRSLEGYCGMDEETMETREPSASAKAKGQ 0 | |||
0 * 0 | |||
</pre> | |||
=== Snakes: Python molarus (python) .. 11 opsins === | |||
The python genome became available in Feb 2011. Snakes fill an important data niche -- while still amniotes, they are sister to the squamate lizard Anolis and together these are sister to the three birds with determined genomes. These determine the diapsid node; when a turtle genome becomes available, that will allow reconstruction of the sauropsid node of mammalian divergence. | |||
In terms of opsins, ten are completely missing from the python assembly: RHO2, SWS2, PIN, PPIN, PARIE, TMTa, TMT2, TMT3, MEL1, and NEUR3. Here TMT2 is definitely expected whereas its paralogs have perhaps already headed out the door, phylogenetically speaking. While this could reflect inadequate coverage, the opsins that are present seem to have good coverage (nearly all their exons). However it should be noted that contigs are very small, often 1-2 kbp, barely an assembly at all. | |||
The 11 opsins from this assembly are shown below. Missing exons are shown in <font color= "blue">blue</font>. | |||
>RHO1_pytMol Python molarus (python) | |||
0 MNGTEGLNFYVPMSNKTGIVRSPFEYPQYYLAEPWKYSALGAYMFLLILLGFPINFLTLYVTIQHKKLRTPLNYILLNLAIANLFMVLVGFTTTMYTSMNGYFVFGTVGCNVEGFFATLG 1 | |||
2 GEMALWSLVVLAIERYVVVCKPMSNFRFTETHAIMGLCFTWIMALACAGPPLVGWSR 2 | |||
1 YIPEGMQCSCGVDYYTPTPEVHNESFVIYMFIVHFVIPLAVIFFCYGRLVCTVKE 0 | |||
0 AAAQQQESATTQKAEKEVTRMVIIMVIAFLICWVPYASVAFYIFTHQGSDFGPVFMTIPAFFAKSSAIYNPVIYIVLNKQ 0 | |||
0 FRNCMITTLCCGKNPLAEDDTSAGTKTETSTVSTSQVSPA* 0 | |||
>SWS1_pytMol Python molarus (python) | |||
0 MSGEEDFYLFENISSVGPWDGPQYHIAPMWAFHFQTLFMGLVFFAGTPLNAIILIVTIKYKKLRQPLNYILVNISFAGLLFCVFAVFTVFLASSQGYFFFGRHVCALEAFLGSVA 1 | |||
2 GLVTGWSLAFLAFERYVVICKPLGNFRFNSKHALLVVVATWVIGVGVSVPPFFGWSR 2 | |||
1 FIPEGLQCSCGPDWYTVGTKYKSEYYSWFLFVFCFIVPLSLIVFSYLRLLGALRA 0 | |||
0 VAAQQQESATTQKAEREVSRMVVVMVGSFCLCYVPYATLAMYMVNHPKHGLDLRLVTIPAFFSKSSCVYNPIIYCFMNKQ 0 | |||
0 FRACIMQTVCGKPLTDESDAGSSAQKTEVSSVSSSQVSPSSGAGATGSPRS* 0 | |||
>LWS_pytMol Python molarus (python) | |||
0 MQTRGKEVAKAMTEAWNVAVFAARRRNDDDTTRESVFIYTNSNNTR 1 | |||
2 GFEGPNYHIAPRWVYNLTSLWMVFVVIASVFTNGLVLVATAKFKKLRHPLNWILVNLAIA DLGETVIASTISVINQFFGYFILGHPLCVVEGYTVSVC 1 | |||
2 GITGLWSLAIISWERWVVVCKPFGNLKFDAKLALAGILFSWVWSATWTAPPIFGWSR 2 | |||
1 YWPHGLKTSCGPDVFSGNEDPGVQSYMIVLMVTCCIIPLSIIILCYLQVWMAIRA 0 | |||
0 VAAQQKESESTQKAEKEVSRMVVVMILAYIFCWGPYTTFACFAAANPGYAFHPMTASLPAFFAKSATIYNPIIYVFMNRQ 0 | |||
0 QFRNCIMQLFGKKVDDGSEVSSTSRTEVSSVSNSSVSPA* 0 | |||
>VAOP_pytMol Python molarus (python) | |||
0 MDGARRESENGSLLLDASAGSSLVDPFSQPLDSIESWNFHLLSALMFLVTSLSLFENFTVILVTLKFRQLRQPLNYVIVNLSVADFLVSLVGGTISFLTNLKGYFYMGRWACVLEGFAVTFF 1 | |||
2 GIVALWSLALLAFERYVVICRPLGNVHFKGKDATLGIAFVWIFSFIWTIPPTMGWSSYTTSKIGTTCEPNW 2 | |||
1 YSGEYSDHTFIITFFTTCFIMPLLIILVSYGKLMRKLRK 0 | |||
0 <font color= "blue">vsntqg</font>RLGSTRKPERQVTRMVVIMILAFLICWTPYAAFSILVTTCPSLELDPRLAAIPAFFSKTATVYNPVIYVFMNNQ 0 | |||
0 FRKCLVQLFRCSRQDIADSNMNQISKRAVLTPTKKYGEVSTAAARVTIFNQRNEDERSSsQSFALLSVLENWLCQFLRG* 0 | |||
>ENC_pytMol Python molarus (python) | |||
0 MYSGNGSGGSLEPTAGKQDLERRRQQPAGDSGTPFSASTYELLALLIAAIGFVGLCNNLLVLVVYTKFKRLRTPTNLFLVNISLSDLLVSLFGVSFTFLSCLRNHWAWDAAGCVWDGFSNSLFG 1 | |||
2 GIVSLMTLTVLAYERYIRVVHARVVDFSWSWRAITYIWLYSLAWTGAPLLGWNHYTLELHGLGCSVDWSSREPGDTSFVFFFFLGCLVAPLGIVVYCYGHILHAVRM 0 | |||
0 FRRVEELQTVHVIKILRYEKKVAKMCFLMITTFLICWMPYAVVSLLIAYGYGHLITPTVAIIPSFFAKSNTAYNPVIFIFMSRK 0 | |||
0 FRRCLVQLFCVQFLRFKRTLKEQPAIESNKPIRPIVMSQKVGDRPKKKVtfssssiifiitsDDTEQIDVSTKCSDTKINVIQVKPL* 0 | |||
>MEL2_pytMol Python molarus (python) | |||
0 MATAHPTKVDAPDYVLYAVGSCVLVIGCIGIIGNLLVLYAFYS 2 | |||
1 NKRLRTPPNYFIMNLAVSDFLMCATQAPVCFFNSIHKEWVLGDT 1 | |||
2 GCNFYAFCGALFGITSMMTLLAISVDRYCVITKPLQSIKRSSKKRSCLIIMFVWLYSLGWSVCPLFGW 1 | |||
2 EKDIPEGLMISCTWDYVTYSPANRSYTMLLCCCVFFIPLIIIFHCYLCMFLAIRNTGR 2 | |||
1 RKHSASQNIKSEWKLAKIAFVAIVVYVVSWSPYACVTLIAWAG 2 | |||
1 YARILTPYSKSVPAIIAKASAIYNPIIYAIIHPSYR 2 | |||
1 <font color= "blue">RTIRSAVPCLRFLIPISKSDLSTSSMSESSFRASVSSRHSFSYRNKSTYISSISAKET 0</font> | |||
0 TWNNVELDPVESVHTKLQPPQSNSFSTNAEEKSELPMKAPGCDVPTEEKVKAVFSNAPIHFTNKSSCVQAQSGVLPPSTVVVVMFIQ 0 | |||
>RGR1_pytMol Python molarus (python) | |||
0 MTVSDSLPEGFTELEMFSFGTVMLAE 1 | |||
2 ALFGFSLNVLTIVSFWKITELQTPGNFLIFNLALSDCGICINALIAAFSTFLR 2 | |||
1 RYWPYGSDGCQIHGFQGFLTALTSINCAAAIAWDRYHQYCT 1 | |||
2 RSKLQWNSVFSMGLCAWCFAGFWSAMPLLGWGTYDYEPLRTCCTLDYTKGDR 2 | |||
1 NYIMFLIPLVLFNFVIPIFIMLMSYQSIDNKFRKTAQVK 0 | |||
0 FNTGLPVKSLVICWGPYSLLSFYAAVENVAFVSPKILM 0 | |||
0 IPALMAKTSPTMNAFIYALGNENYRGGMWQFLTGEKIEKAKIDDKTN* 0 | |||
>PER1_pytMol Python molarus (python) | |||
0 <font color= "blue">MRRNDSANLLESEHHDRSAFSQTDHNIVAAYLITA 1</font> | |||
2 GVISLLSNIVVLSIFIKFKELRTPSNTIIIHLAFTDIGVSSIGYPMSAASDLHGSWKFGYMGCQ 0 | |||
0 <font color= "blue">IYAGLNIFFGMSSIGLLTVVAVDRYLTICRPAI 1</font> | |||
2 GNRLTAHNYIALIFAAWTNAVFWASMPVVGWASYAPDPTGATCTINWRENDM 2 | |||
1 SFISYTMTVIAVNFAVPLIVMFYCYYNVSKAMRQYPASRVLENLNIDWSEQVDVTK 0 | |||
0 MSVIMILMFLLAWSPYSIVCLWSSFGDPKKIPPTMAIIAPLFAKSSTFYNPCIYVIANK 2 | |||
1 <font color= "blue">FRRAMLSMVQCQTHREITITDVLPMNRSRSPH* 0</font> | |||
>NEUR1_pytMol Python molarus (python) | |||
0 MYLRQNASSQDINLPHYQRDGDPFASKLSKEADIIAGIYLLII 1 | |||
2 GIMSTLGNGYVIYMSIQRKRKLRPAEIMTVNLAVCDLGIS 1 | |||
2 VTGKPFSVIAFLSHRWIFGWSGCRWYGWAGFFFGVGSLITMTAVSLDRYFKICYLAY 1 | |||
2 GIWLKRHHAFICLGIIWSYAVFWATIPFAGLGNYAPEPFGTSCTLDWWLAQGSVAGQAFILNILFFCLVFPTAVIVFSYIKIIAKVKSSSKEVAHYDNRFQNSHELEMKLTK 0 | |||
0 VAVLICAGFLIAWIPYAAVSVWSAFGKPDSVPIKVSVVPTLLAKSAAMYNPVIYQVINCKSVCCQPEALRPLQKKNSLNKSR 2 | |||
1 <font color= "blue">VYTISTFRKSTTSAR* 0</font> | |||
>NEUR2_pytMol Python molarus (python) | |||
0 <font color= "blue">MDPSFANSTFQSKITEAADIVVGTCYMVF 1</font> | |||
2 GVVSLFGNSLLLWVAYRKRAILKPAEFFIVNLAVSDIGMTVILFPLATPSFFAHR 2 | |||
1 <font color= "blue">WLYGKHICLFYAFCGLFFGICSLSTLTLLSVVCCLKICFPAY 1</font> | |||
2 GNKFSPPYAGVLLVCVWIYAFIFAAAPLADWGSYGPEPYGTACCIKWKASTREAKFYIMALFVFCYIIPCILILISYSLILWTVKVSRRAVRQHMSPQSKHNSVHSLIVK 0 | |||
0 LSISVCIGFLAAWTPYAIVAMWAAFGDPSKVPILAFALSAVFAKSSATYNPLVYLLFKPNFQKFLSKDLSFFQAVCAVCCCSRSRVITLQSFHTRDGRASMRFSTA | |||
FTDHRGSCRNCSDTFECFSNYPRCYRLTQKSDPASKTNRLAILTDGRACRPSFKRTVQVMVLMTRKKTGIGTMNVAGDVLPSNIVRDLM* 0 | |||
>NEUR4_pytMol Python molarus (python) | |||
0 MSFQVSVQAPWRNNNMTFLNKDHPVSEQGETIIGFYLLTL 1 | |||
2 GWMSWIGNSIVIFVLYKQRAVLQPTDYLTFNLAVSDGSIAIFGYSRGIIDIFNVFQDDGFLVTSIWTCQV 0 | |||
0 VDGFLTLLFGLSSINTLTVISVTRYIKGCHPNR 1 | |||
2 GHCISTSSISVAIFLIWTAALFWSLAPFLGWGSYr 1 | |||
2 DRMYGTCEIDWTKANFSTTYKSYIVSIFICCFFMPVMVMVFSYMSIINTVKSSHALSGMGDPTDRQRRMEQSVTR 0 | |||
0 VSLVVCTCFIAAWSPYAVISMWSASGCPVPNLTSIFASLFAKSASFYNPIIYFGMSSKFRKDIAVLFHFAKEIKDPVKLKQFKILKQKLDSSPSHGEEKGAIDAQPALHSDSGVGSHPNTPPPVNRKGYFVVFDNWPQNPDFECDRL* 0 | |||
=== <span style="color: #990099;">Chondrichthyes: Callorhinchus milii (elephantshark) .. 17 opsins</span> === | |||
Six ray-finned fish genomes and massive transcript studies in yet other Clupeomorpha are available but the usefulness of this data is complicated by lineage-specific expansions and very rapid evolution of protein sequences. Little data is available from lobed-finned fish; whereas coelacanth genome has been proposed, that has stalled at 169,000 traces as of May 2008. | |||
This makes the [http://www.sciencemag.org/cgi/content/full/314/5807/1892 preliminary genome assembly] of the much earlier diverging Callorhinchus (oft-misspelled) and numerous skate transcripts very special because they are the "last stop" before the very difficult lamprey genome (currently assembled but with contigs seldom larger than 1-2 exons and only rarely containing syntenic information). | |||
[[Image:CalMil.png|left|]] | [[Image:CalMil.png|left|]] | ||
This [http://www.flmnh.ufl.edu/fish/Gallery/Descript/GhostShark/GhostShark.html large-eyed cartilaginous fish] lives to depths to 200m on the continental shelf of southern Australia and New Zealand but migrates into coastal estuaries to lay egg cases in sand and muddy substrates. The distinctively-shaped egg cases are sometimes found washed ashore after storms. They are up to 25cm long, 10cm wide, and take up to eight months to hatch. The one member of the genus studied has a vitamin A1-based photopigment with maximum absorbance at 499 nm presumably adapted to the overall photic environment at that depth. | This [http://www.flmnh.ufl.edu/fish/Gallery/Descript/GhostShark/GhostShark.html large-eyed cartilaginous fish] lives to depths to 200m on the continental shelf of southern Australia and New Zealand but migrates into coastal estuaries to lay egg cases in sand and muddy substrates. The distinctively-shaped egg cases are sometimes found washed ashore after storms. They are up to 25cm long, 10cm wide, and take up to eight months to hatch. The one member of the genus studied has a vitamin A1-based photopigment with maximum absorbance at 499 nm presumably adapted to the overall photic environment at that depth. | ||
Sequencing of the Callorhinchus genome resumed in the fall of 2008 with 454 technology; those reads are released at GenBank short read archive but are not blastable online. New Sanger 'finishing' reads from WUGSC became available at NCBI trace archives in mid-May 2009. From an exhaustive search of elephantfish data in WGS and Trace divisions of GenBank on 5 Nov 2007, many opsin exons but few complete genes were recovered and posted here. The opsin classifier reliably assigns these fragments to their ortholog class. | |||
In Jan 2009, Davies et al reported sequencing [http://genome.cshlp.org/content/early/2009/02/04/gr.084509.108.full.pdf+html full length RHO1, RHO2 and duplicated LWS1 and LWS2 genes] in Callorhinchus, further establishing that SWS1 and SWS2 are missing in this basal chondrichthyes. That reflects a lineage-specific loss because Geotria lamprey has both.(These sequences have not yet been released from hold by GenBank but can be recovered from Fig. 3.) | |||
Overall, Callorhinchus has good complement of vertebrate opsin genes. Parietopsin is also missing to date. Two encephalopsin- and two melanopsin-class opsins were found. The RGR, peropsin, and neuropsin genes will prove important in better determining their unresolved overall gene tree placement (which an October 2007 [http://www.plosone.org/article/info:doi/10.1371/journal.pone.0001054 opsin phylogeny paper] placed deeply within rhabdopsins) but more commonly classify as very basal ciliary opsins. | |||
There are 17 Callorhinchus opsins currently available in the reference collection: RHO1, RHO2, LWS1, LWS2, PIN, PPIN, RGR1, TMTa1, TMTa2, VAOP, ENCEPH, PER1, NEUR1, NEUR2, NEUR4, MEL1b, and MEL1. | |||
=== <span style="color: #990099;">Agnatha: Petromyzon marinus (lamprey) .. 9 opsins + 2 pseudogenes</span> === | === <span style="color: #990099;">Agnatha: Petromyzon marinus (lamprey) .. 9 opsins + 2 pseudogenes</span> === | ||
Lamprey (and less-studied hagfish) are the closest surviving | Lamprey (and less-studied hagfish) are the closest surviving outgroups to jawed vertebrates and thus central to reconstructing that last common ancestor. This importance is accentuated by the considerable temporal and evolutionary distance to the preceding two divergence nodes, cephalochordate (Branchiostoma) and urochordate (Ciona), especially for opsins because imaging eyes and advanced color vision emerged within a fairly compressed time frame within the lamprey stem. | ||
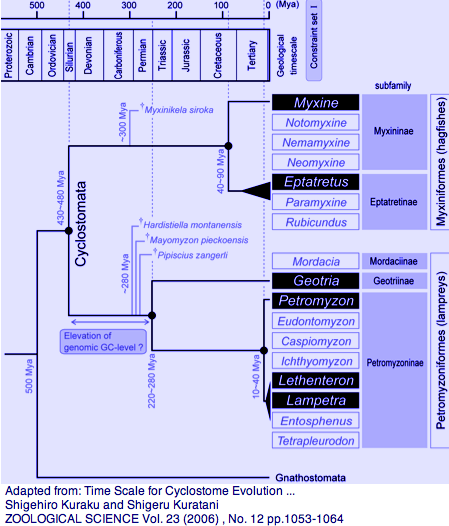
[[Image:LampreyChron.png|left|]] | [[Image:LampreyChron.png|left|]] | ||
| Line 19: | Line 251: | ||
Among extant lamprey, the genera Geotria and Petromyzon split 280-220 myr ago (helpful in breaking up the 500 myr long branch) whereas Lethenteron/Petromyzon split much later at 20 myr (helpful in locating orthologs from the former in the sequenced genome of the latter). There are no opsin sequences available for the third lamprey family, represented by Mordacia praecox, but it too is observed to have [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=18456882 multifocal crystalline lenses that compensate for longitudinal chromatic aberration.] | Among extant lamprey, the genera Geotria and Petromyzon split 280-220 myr ago (helpful in breaking up the 500 myr long branch) whereas Lethenteron/Petromyzon split much later at 20 myr (helpful in locating orthologs from the former in the sequenced genome of the latter). There are no opsin sequences available for the third lamprey family, represented by Mordacia praecox, but it too is observed to have [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=18456882 multifocal crystalline lenses that compensate for longitudinal chromatic aberration.] | ||
Photoreceptor systems have been studied extensively [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=9427550,17463225,15683562,15312025,11774340,9210581,9082962,2776863,2834011,2,12916734 in both larva and adults.] Geotria australis has a full complement of sequenced imaging porphyropsins LWS, SWS1, SWS2, RHO2, and RHO1 (RHO2 not so clearly a | Photoreceptor systems have been studied extensively [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=9427550,17463225,15683562,15312025,11774340,9210581,9082962,2776863,2834011,2,12916734 in both larva and adults.] Geotria australis has a full complement of sequenced imaging porphyropsins LWS, SWS1, SWS2, RHO2, and RHO1 (RHO2 not so clearly a straightforward ortholog to jawed vertebrate counterparts), though its non-pineal opsin classes are not as well characterized as Petromyzon. This implies that the ancestral vertebrate possessed already possessed full photopic (bright light) cone-based color vision with the potential for pentachromacy, multi-focal lenses, pigment filtration, and likely circadian rhythm, pupillary reflex, and pineal and parapineal functions. Photoreceptor morphology and spectral sensitivity can change dramatically during various phases of the lamprey lifecycle, presumably adaptively; lineage-specific issues are not a central focus here. | ||
An April 2009 paper directly characterizes [http://www.ncbi.nlm.nih.gov/pubmed/19398493 Petromyzon imaging opsins]. RHO1 and LWS, regenerated with 11-cis retinal rather than 3-dehydroretinal porphyropsin, give λmax values of 501 nm and 536 nm, respectively. Bidirectional experimental substitution show the S164P substitution accounts for the LWS blue-shift of 19 nm relative to Geotria. | |||
This site normally [[Opsin_evolution:_LWS_PhyloSNPs|waffles between serine and alanine]] but proline also occurs in pufferfish, turbot and japanese lamprey (4% of 94 available LWS sequences). Consequently the ancestral lamprey state was proline in the Lethentron/Petromyzon ancestor at 20 myr and S/P at the ancestral node with Geotria with serine favored because the two available chondrichthyian LWS sequences have serine. The timing is consistent with, but not proof of, a compensatory shift in Petromyzon that accompanied loss of SWS2 and/or SWS1. Six of eight lamprey GenBank genera remain unsequenced. If LWS is absent or lost from hagfish, then no earlier diverging outgroup can ever be found. It is better to view the value of each ancestral residue as a compositional vector that accommodates natural population polymorphism abundances and uncertainties in reconstruction. | |||
The authors confirm the August 2008 report here that RHO2 is completely lost, SWS2 a very deteriorated pseudogene, and SWS1 a fairly recent pseudogene, ie the pentachromat Geotria better represents the ancestral lamprey state. Pseudogenes presumably evolve at the neutral mutation rate subsequent to the initial inactivation event, yet this could be influenced by chromosomal context and base composition issues. | |||
However dating may work out quantitatively, it seems clear that Petromyzon passed through tetrachromat and trichromat post-Cretaceous stages separated by perhaps tens of millions of years. The loss of these opsins may not be adaptive so much as reflect 'use it or lose it'. Some advantage might accrue if the number of cones remained fixed and SWS vacancies were occupied by additional LWS, yet it is not so clear how LWS would displace suboptimal SWS in the early stages of pseudogenization. | |||
Should lampreys be considered living fossils? The 360 mya fossil Priscomyzon riniensis [http://www.ncbi.nlm.nih.gov/pubmed/17066033 closely resembles] living lampreys in external morphology, large oral disc, circum-oral teeth and branchial basket. However many aspects of soft tissue anatomy are not preserved, leaving their constancy a matter of speculation. Lamprey larva being filter feeders, the question arises of what ancestral adult lamprey ate prior to the evolution of fish. | |||
Lamprey are definitely not living fossils from the molecular point of view. Lamprey proteins evolve rapidly -- it is the norm for a cephalochordate protein to match human ortholog better than lamprey despite greatly increased branch length. This divergence of lamprey proteins is not just pointwise residue change but also involves indels and seemingly radical changes in protein character. This change could be mostly neutral relative to the ancestral lamprey state (function molecular fossil), but the fact is, Branchiostoma (sequence molecular fossil) sheds far more light on its ancestral divergence node than does Petryomyzon. At some point the ability to reconstruct ancestral genomes will be limiting for our understanding of craniate evolution. | |||
Opsins are by no means the only fully developed gene system to have emerged in vertebrates by the time of lamprey divergence. The rod and cone cyclic nucleotide phosphodiesterases of Petromyzon consist of a single PDE6 catalytic subunit EF432251 but two inhibitory Pgamma subunits expressed in the long and short photoreceptors, EF427669 and EF470978 respectively. These are not in tandem position in the current assembly. Recently it has also become clear that the androgen receptor, GABA receptor network, and many other systems need to be backdated to agnathans. | Opsins are by no means the only fully developed gene system to have emerged in vertebrates by the time of lamprey divergence. The rod and cone cyclic nucleotide phosphodiesterases of Petromyzon consist of a single PDE6 catalytic subunit EF432251 but two inhibitory Pgamma subunits expressed in the long and short photoreceptors, EF427669 and EF470978 respectively. These are not in tandem position in the current assembly. Recently it has also become clear that the androgen receptor, GABA receptor network, and many other systems need to be backdated to agnathans. | ||
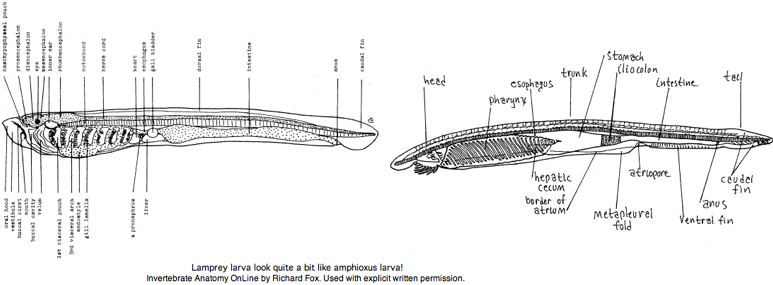
The larvae (ammocoetes or 'sand dweller') are often said blind. It is more accurate to say larva have developmentally arrested eyes but with a small region of differentiated retina. That potentially allows for limited imaging vision -- opsin expression has not been examined. The larval stage, which can last up to 17 years, was considered a separate species by early workers. The [http://webs.lander.edu/rsfox/invertebrates/ammocoetes.html anatomical resemblance] of lamprey larva to (adult) amphioxus is quite remarkable, a case perhaps of ontogeny recapitulating phylogeny. | The larvae (ammocoetes or 'sand dweller') are often said blind. It is more accurate to say larva have developmentally arrested eyes but with a small region of differentiated retina. That potentially allows for limited imaging vision -- opsin expression has not been examined. The larval stage, which can last up to 17 years, was considered a separate species by early workers. The [http://webs.lander.edu/rsfox/invertebrates/ammocoetes.html anatomical resemblance] of lamprey larva to (adult) amphioxus is quite remarkable, a case perhaps of ontogeny recapitulating phylogeny. | ||
<br clear ="all"> | <br clear ="all"> | ||
The metamorphosis into adult lamprey is every bit as profound as that of amphibians, involving a radical rearrangement of internal organs, continued development of eyes and transformation from sediment-dwelling filter feeder into pelagic marine vertebrate predator. Adult lampreys, like many early-diverging vertebrates, have three eyes: a pair of lateral eyes on the sides of the head and a single, unpaired median eye on top of the head. Each lateral eye is a spherical black structure lying beside the posterior prosencephalon that marks its boundary to the mesencephalon. | |||
The metamorphosis into adult lamprey is every bit as profound as that of amphibians, involving a radical rearrangement of internal organs, continued development of eyes and transformation from sediment- | |||
[[Image:Opsin Lamp Amph.png|left|]] | [[Image:Opsin Lamp Amph.png|left|]] | ||
| Line 39: | Line 281: | ||
The lamprey genome project has resulted in an [http://www.genome.wustl.edu/genome/other_vertebrates/Petromyzon_marinus/seq_plan.cgi anemic assembly] despite accruing 19 million traces. It seems a 16 kbp retroposon has expanded enormously, which in conjunction with very high GC content and high levels of heterozygosity makes assembly of the 2.4 Gbp genome quite difficult. However the [http://www.genome.wustl.edu/tools/blast/ blast page at WUSTL] allows a Petromyzon "3.0 supercontig" tblastn option. UCSC provided a lamprey browser In January 2008. Gene name searches work better than Blat because of very considerable divergence of protein sequences. | The lamprey genome project has resulted in an [http://www.genome.wustl.edu/genome/other_vertebrates/Petromyzon_marinus/seq_plan.cgi anemic assembly] despite accruing 19 million traces. It seems a 16 kbp retroposon has expanded enormously, which in conjunction with very high GC content and high levels of heterozygosity makes assembly of the 2.4 Gbp genome quite difficult. However the [http://www.genome.wustl.edu/tools/blast/ blast page at WUSTL] allows a Petromyzon "3.0 supercontig" tblastn option. UCSC provided a lamprey browser In January 2008. Gene name searches work better than Blat because of very considerable divergence of protein sequences. | ||
New sequencing technologies make affordable an immense collection of cDNA a potential solution in lamprey. It would permit a pseudo-assembly of exons flanked by some at least genomic dna. That is, transcripts are aligned into the trace archives to obtain non-coding context for exons. This would allow restricted topics to be studied (intron retention, invariant non-coding) but not others (upstream regulatory, chromosomal gene order). | New sequencing technologies make affordable an immense collection of cDNA a potential solution in lamprey. It would permit a pseudo-assembly of exons flanked by some at least genomic dna. That is, transcripts are aligned into the trace archives to obtain non-coding context for exons. This would allow restricted topics to be studied (intron retention, invariant non-coding) but not others (upstream regulatory, chromosomal gene order). Labeling techniques such as FISH could provide some linkages but not gene-level order; perhaps this could be done at the level of BACs using exons from all possible gene pairs. Alternatively, the genome of Geotria might present fewer issues; it would certainly be better for the study of opsins. | ||
Petromyzon sequences are very disorganized at GenBank. Some 147,000 conventional ESTs are stored inappropriately at the trace archives. These are attributed to WUGSC genome sequencing center but that site makes no mention of the project. Another 120,731 short ESTs from a MSU group are stored in the conventional location but the individual lamprey used may be only distantly related to that used in the genome project. Some 32,000 454-technology cDNA reads are also available but stored in the trace archives but not where they belong (short read archive SRA) but rather in the Petromyzon_marinus_OTHER database. | Petromyzon sequences are very disorganized at GenBank. Some 147,000 conventional ESTs are stored inappropriately at the trace archives. These are attributed to WUGSC genome sequencing center but that site makes no mention of the project. Another 120,731 short ESTs from a MSU group are stored in the conventional location but the individual lamprey used may be only distantly related to that used in the genome project. Some 32,000 454-technology cDNA reads are also available but stored in the trace archives but not where they belong (short read archive SRA) but rather in the Petromyzon_marinus_OTHER database. | ||
| Line 45: | Line 287: | ||
GenBank opsin data for Geotria and Lethenteron can be mapped into Petromyzon orthologs using its traces to obtain a supplemental set of opsin sequences parsed for intron breaks and phases. Opsin classes not available from these other lamprey can be queried using chondrichthyes counterparts, with most genes only recoverable as fragments because of gaps in coverage (rather than divergence). In all cases, intron placement remains perfectly conserved from lamprey to mammal, indeed to amphioxus with some complications. | GenBank opsin data for Geotria and Lethenteron can be mapped into Petromyzon orthologs using its traces to obtain a supplemental set of opsin sequences parsed for intron breaks and phases. Opsin classes not available from these other lamprey can be queried using chondrichthyes counterparts, with most genes only recoverable as fragments because of gaps in coverage (rather than divergence). In all cases, intron placement remains perfectly conserved from lamprey to mammal, indeed to amphioxus with some complications. | ||
In evolutionary accounts, imaging opsins are often claimed to have appeared abruptly in lamprey as a complete color vision set orthologous to fish and amniotes (actually far exceeding the more restricted mammalian set). It is fair to say that the lamprey opsin complement establishes that the ancestral node animal had full-blown color vision and | In evolutionary accounts, imaging opsins are often claimed to have appeared abruptly in lamprey as a complete color vision set orthologous to fish and amniotes (actually far exceeding the more restricted mammalian set). It is fair to say that the lamprey opsin complement establishes that the ancestral node animal had full-blown color vision and auxiliary photoreception (ie the gene family had finished expanding on its way to seriously contracting in mammalian clades). | ||
Had one key species of lamprey happened to have gone extinct, namely Geotria, this conclusion would be seriously weakened because RHO2 orthologs are absent to date from the Petromyzon and other lamprey; SWS1 could almost be added to this list as it is known outside of Geotria only as a recent and rapidly disappearing pseudogene, an ironic parallel to later loss of SWS1 in monotremes. Additionally, SWS2 has a much older pseudogene in Petromyzon as discussed below; its loss may have predated the divergence from Lethenteron. | Had one key species of lamprey happened to have gone extinct, namely Geotria, this conclusion would be seriously weakened because RHO2 orthologs are absent to date from the Petromyzon and other lamprey; SWS1 could almost be added to this list as it is known outside of Geotria only as a recent and rapidly disappearing pseudogene, an ironic parallel to later loss of SWS1 in monotremes. Additionally, SWS2 has a much older pseudogene in Petromyzon as discussed below; its loss may have predated the divergence from Lethenteron. | ||
| Line 63: | Line 305: | ||
Energy-intensive processes such as swimming muscles and imaging vision (with its intensive consumption of ATP via cyclic GMP, nerve conduction, and CNS processing) may have first become feasible at some threshold of oxygen partial pressure. While accompanying caloric intake must sustain the energy drawdown, that is readily provided without predation (eg cows). We don't have an energy budget for photoreception in non-cone species such as Branchiostoma to assess whether its 7 opsins use more or less ATP than 3 retinal opsins in mammals. We don't have an energy budget for jellyfish bell-flexing swimming to compare cnidarian oxygen consumption with myotome swimming -- recall the predation direction here is often cnidarian-on-vertebrate. | Energy-intensive processes such as swimming muscles and imaging vision (with its intensive consumption of ATP via cyclic GMP, nerve conduction, and CNS processing) may have first become feasible at some threshold of oxygen partial pressure. While accompanying caloric intake must sustain the energy drawdown, that is readily provided without predation (eg cows). We don't have an energy budget for photoreception in non-cone species such as Branchiostoma to assess whether its 7 opsins use more or less ATP than 3 retinal opsins in mammals. We don't have an energy budget for jellyfish bell-flexing swimming to compare cnidarian oxygen consumption with myotome swimming -- recall the predation direction here is often cnidarian-on-vertebrate. | ||
A fairly recent SWS1 pseudogene is readily locatable in Petromyzon maritimus. Internal stop codons and frameshifts are not | A fairly recent SWS1 pseudogene is readily locatable in Petromyzon maritimus. Internal stop codons and frameshifts are not artifacts of assembly because each has multiple supporting raw trace reads but none supporting a functional reading frame. A not wholly satisfactory EST sequence EB722598 suggests, not unusually, that transcription of the pseudogene still occurs. Using Geotria as template, the coding sequence can be corrected from fragments to give a better idea of what Petromyzon SWS1 looked like prior to loss of function, though recovery of the spectrum seems like a dubious proposition. | ||
<pre> | <pre> | ||
>SWS1_petMar Petromyzon marinus (lamprey) pseudogene, sequence from genome: 7 frameshifts (^) and 2 stop codons | >SWS1_petMar Petromyzon marinus (lamprey) pseudogene, sequence from genome: 7 frameshifts (^) and 2 stop codons | ||
| Line 80: | Line 322: | ||
</pre> | </pre> | ||
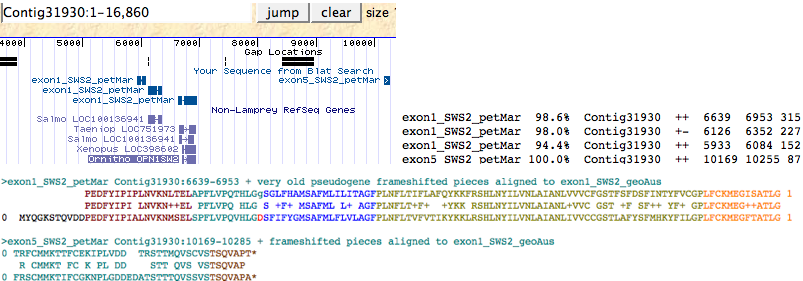
A very old SWS2 pseudogene can still be detected in Petromyzon maritimus either by keyword search in the lamprey genome browser | A very old SWS2 pseudogene can still be detected in Petromyzon maritimus either by keyword search in the lamprey genome browser or by trace archive blastn using SWS2_geoAus dna as query, in both methods verified by blastx against reference opsin sequences. Only exon 1 and exon 5 are still detectable; these lie in the same 16 kbp Contig 31930 and still retain their splice donor and acceptor locations and phases (indicating this is not processed pseudogene debris). Two further pieces of adjacent tandem debris related to exon 1 also occur in this contig. These exon fragments, when assembled onto a SWS2_geoAus template and corrected for frameshifts below, have greatly higher blastp matches to SWS2 sequences from a variety of species than to any other opsin class. | ||
[[Image:SWS2petMarps.png|left|]]<br clear="ALL"> | [[Image:SWS2petMarps.png|left|]]<br clear="ALL"> | ||
<pre> | <pre> | ||
>exon1:5_SWS2_petMar Petromyzon marinus (lamprey)Contig31930:6639-6953 + very old pseudogene frameshifted assembly | >exon1:5_SWS2_petMar Petromyzon marinus (lamprey) Contig31930:6639-6953 + very old pseudogene frameshifted assembly | ||
0 PEDFYIPIPLNVKNLTELAPFLVPQTHLGgSGLFHAMSAFMLILITAGFPLNFLTIFLAFQYKKFRSHLNYILVNLAIANLVVVCFGSTFSFDSFINTYFVCGPLFCKMEGISATLG 1 | 0 PEDFYIPIPLNVKNLTELAPFLVPQTHLGgSGLFHAMSAFMLILITAGFPLNFLTIFLAFQYKKFRSHLNYILVNLAIANLVVVCFGSTFSFDSFINTYFVCGPLFCKMEGISATLG 1 | ||
0 TRFCMMKTTFCEKIPLVDD TRSTTMQVSCVSTSQVAPT* | 0 TRFCMMKTTFCEKIPLVDD TRSTTMQVSCVSTSQVAPT* | ||
| Line 107: | Line 349: | ||
RHO1_geoAus Geotria australis full P497 AY366493 PMed:17463225 | RHO1_geoAus Geotria australis full P497 AY366493 PMed:17463225 | ||
RHO1_letJap Lethenteron japonicum full P--- AB116382 PMed:15096614 | RHO1_letJap Lethenteron japonicum full P--- AB116382 PMed:15096614 | ||
RHO1_petMar Petromyzon marinus full | RHO1_petMar Petromyzon marinus full P501 AB116382 PMed: | ||
RHO2_geoAus Geotria australis full P492 AY366494 PMed:17463225 | RHO2_geoAus Geotria australis full P492 AY366494 PMed:17463225 | ||
SWS2_geoAus Geotria australis full P439 AY366492 PMed:17463225 | SWS2_geoAus Geotria australis full P439 AY366492 PMed:17463225 | ||
| Line 115: | Line 357: | ||
LWS_geoAus Geotria australis full P560 AY366491 PMed:17463225 | LWS_geoAus Geotria australis full P560 AY366491 PMed:17463225 | ||
LWS_letJap Lethenteron japonicum full P--- AB116381 PMed:15096614 | LWS_letJap Lethenteron japonicum full P--- AB116381 PMed:15096614 | ||
LWS_petMar Petromyzon maritimus full | LWS_petMar Petromyzon maritimus full P536 genome PMed:no_ref | ||
PPIN_petMar Petromyzon maritimus full P370 genome PMed:15096614 | PPIN_petMar Petromyzon maritimus full P370 genome PMed:15096614 | ||
| Line 127: | Line 369: | ||
NEUR_petMar Petromyzon marinus frag1:6 P--- genome PMed:no_ref | NEUR_petMar Petromyzon marinus frag1:6 P--- genome PMed:no_ref | ||
=== <span style="color: #990099;">Chordate: Eptatretus burgeri (hagfish) .. 0 opsins</span> === | |||
Hagfish, after [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=18499072,19402130,19267641,18026166,9866205,12927130,17276090,16641994,11820840,17301331,1129982,17377535,15288047,7566650,17051155 decades of back-and-forth], are often sistered with lamprey, news not accepted yet by the [http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Tree&id=7711&lvl=3&p=nuccore taxonomy division] of GenBank nor [http://www.springerlink.com/content/n4t036300571k8j4/fulltext.html numerous researchers] and [http://tolweb.org/Hyperotreti paleontologists]. Given that common phylogenetic algorithms already grossly misplace mouse within the mammalian tree, we cannot expect nuclear coding genes to perform better at 500 myr back (five times the distance). Mitochondrial genome analysis has a similar history, though hagfish was found basal to lamprey + jawed vertebrates according to a [http://www.ncbi.nlm.nih.gov/pubmed/11820840? 2001 analysis]. | |||
Branchiostoma could serve as an outgroup (provided the first assembly is not used) but including radically evolving tunicates would skew results. The unfortunate lamprey assembly itself raise serious obstacles to determining topology. At this distance 28S, 18S rRNA and mitochondrial 16S rRNA may work better yet secondary and tertiary co-evolutionary considerations have not yet been incorporated. Hagfish experienced an extra round of HOX gene expansion, undercutting both HOX cluster copy number as a hallmark of supposed vertebrate body plan innovation and relentless speculation on 1R and 2R whole genome duplication in the vertebrate lineage. | |||
Smallest lamprey genome: 1.29 Mbp, Lampetra fluviatilis | |||
Largest lamprey genome: 2.50 Mbp, Petromyzon spp. | |||
Smallest hagfish genome: 1.29 Mbp, Eptatretus cirrhatus | |||
Largest hagfish genome: 4.59 Mbp, Myxine garmani | |||
None of the 660 nucleotide entries at GenBank as of May 2009 pertain to any component of vision, frustrating given that hagfish eye anatomy has been studied since 1886 (Krause, Die Retina der Fische. Cyclostomata. Internationale Monatschrift fur Anatomie und Histologie v3 p8-21) and [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=AbstractPlus&list_uids=6241039 opsin antibody labeling] was demonstrated in Myxine glutinosa in 1984 by Vigh-Teichmann. | |||
Jawless fish first appear 358 myr back in the [http://www.sciencemag.org/cgi/content/abstract/254/5032/701 Late Devonian fossil record]]. Hagfish and lamprey split well after the Cambrian, roughly 430 myr ago according to much-questioned molecular clocks. That's a time span comparable to divergence of human from shark. The oldest fossil hagfishes are Late Carboniferous (330 myr). The two surviving hagfish groups split some 75 myr ago (similar to human/mouse). Only recently has it been possible to obtain hagfish eggs and embryos and revisit the neural crest issue (establishing the conventional vertebrate series of events). | |||
Hagfish are nocturnal in aquaria and deep-sea in their natural habitat -- a [http://www.biolbull.org/cgi/content/full/212/1/55?view=long&pmid=17301331 new basal Eptatretus species] was recently captured at a deep hydrothermal vent. Such habits do not suggest that the ancestral imaging opsin portfolio will be retained (even as recognizable pseudogenes), yet hagfish still have circadian rhythm (based in the preoptic nucleus) and dermal photoreceptors (despite no pineal gland) as well as eyes. | |||
[[Image:HagfishEye.jpg|left]] | |||
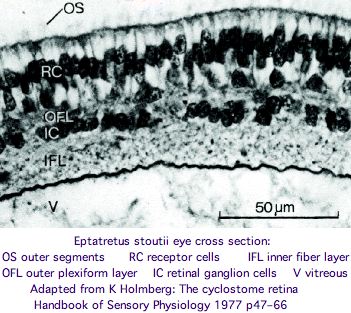
These non-imaging but paired eyes lack cornea, lens, vitreous body, and extrinsic eye muscle but nonetheless the retina and optic nerve react with RHO1 antibody. The eyes are larger in Eptatretus than in Myxine, where they are partly covered by the trunk musculature. However 1.3 mm is still quite small for an eye. | |||
After comparison of all extant genera, Fernholm and Holmberg concluded in 1975 that the hagfish eyes are secondarily degenerated from more conventional eyes adapted for shallow water (for example an early lens placode disappears). However bipolar neurons are never present and photoreceptors connect directly to ganglion cells, calling homologization to protostome wiring [http://www.ncbi.nlm.nih.gov/pubmed/19081043? into question]. The comparative anatomy of hagfish eyes has an excellent 1977 review in The Biology of Hagfishes (JM Jorgensen), pages 542-555 available by google book search. | |||
However this view is [http://www.springerlink.com/content/n4t036300571k8j4/fulltext.html disputed] in a wide-ranging 2008 proposal for imaging eye evolution: | |||
"It is often suggested that the hagfish eye is degenerate, having regressed from a more lamprey-like eye that existed in the common ancestor of hagfish and lampreys. We find this view implausible. Given that the hagfish eye has survived for hundreds of millions of years, in comparison with the degeneration that has occurred in just thousands of years in cavefish. The hagfish eye must have had considerable survival value. Degeneration cannot explain how it was advantageous to hagfish for: (a) a three-layered retina to revert to two-layered; (b) the processing power of the bipolar cells to be lost; (c) reversion to a more rudimentary photoreceptor structure; (d) the disappearance of the lens, cornea, iris, and ocular muscles, all without trace; (e) re-covering of the eye by skin; and (f) re-projection of the retinal ganglion cells from tectum to the hypothalamus. It seems more parsimonious that the hagfish eye would simply have been lost..." | |||
These authors argue further that the hagfish eye is more like a pineal gland (which hagfish lack but lamprey have) and functions in circadian rhythm rather than vision. Here we might wonder why bilateral eyes are needed with pigmented epithelial backstop giving forward and side directional capability, whereas mere light photoperiod detection would be more sensitive without it. At a minimum, paired hagfish eyes determine orientation with respect to light source (sky) and enable consistency of forward motion. | |||
No hagfish opsins have been sequenced; no genome project is scheduled (even as the price drops in 2009 to $3,000). Even if hagfish imaging opsins are mostly gone, residual ciliary and rhabdomeric opsins could be quite informative. If so, hagfish have information about a critical stage in vertebrate imaging opsin evolution. | |||
How will we interpret hagfish opsins as sequence data becomes available? That data ideally will come from multiple hagfish genera because gene loss can take different patterns in individual species causing the ancestral state to be underestimated. The critical issue is how eye opsins classify (root among ciliary opsins) -- that's just a quick blastp at the opsin classifier. Hagfish may well use a melanopsin in the preoptic nucleus for circadian rhythm but cannot plausibly utilize rhadomeric opsins in the eye as do protostomes. | |||
* If hagfish diverged very early, only earlier opsins classifying to PIN, VAOP or PPIN (the likeliest of these to localize to the eye) will be found. While still ciliary in terms of their signaling transduction cascade, these are perhaps more likely in the dermal photoreceptors, which highlights the need for in situ hybridization should the data come from whole genome determination or whole animal transcripts. If confirmed as primary photoreceptor in hagfish eye, this would rule out Cyclostoma since <font color="#990099">regression from LWS etc (cone opsins) to PPIN is an unacceptable scenario.</font> | |||
* If hagfish diverged early, LWS is most plausible among imaging opsins given its basal position in the sequential duplication cascade of cone opsins (already finalized prior to last common ancestor of lamprey and gnathostomes). However its lambda-max may be adaptively shifted by now to a shorter wavelength better suited to dim light at depth. This outcome does not favor sistering with lamprey but does not quite rule it out either (other cone opsins could be lost). It does however require appropriately localized genes of the cis-retinal regenerative cycle such as RPE65. | |||
* If hagfish diverged late and experienced opsin gene loss after a larger complement of opsins had evolved in a common stem ancestor, then any of LWS, SWS2, SWS1, RHO2 or RHO1 could still be present. As an example, suppose only a single opsin is found, classifying as SWS1. This implies, given the gene history, a 'lower bound' of LWS and SWS2 (ie trichromaticity at some point) having been lost but says nothing about whether RHO2 or RHO1 had ever been present in a hagfish ancestor. | |||
* If hagfish diverged very late, the residual opsin repertoire could be either large or mostly lost. The three species of lamprey studied establish a full complement of opsins at agnathan ancestor with gnathans. These opsins include 5 imaging opsins, parietopsin, pinopsins VAOP, encephalopsin, melanopsin, and neuropsins. Lamprey TMT, RGR and peropsin appear to have been lost but may merely be too diverged to recognize. The surviving opsins in hagfish then provide clues to what opsins were lost. | |||
It's worth reviewing the opsin classes found in the two earlier diverging clades Ciona and Branchiostoma and their tissue localization. Between extensive genome, transcript projects and expression experiments, we can be sure of having complete opsin sets and expression assignments. | |||
* Tunicate opsins (below) include an advanced ciliary opsin classifying to PPIN expressed appropriately in its ocellus and containing the terminal [[Opsin_evolution:_Cytoplasmic_face#Melanopsins|VAPA* cilium targeting sequence]]. That implies hagfish at one point in its history had a similar opsin classifying to PPIN VAOP PIN. A second PPIN, two RGR and a highly diverged melanopsin complete the repertoire of both Ciona intestinalis and the distantly related 'congeneric' Ciona savignyi. Here all GPCR containing a lysine at Schiff base homologous position (a K-rhodopsin) have been evaluated as potential photoreceptors. Ciona has not retained the neuropsin and peropsin opsin classes. | |||
* Branchiostoma has only basal ciliary opsins classifying to the TMT/ENCEPH group (in addition to melanopsins, neuropsins and peropsins). Its highly diverged encephalopsin lacks the fixed conserved length and VAPA targeting sequence also [[Opsin_evolution:_Cytoplasmic_face#Melanopsins|characteristic of the C-terminus of encephalopsins]] (at least from Callorhinchus on; lamprey coverage is incomplete here). While it is implausible that a PPIN opsin could have been overlooked given the extensive trace coverage reflected in the second assembly and the intense experimental targeting of opsins in B. belcheri, other cephalochordates should also be sequenced. | |||
*While this observation could reflect loss of PPIN in amphioxus, that implies <font color="#990099">regressive replacement of PPIN by ENCEPH in the eye</font>, far more complicated genetically than simply losing a cone opsin or two (eg Petromyzon vs Geotria). Thus the overall opsin picture strongly favors the phylogenetic tree with urochordates closer to lamprey than cephalochordates. We may be better off with such considerations because published [http://www.ncbi.nlm.nih.gov/pubmed/18563158? large-scale molecular studies] to date have not been definitive for chordate tree topology: | |||
* the sea urchin and hemichordate genomes are too diverged to work as outgroups, drosophila is too far back and its proteome too derived | |||
* Ciona assemblies have gotten worse, the KH assembly must be used | |||
* the half-baked Oikopleura genome project indicates a very rapidly evolving and immensely diverged species | |||
* the first Branchiostoma assembly had [http://www.ncbi.nlm.nih.gov/pubmed/19379521? mediocre gene models], the second remains unannotated | |||
* lamprey genome is a mess due to retroposons, poor exon coverage, and lack of gene models | |||
* chondrichthyes genome is not far enough along | |||
* teleost fish genomes are highly diverged, unfinished, and confounded by lineage-specific duplication | |||
* orthologs are difficult to establish without synteny, complete genomes, and extensive prior gene family annotation | |||
* mutational models used in ML are extremely dubious over this time scale and diversity of population genetics | |||
* immense long branches have been used throughout, a risky strategy | |||
[[Image: | Hagfish auxiliary genes are an important further consideration because ciliary opsins appear incapable of replenishing themselves, unlike rhabdomeric opsins which can regenerate cis-retinals in situ. RPE65, RBP3 (IRBP), and Galpha signaling factors are three interesting gene families to consider here. The first should localize to retinal pigmented epithelia, the second to interstitial matrix. and the third should cluster with vertebrate cone transducins. | ||
* [[Opsin_evolution:_RBP3_(IRBP)|IRBP]] presents a timing opportunity because of the rare genomic event raising the domain count from one in Branchiostoma to four in lamprey (and all gnathans) with a concomitant radical change in exon structure and probable improvement in retinal shuttling. Unfortunately the gene seems lost in those tunicates for which there is data. No expression data is available for cephalochordate IRBP. No data is available for hagfish but if IRBP here proved structured like amphioxus rather than lamprey, that would unequivocally resolve hagfish position on the phylogenetic tree. | |||
* [[Opsin_evolution:_RPE65|RPE65]] is more favorable in terms of data availability but is complicated by two additional ancestral paralogs and lacks the 'smoking gun' of IRBP non-convergent evolution. No data is available for hagfish. The amphioxus genome shows an inconvenient expansion of this gene family, not all of which can be attributed to high-polymorphism assembly artifacts. Lamprey genome is woefully incomplete in its coverage. Ciona has two paralogs with the expected intronation. | |||
* [[Opsin_evolution:_transducins|Only GNAI]] sequence data (from leukocytes) is available for hagfish as of May 2009. Here hagfish eye needs to be specifically screened for imaging transducins as absence of evidence is hardly evidence of absence. This inhibitory Galpha is unsuitable for imaging opsin transduction but [http://www.ncbi.nlm.nih.gov/pubmed/18832159? might work for parapinopsin]; it clusters equally among vertebrate GNAi1/i2/i3 (analogously to its [http://www.ncbi.nlm.nih.gov/pubmed/15236930? GATA3-like and BTK-like genes]) whereas lamprey has the contemporary complement of [http://www.ncbi.nlm.nih.gov/pubmed/18687354? clearly separated GNAT1 and GNAT2]. Such data has been cited for [http://www.ncbi.nlm.nih.gov/pubmed/19267631? wholesale gene duplication] between hagfish and lamprey; observe the [[Opsin_evolution:_transducins#Curious_evolutionary_origins_of_alpha_subunit_multiplicity|Galpha gene tree]] is, like that for imaging opsins, inconsistent with whole genome duplication and a better fit to simple tandem gene expansions during a very rapid adaptive era in eye evolution. | |||
=== <span style="color: #990099;">Urochordata: Ciona intestinalis (tunicate) .. 6 opsins</span> === | |||
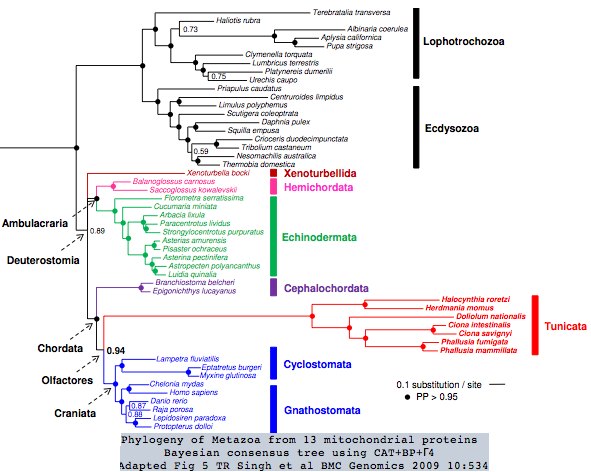
Tunicates occupy a strategic urochordate position in the phylogenetic tree as [http://en.wikipedia.org/wiki/Urochordate immediate outgroup] to vertebrates. Three tunicate genomes have been sequenced but these proved disappointing for comparative genomics due to their derived nature, which adversely impacts coding sequence divergence, gain and loss of genes, overwriting of ancestral introns, almost total loss of gene order, and high positional heterozygosity. The situation in the seven sequenced tunicate mitochondrial genomes is also [http://www.biomedcentral.com/1471-2164/10/534 extreme]. | |||
[[Image:MetTree.jpg|left]] | |||
It may not be possible to find more conservatively evolving tunicates if rapid generation time and free spawning are characteristics of all extant urochordates. Yet in other aspects, such as reconstructing the evolutionary trajectory of the vertebrate eye, contemporary tunicates may have retained critical information. | |||
The most useful of these rogue [http://genomebiology.com/2008/9/10/R152 genomes] to is Ciona intestinalis -- Ciona savignyi and Oikopleura dioica have meagre transcript data and see low annual use as model organisms. PubMed abstracts mention these species in the publication ratio 886:86:46 overall and 62:8:7 for calendar year 2008. | |||
Halocynthia roretzi has many cdna but has not been evaluated for genomic characters whereas Ciona intestinalis has been developed extensively as an experimental system; its massive cdna coverage allow better contig joining and recovery of complete coding gene models which would be nearly impossible (because of divergence and intron gain/loss) from mere homological alignment to other deuterostome proteins. | |||
The first two assemblies of Ciona intestinalis were highly defective, having some 5,109 missing genes, faux duplications and truncated gene models (a third of the 15,254 total) with the second JGI assembly ironically [http://genomebiology.com/2008/9/10/R152 worse than the first.] This raises serious questions about recent papers in comparative genomics that relied on highly defective early gene sets, notably those revising urochordate taxonomic position and comparing rates of protein evolution to Branchiostoma (whose initial assembly also required serious revision). | |||
Note however that the new Ciona assembly KH is available on 7 Nov 08 only as raw download (though a [http://hoya.zool.kyoto-u.ac.jp/blast_kh.html blast server] with username = guest, password = welcome is available) and that the June 2008 [http://www.nature.com/nature/journal/v453/n7198/full/nature06967.html Branchiostoma genome paper] refers to v1.0, whereas the much-revised release 2.0 became available at its [http://genome.jgi-psf.org/cgi-bin/runAlignment?db=Brafl1&advanced=1 public blast site] in Nov 2008 (gene models are not yet recalculated but simply lifted from 1.0). Obviously Ciona KH and Branchiostoma 2.0 are what need to be compared (and to a better lamprey assembly which is not even underway). | |||
A complete set of Ciona opsin genes cannot necessarily be recovered even from the KH assembly because the article notes "it is still possible that a minor fraction of genes, such as genes expressed only under particular environmental conditions, are not covered by these ESTs. A fraction of previous models not supported by paired ESTs were excluded from the KH model set. A part of them may be real genes or unannotated fragments of genes represented by the KH models, because the encoded protein shows sequence similarity to proteins known in other species (approximately 1,641 loci with <1E-05 blast hits in the human proteome), These are provided as a supplemental model set (see Materials and methods) along with other unsupported or incompletely supported models.... probable that a minority of additional genes reside within gaps in the current assembly (48 EST-supported loci)... 47,511 ESTs (4%) were not mapped anywhere in the KH assembly... Moreover, we estimate that at least 84% of the KH transcript models contain the complete protein-coding ORF..." | |||
Fortunately, both larval and adult photoreception have been [http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Retrieve&db=pubmed&dopt=Abstract&list_uids=10483365,1918443,11591373,12687683,12373590,12875912,12392543,16118796,14611940,17245701,16169492,15671331,15176085,16930093 thoroughly studied.] Ciona lacks imaging eyes and thus any counterparts to rod and cone opsins (as with the cephalochordate Branchiostoma). The relative topology of these two with respect to the vertebrates has tilted in recent years towards amphioxus as immediate outgroup but the lack anything beyond encephalopsins in Branchiostoma yet a parapinopsin in tunicates argues strongly against this. | |||
[[Image:Opsins_cii_larval_eye.png|left|]] | |||
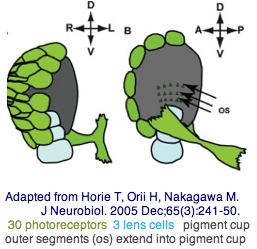
The tadpole larva CNS contains 335 cells of 13 types. These include 30 retinal photoreceptor cells in an unpaired ocellus and 5 accessory cells -- 3 for a ocellus lens-like structure, 1 for the pigment cup, and 1 pigment cell in the otolith (inconsistently with a hydrostatic sensing role for its 19 receptors). The pigment cells of the ocellus and otolith form an equipotent developmental equivalence group -- a bilateral pair of cells in the blastula gives rise to the otolith and ocellus melanocytes whereas the retina arises from both left and right cell lineages. The observed genomic complement of opsins may largely come into play in the larva because the adult is sessile with little resemblance to vertebrates. The larva are non-feeding which scarcely fits a super-predator role for early deuterostomes opsins. | |||
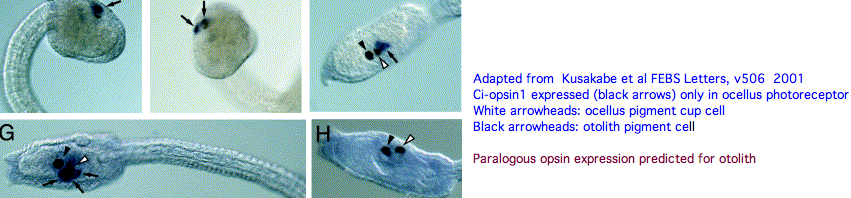
An evidently ciliary opsin called Ci-opsin1 is expressed in the larval ocellus (stored here as PPINa_cioInt). The opsin | An evidently ciliary opsin called Ci-opsin1 is expressed in the larval ocellus (stored here as PPINa_cioInt). The opsin classifier places this in the PPIN/PIN/VAOP group with best match 44% identity, quite respectable given a billion years of roundtrip evolutionary time. As noted initially by Kusakabe et al in 2001, this opsin shares 3 identical introns with the vertebrate group. | ||
Today there are 25x as many opsin sequences available with much greater phylogenetic dispersion. It appears Ci-opsin has 2 new intron insertions relative to the ancestral Gq ciliary opsin 4-intron pattern 0.2.0.0. This pattern is specific (not shared by Gt melanopsins nor Go retinal isomerases) and diagnostic (disregarding a few lineage-specific gains and losses) -- see [ | Today there are 25x as many opsin sequences available with much greater phylogenetic dispersion. It appears Ci-opsin has 2 new intron insertions relative to the ancestral Gq ciliary opsin 4-intron pattern 0.2.0.0. This pattern is specific (not shared by Gt melanopsins nor Go retinal isomerases) and diagnostic (disregarding a few lineage-specific gains and losses) -- see [[Opsin_evolution:_annotation_tricks|documentation]]. | ||
Three ancient ciliary introns were already established at the time of amphioxus and tunicate encephalopsin divergences. Indeed they already occur in sea urchin, ragworm, mosquito, moth, and beetle ciliary opsins. Consequently they were present in the parent ciliary opsin of | Three ancient ciliary introns were already established at the time of amphioxus and tunicate encephalopsin divergences. Indeed they already occur in sea urchin, ragworm, mosquito, moth, and beetle ciliary opsins. Consequently they were present in the parent ciliary opsin of Ur-bilatera and no doubt Cnidaria. There's nothing surprising about this because the vast majority of (human) coding introns originated far earlier in unicellular eukaryotes and have been conserved ever since. Outside of rogue lineages such as drosophila, nematode, and tunicates, event rates for intron gain and loss are perhaps 1-2 per five billion years of branch length. Convergence is not favored because 333 aa sites x 3 intron phases = 1000 distinct possibilities in an opsin-sized protein -- for an already very rare event to happen twice in the available branch length requires predisposing factors. | ||
We will use these deep intron characters later to supplement -- and even trump -- maximal likelihood inference from primary sequence divergence which captures the broad picture but fails to resolve the issues of most interest. With opsins, alignment (at these time depths and rates of change) hits the wall of generic rhodopsin superfamily and indeed generic GPCR proteins, which numbered many hundreds at the time of | We will use these deep intron characters later to supplement -- and even trump -- maximal likelihood inference from primary sequence divergence which captures the broad picture but fails to resolve the issues of most interest. With opsins, alignment (at these time depths and rates of change) hits the wall of generic rhodopsin superfamily and indeed generic GPCR proteins, which numbered many hundreds at the time of Ur-bilatera. There are already many constraints on proteins which must have seven transmembrane helical segments, covalently bind retinal with a lysine and counterion, and interact with heterotrimeric signaling protein. | ||
With the genome in hand, we can see Ci-opsin1 has an unstudied paralog (here called PPINb_cioInt) of 58% identity and identical introns (other than a new phase 21 intron breaking exon 4). There is no expression data for the paralog in the [http://genome.cse.ucsc.edu/ UCSC browser track] but it cannot plausibly be a pseudogene due to the conserved nature of amino acid replacements, so we wonder about subfunctionalization. <span style="color: #990099;">The hybridization experiment will have to be repeated</span> at various life stages. Paralog lambda max might be computable or measurable in a construct. The 1999 experiment (which measured speed-up in swimming after light decreased, reminiscent of the pineal-mediated frog tadpole response) deduced a lambda max of 505 nm -- perhaps that was a composite action spectrum. The new paralog in fact conserves the key lysine and counterion. | With the genome in hand, we can see Ci-opsin1 has an unstudied paralog (here called PPINb_cioInt) of 58% identity and identical introns (other than a new phase 21 intron breaking exon 4). There is no expression data for the paralog in the [http://genome.cse.ucsc.edu/ UCSC browser track] but it cannot plausibly be a pseudogene due to the conserved nature of amino acid replacements, so we wonder about subfunctionalization. <span style="color: #990099;">The hybridization experiment will have to be repeated</span> at various life stages. Paralog lambda max might be computable or measurable in a construct. The 1999 experiment (which measured speed-up in swimming after light decreased, reminiscent of the pineal-mediated frog tadpole response) deduced a lambda max of 505 nm -- perhaps that was a composite action spectrum. The new paralog in fact conserves the key lysine and counterion. | ||
| Line 174: | Line 480: | ||
<br clear="all" /> | <br clear="all" /> | ||
Possibly this ciliary paralog pair descended from a gene duplication already present in the last common | Possibly this ciliary paralog pair descended from a gene duplication already present in the last common ancestor, leading after still more gene duplications to the current portfolio of vertebrate ciliary opsins. This would account for its ambivalent behavior in the Opsin Classifier with respect to the PPIN/PIN/VAOP group. Alternately the pair might represent a tunicate-specific duplication of secondary interest. Ciona savignyi has a clear ortholog (88% identity) to PPINa_cioInt but a lesser match at 59% to PPINb_cioInt, in both cases with identical introns (not an unusual pattern in gene duplications assuming PPINa_cioInt continues the original function). C. savignyi -- which is only in the same genus from a severe anthropocentric perspective -- helps gauge the rate of evolution of C. intestinalis opsins. | ||
Photoreception in the adult ascidian, which might seem gratuitous in a sessile filter-feeder, has not been studied in quite such detail. However several non-opsin expression studies suggest that adult photoreceptors may develop about pigmented spots around oral and atrial siphons, epithelial cells of sperm duct and cerebral | Photoreception in the adult ascidian, which might seem gratuitous in a sessile filter-feeder, has not been studied in quite such detail. However several non-opsin expression studies suggest that adult photoreceptors may develop about pigmented spots around oral and atrial siphons, epithelial cells of sperm duct and cerebral ganglia, involving behaviors such as siphon contraction, phototropism, and gamete release. The anterior photoreceptor of the oral siphon has even been [http://www.sci.u-hyogo.ac.jp/life/bioinfo/kusakabe/research.html homologized to vertebrate lateral eyes.] | ||
We'll see below that exactly the same problem as above (undocumented paralog) may affect interpretation of a comprehensive experimental study of Ciona Ci-opsin3 (RGR1_cioInt at the Opsin Classifier). Here too I was able to recover a related second gene in both C. intestinalis and C. savignyi. This illustrates the power of genomics -- provided coverage is complete, a full complement of bioinformatically extracted opsins can guide experimental design from the beginning. A full set of opsin classes should be sought in the genome, even if their degree of sequence divergence and lack of transcripts makes this difficult. | We'll see below that exactly the same problem as above (undocumented paralog) may affect interpretation of a comprehensive experimental study of Ciona Ci-opsin3 (RGR1_cioInt at the Opsin Classifier). Here too I was able to recover a related second gene in both C. intestinalis and C. savignyi. This illustrates the power of genomics -- provided coverage is complete, a full complement of bioinformatically extracted opsins can guide experimental design from the beginning. A full set of opsin classes should be sought in the genome, even if their degree of sequence divergence and lack of transcripts makes this difficult. | ||
| Line 186: | Line 492: | ||
[[Image:RhodCiona.jpg|center|]] | [[Image:RhodCiona.jpg|center|]] | ||
It turns out (using closest homologs) that Ci-BCO is predominantly expressed in larval ocellus photoreceptor cells, whereas Ci-RPE65 is not significantly expressed there nor in larval brain vesicle but rather in photoreceptor cells of the neural complex (a photoreceptor organ of the adult) right along with Ci-opsin3 and Ci-CRALBP (ie, like cephalopod). It appears the larval visual cycle uses Ci-opsin3 as restorative photoisomerase whereas the adult visual cycle Ci-RPE65. The remote paralog RGR2_cioInt was not studied and its role remains speculative. Given its degree of divergence yet | It turns out (using closest homologs) that Ci-BCO is predominantly expressed in larval ocellus photoreceptor cells, whereas Ci-RPE65 is not significantly expressed there nor in larval brain vesicle but rather in photoreceptor cells of the neural complex (a photoreceptor organ of the adult) right along with Ci-opsin3 and Ci-CRALBP (ie, like cephalopod). It appears the larval visual cycle uses Ci-opsin3 as restorative photoisomerase whereas the adult visual cycle Ci-RPE65. The remote paralog RGR2_cioInt was not studied and its role remains speculative. Given its degree of divergence yet persistence in a second ascidian, it is an old gene duplication maintained somewhat by selective pressure. | ||
What about rhabdomeric opsins in Ciona? We know that melanopsin persisted into vertebrates so it must have been present at the common ancestor with ascidians. Rhabdomeres themselves as a subcellular opsin housing specialization did not persist so their apparent absence in Ciona does not imply the absence of melanopsin. | What about rhabdomeric opsins in Ciona? We know that melanopsin persisted into vertebrates so it must have been present at the common ancestor with ascidians. Rhabdomeres themselves as a subcellular opsin housing specialization did not persist so their apparent absence in Ciona does not imply the absence of melanopsin. | ||
A Ciona melanopsin could be very diverged. The best possible search involves tblastn of the [http://genome.jgi-psf.org/cgi-bin/runAlignment?db=Cioin2&advanced=1improved Ciona assembly] and GenBank est_others with a variety of queries (since the best query is not known in advance; after the fact it is provided by the Opsin | A Ciona melanopsin could be very diverged. The best possible search involves tblastn of the [http://genome.jgi-psf.org/cgi-bin/runAlignment?db=Cioin2&advanced=1improved Ciona assembly] and GenBank est_others with a variety of queries (since the best query is not known in advance; after the fact it is provided by the Opsin Classifier). Reconstructed ancestral melanopsins can improve on specific species queries by eliminated half of the roundtrip divergence. | ||
However overly sensitive queries have the risk of merely returning generic rhodopsin-superfamily members (notably ADRA1A adrenergic receptor). While these won't receive clean approval from the Opsin Classifier, any putative melanopsin must be secondarily validated by retention of intron pattern, synteny with vertebrate melanopsins (unlikely in Ciona), and internal amino acid signatures of authentic melanopsin-type photoreceptors. | However overly sensitive queries have the risk of merely returning generic rhodopsin-superfamily members (notably ADRA1A adrenergic receptor). While these won't receive clean approval from the Opsin Classifier, any putative melanopsin must be secondarily validated by retention of intron pattern, synteny with vertebrate melanopsins (unlikely in Ciona), and internal amino acid signatures of authentic melanopsin-type photoreceptors. | ||
A more powerful search technique evaluates K-rhodopsins, defined as any GPCR with lysine in Schiff base homologous position (ie the lysine continues past a NAxxY motif to the YR motif at the deeply invariant length of 19 residues). In May 2009, two GPCR classifying as melanopsins were recovered here from Ciona intestinalis and Ciona savignyi. Those sequences are still rough despite three available transcripts because of divergence and a scrambled assembly in this region. The transcripts originated in blood cells and juvenile whole animal according to their annotation, leaving their role in photoreception undemonstrated. | |||
Opsin sequences available from urochordates in Nov 2009. Renamed from original literature using the opsin classifier to place. | |||
<font color="magenta"> | |||
PPINa_cioInt PPINa_cioSav ciliary NM_001032555 Ci-opsin1 ERY KTATIYNPLIYIGLNRQFR | |||
PPINb_cioInt PPINb_cioSav ciliary XM_002119927 --- ERY KTANIYNPLIYIGLNKQFR</font><font color="blue"> | |||
MEL1_cioInt MEL1_cioSav melanopsin --- --- DRH KASCVHSSFAYITNAHFR</font><font color="brown"> | |||
RGRa_cioInt RGRa_cioSav other NM_001032468 Ci-opsin3 DKY KVFVGSNPFIYIYFDPELR | |||
RGRb1_cioInt --- other NM_001032464 CiNut DRY KVISVVNPYLYMRSDPELLA | |||
RGRb2_cioInt RGRb2_cioSav other XM_002121277 --- DRY KFISVMNPYMYMRSDPELLR</font> | |||
=== <span style="color: #990099;">Cephalochordata: Branchiostoma (amphioxus) .. 12 opsins</span> === | |||
The [http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=2034537 phylogenetic tree] of cephalochordates, based on mitochrondrial dna, shows 9 living species coalescing at [http://www.ncbi.nlm.nih.gov/pubmed/15883887 162 myr] whereas the last common ancestor with vertebrates is some 400 myr earlier, thus limiting our ultimate ability to ever accurately reconstruct this node. Cephalochordates species are anthropocentrically grouped into 'genera' at divergences that applied evenhandedly to placentals would require lumping them all into a [http://www.ncbi.nlm.nih.gov/pubmed/19411599 single genus]. | |||
[[Image:Ceph div.jpg|left]] | |||
Cephalochordate opsin data is entirely concentrated in detailed studies of [http://www.ncbi.nlm.nih.gov/pubmed/12435605,18513236,18088357,15936279,15851682,18563158 Branchiostoma belcheri] and bioinformatic efforts on the genomic species [http://www.ncbi.nlm.nih.gov/pubmed/18562680 Branchiostoma floridae]. The seven full-length B. belcheri opsin genes can be mapped into 'best-blast orthologs' in B. floridae where intron structure can be determined assuming GT-AG junctions, hopefully with accurate transfer of experimental annotation to these unstudied genes. B. floridae has 4 additional opsins whose counterparts in B. belcheri are unknown; the total of 12 opsins exceeds human by 4. | |||
Every study to date must be repeated using the full genomic complement of 12 opsins (rather than just the initial 7). For example, it [http://www.pnas.org/content/early/2009/05/15/0811476106.abstract cannot be concluded] when two melanopsin paralogs exist that rhabdomeric Joseph cells and pigmented dorsal ocelli (Hesse organ) photoreceptors must use one melanopsin but not the other, simply from rough action spectrum consistency. (The other melanopsin could not have endured 500 myr without contributing a function.) | |||
The [http://www.brain-map.org/ Allen Brain Atlas] of mouse has exposed the great fallacy of assuming photoreceptor genes are functionally expressed only in anatomically obvious cell types. Thus frontal eye and lamellar body (pineal) for the ciliary opsins and Joseph cells and Hesse ocelli for microvillary opsins may not be the entire photoreceptor repertoire. To understand photoreception in amphioxus, it will prove necessary to revisit all life stages of all amphioxus cell types with hybridization with all 12 opsins to see what else expresses them. | |||
=== <span style="color: #990099;">Echinodermata: | Four B. floridae opsins are ciliary (same count as human), three are rhabdomeric melanopsins, and the rest peropsins and neuropsins. Lancelet ciliary opsins all classify to the basal TMT/Enceph type; no counterparts to mid-opsins (PPIN, VAOP, PIN, PARIE) or cone/rod imaging opsins occur even allowing for immense divergence. Conceivably this is attributable to genome incompleteness or lineage-specific loss; far more likely, these specialized opsin classes never evolved in cepholochordates in view of photoreceptor anatomy and phylogenetic position. Observe Ciona has an opsin classifying to PPIN, consistent with later divergence of urochordates. | ||
The sea urchin genome carried a big surprise: the | <br clear="all"> | ||
The 12 cephalochordate opsins classified, with [[opsin_evolution:_transducins|signaling class]], synonymous terminology, and [[Opsin_evolution:_Cytoplasmic_face|C2-TM7 switch]]; full sequences [[Opsin_evolution#Opsin_Dataset|here]]. | |||
<font color="magenta">TMT5_braFlo Branchiostoma floridae Gt Amphiop5 ERY KSSTCYNPLVYFAMNNQFR | |||
TMTx_braFlo Branchiostoma floridae Gt ..new .. ERY KSSVVYNAAIYVAMNNQFR | |||
TMTy_braFlo Branchiostoma floridae Gt ..new .. SRY KTSYIVNTIIYLVMEKEFR | |||
ENCEPH4_braFlo Branchiostoma floridae Gt Amphiop4 ERY KSSTAYNPIIYVLMNNQFR</font> | |||
<font color="blue">MEL6_braFlo Branchiostoma floridae Gq Amphiop6 ERY KLSVLFNPVAYVLSIPSFR | |||
MELmop_braFlo Branchiostoma floridae Gq Amphi-mop DRY KSSAVYNPIVYAITHPKFR | |||
MELx_braFlo Branchiostoma floridae Gq ..new.. ERF KLSVLINPVAYVFSIPSFR</font> | |||
<font color="brown">NEUR1a_braFlo Branchiostoma floridae .. ..new .. MRF KSNSLWNPIIYLGMNERFR | |||
NEUR1b_braFlo Branchiostoma floridae .. ..new .. MRF KSSSLWNPIIYLGMNDRFS | |||
PER1_braFlo Branchiostoma floridae Go Amphiop1 YRY KSSCMMNPIIYSCCNGKFR | |||
PER2_braFlo Branchiostoma floridae Go Amphiop2 HRY KTHCAFNPILYMLMSEVYR | |||
PER3_braFlo Branchiostoma floridae Go Amphiop3 DRY KSSALYNPIIYIIANRRFR</font> | |||
[[Image:CephOpTree.jpg|left]] | |||
The bioinformatics of B. floridae is very difficult, first because the individual sequenced was highly polymorphic (not inbred). This caused misassembly on a vast scale, with thousands of genes duplicated either in tandem (due to mate pair logic) or to unanchored contigs. These extra copies -- really just alleles -- can differ by several percent at the amino acid level and more within corresponding introns (due to different retroposon and indel histories). It is very difficult to identify recent true tandem duplications in this environment. The second assembly discarded one copy arbitrarily (causing cDNAs to sometimes mismatch) whereas what is needed is a diploid assembly. | |||
Assembly 2.0 is available for [http://genomeportal.jgi-psf.org/cgi-bin/runAlignment?db=Brafl1&advanced=1 blast search] but its gene content had not been re-analyzed as of May 2009. The published gene count of 23,000 surely needs major downward revision. Synteny relations should still hold, though that was [http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pubmed&pubmedid=18562680 loosely defined] under twin dubious assumptions allowing unlimited 'local' inversions and admitting weak uncurated partial homology to establish orthology. Branchiostoma genes remain uncurated; nearly all pipeline gene models are wrong in significant details. However the above set of 12 probably reflects a complete set of all K-rhodopsins in this species. | |||
Considerable continuity can be seen to protostome invertebrates. The best ciliary opsin matches to lophotrochozoan and arthropod opsins have 38% identity, whereas melanopsins do better at 46%. To call these orthologs is not helpful (in the absence of synteny) when each clade has experienced lineage-specific gains and losses. | |||
The TMT sequences in Branchiostoma contain an extra phase 00 intron relative to exon 2 of vertebrate counterparts; the encephalopsin has additionally an extra phase 12 intron in exon 1. Neither of these introns is seen from lamprey on, nor do they occur in lophotrochozoan TMT (which have their own novel phase 12 intron in exon 2). It is parsimonious to take vertebrate intronation as ancestral and invoke two intron gain events in cephalochordate. | |||
None of the six available Branchiostoma ciliary opsins contain the VAPA* targeting signal. This may have evolved later -- it is first seen in the Ciona PPIN homolog and, in the gene tree, [[Opsin_evolution:_Cytoplasmic_face#Encephalopsin|encephalopsins]]. Note the specialized ciliary membranes housing opsins arose far earlier as it is seen already in the protostome Platynereis (indeed its opsins are close to the signal with VAAA* in TMT1_plaDum and VAAT* in TMT2_plaDum). | |||
Thus ciliary opsins in amphioxus seem not to have this specialization (in addition to great divergence at the sequence level). It's also unclear if any member of the expanded TMT/ENCEPH gene set has a special homology relation to the PPIN opsin in tunicates (their only ciliary opsin). Here the best match is amphioxus TMTx to Ciona PPIN but the percent identity is only 36%. Perhaps it is better to look backwards to echinoderm, yet no special affinities are observed there either (TMTPIN_stoPur and ENCEPH_strPur are its only two ciliary opsins). | |||
[[Image:BranchioJoeHes.jpg|left]] | |||
A May 2009 [[http://www.pnas.org/content/early/2009/05/15/0811476106.abstract PNAS article] provides refreshingly direct experimental evidence that the Joseph laminar body and Hesse ocelli function as photoreceptors. It provided possible to isolate individual cellular units and make observations on them. In conjunction with three previous cdna hybridization and ultrastructural [http://www.ncbi.nlm.nih.gov/pubmed/18513236,18088357,15936279 detailed studies], the evidence seems overwhelming. However that seemed the case to comparative anatomists already in 1904. The issue today: if MELmop_braFlo (Amphi-mop) provides the primary photoreceptive opsin in both organs, what then is the role of the other melanopsic opsin MEL6_braFlo (Amphiop6)? | |||
<br clear="all"> | |||
(to be continued) | |||
=== <span style="color: #990099;">Echinodermata: Strongylocentrotus purpuratus (sea urchin) .. 7 opsins</span> === | |||
The sea urchin genome carried a big surprise: the long-belittled echinoderm has a large set of genes for sensory and signaling capability (comparable in number to human). These include [http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=16965768,17095691,17067569 seven opsins] considered here. | |||
Adult sea urchins exhibit a variety of responses to light intensity: predator avoidance, shelter seeking, covering reactions, diurnal migrations, testis response to light, and spine defense reaction to shadowing. Various pedicellariae (jaw-like appendages around the base of spines) keep the body surface clear of encrusting organisms and aid in food capture. Larva exhibit negative phototaxis and photosensitive vertical migration [http://www.ncbi.nlm.nih.gov/pubmed/20067495 apparently mediated] by a encephalopsin-class opsin. | |||
[[Image:Opsin_urchin_expr.png|left|]] | [[Image:Opsin_urchin_expr.png|left|]] | ||
[[Image:UrchinOpsinCil.jpg|left]] | |||
Because sea urchin proteins are quite diverged from those of other deuterostomes (including sister hemichordates), it is difficult to recover accurate full-length sequences by homology alone, especially poorly conserved or short exons, in view of the so-so genome assembly. As of June 2011, only three of seven urchin opsins have any transcript support and two of these are apparent orthologs in related species, Hemicentrotus pulcherrimus and Strongylocentrotus droebachiensis. The single S. purpuratus opsin with experimental transcripts (3 fragmentary reads, no publication) is a neuropsin entirely missed by the urchin annotation effort. | |||
The | The H. pulcherrimus opsin [http://www.ncbi.nlm.nih.gov/nuccore/224809166 224809166] is 95% identical to an unstudied S. purpuratus opsin TMT1_strPur (represented at GenBank by two poor quality pipeline models XM_778209 and XM_001177470, the largely correct [http://annotation.hgsc.bcm.tmc.edu/Urchin/cgi-bin/viewGleanGene.cgi?gene_id=GLEAN3_05569 GLEAN3_05569] stored elsewhere), enabling the experimental picture to be consolidated over in S. purpuratus. | ||
The initial methionine remains somewhat uncertain and the DRY region is anomalous (QRC) but probably retains most motif functionality. Glycosylation occurs as in other ciliary opsins. The intron pattern 0.2.2.0.0 is a perfect match in position and phase to pinopsins. There is no detectable counterpart within the one hemichordate genome available (Saccoglossus). The best <span style="color: #990099;">Opsin Classifier</span> match 41% identity is to frog, TMTa_xenTro, which has a conventional ERY motif. | |||
The two remaining opsins classify as | Expression of MEL2_strPur begins during larval development at the 14 hours postfertilization swimming blastula stage initially in cells around the tip of the [http://en.wikipedia.org/wiki/Archenteron archenteron] in early gastrulae. By pluteus stage, cells expressing the encephalopsin are found in the aboral ectoderm but by eight-arm pluteus stage are restricted to the tips of the larval arms and posterior body. Pigment cells are present. Knockdown inhibits vertical swimming of the larvae, suggesting the opsin has a role in [http://onlinelibrary.wiley.com/doi/10.1111/j.1440-169X.2009.01154.x/full photosensitive larval swimming vertical migration] (a functionality commonly reported in earlier diverging species). In adult urchins, the gene is expressed exclusively in tube feet. | ||
The sea urchin genome contains a second potential ciliary opsin (ENC_strPur below). It aligns best to Branchiostoma and Platynereis ciliary opsins but only at 33% identity and not that distant to certain melanopsins. This opsin too is likely to be involved in some aspect of photoreception, though that won't advance vertebrate photoreceptor understanding. | |||
The two peropsin-class Go urchin sequences are adjacent in parallel tandem configuration with identical intron pattern but have only 64% amino acid identity, consistent with a moderately old tandem duplication and inconsistent with assembly stutter. Despite weak percentage identity among members of this group, these are K-296 opsins quite distinct from miscellaneous 'rhodopsin' superfamily GPCRs which have ([http://www1.elsevier.com/homepage/sah/emails/genome_si/pdf/sdarticle18.pdf 979 genes in 70 families] annotated in urchin). | |||
The two remaining opsins classify as rhabdomeric melanopsins. One of them, MEL2_strPur, has a GenBank transcript DQ285097 alluding to an unpublished expression study concerning tube feet photoreceptors. The other melanopsin is expressed post-oral arm of two-week-old larvae. Since the ancestral melanopsin has four exons, this evidently arose as a retroposed gene that retained functionality and displaced the now-vanished parental gene, an uncommon event but with numerous precedents in bilateran genomics. | |||
<br clear=all> | |||
cDNA Name Model Class DRY K246 Motif PubMed | |||
AB458218 ......... ............ <font color="blue">TMT1_hemPul QRC KLCTIHNPIIYFLLNKQFK [http://onlinelibrary.wiley.com/doi/10.1111/j.1440-169X.2009.01154.x/full Ooka]</font> | |||
........ Sp-Opsin1 GLEAN3_05569* <font color="blue">TMT1_strPur QRC KLCTIHNPIIYFLLNKQFK </font> | |||
........ Sp-opsin2 GLEAN3_03451** <font color="blue">ENC_strPur DRY KSSVMINPIIYAVTSRVFR</font> | |||
........ Sp-Opsin4 GLEAN3_22851 <font color="magenta">MEL1_strPur DRY KCSAIWNPIIYCLSHEKFN</font> | |||
........ Sp-opsin5 GLEAN3_06737** <font color="magenta">MEL2_strPur FRY KSSVIYNPLIYVVLNSKFR</font> [http://www.pnas.org/content/108/20/8367.full Ullrich] | |||
DQ285097 ......... ............ <font color="magenta">MEL2_strDro FRY KSSVIYNPLIYAVLNSKFR</font> [http://rspb.royalsocietypublishing.org/content/early/2011/03/28/rspb.2011.0336.abstra Lesser] | |||
........ Sp-Opsin3.1 GLEAN3_27634 <font color="green">PER2a_strPur YRY KSSCMVNPIIFLTSSSKFR</font> | |||
........ Sp-Opsin3.2 GLEAN3_27633* <font color="green">PER2b_strPur YRY KSSCMINPIIFLTSSSKFR</font> | |||
CX690664 ......... ............ <font color="brown">NEUR_strPur LRY KTSSVYNPIIYCIFNKSFR</font> | |||
* discrepancies vis-a-vis curated gene models provided below (typically different iMet or extraneous exons | |||
>TMT1_hemPul Hemicentrotus pulcherrimus (sea_urchin) Deut.Echi.Eleu AB458218 20067495 full G? Hp-encephalop larval vertical movement exons by homology | |||
0 MNYSTPVMTPTASFSGSWTSTIESTAMSNLMMNIVTNVNALSGIGNETPTTLGPSSLVVPVSRSTYNYLTVYTGFLTIFGILNNGIVMVLFARFPSLRHPINSFLFNVSLSDLIISCLASPFTFASNFAGRWLFGDLGCTIYAFLVFVA 1 | |||
2 GTEQIVILAALSIQRCMLVVRPFTAQKMTHRWALFFISLTWIYSLIICLPPLFGWNHYTYEGPGT 1 | |||
2 ACSVAWNSPLPGDTSYIIFIFVMVLVIPFGIIIFCYGLLVYAVKK 0 | |||
0 ISRTQAALSSEAKADRKVSKMIFIMILFFLIAWTPYTGFSLYVTFKKNVVITPLAGTFPPFFAKLCTIHNPIIYFLLNKQ 0 | |||
0 FKDALIQLLCCGENPFDRDESEHEGRGGRHRHRTAPSATAHIGGRGRASSLPTATSMLDIPQAASTTVSSSGKTQNKENLEKGPSTSETTNERVFQLSSKVQKFEISDKNKMPSSSELPGASSLSGALMPPRRAMKNQVGCLPPVDN* 0 | |||
>TMT1_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu AAGJ03036958 full G? encephalopsin-type introns Sp-Opsin1 GLEAN3_05569 no sacKow | |||
0 MNYSTPVMTSTASVSGPSPWTSTLESKAMSNLMTGLVTNVNALSGIGNETPTTIGLSSLVVPVSRTTYNYLTVYTGFLTIFGILNNGIVMILFARFPSLRHPINSFLFNVSLSDLIISCLASPFTFASNFAGRWLFGDLGCTLYAFLVFVA 1 | |||
2 GTEQIVILAALSIQRCMLVVRPFTAQKMTHRWALFFISLTWIYSLIICVPPLFGWNRYTYEGPGT 1 | |||
2 ACSVAWNSPSPGDTSYIIFIFVLVLVIPFGIIIFCYGLLVYAVKK 0 | |||
0 ISRTQAALSSEAKADRKVSKMIFIMILFFLIAWTPYTGFSLYVTFGKNVVITPLAGTFPPFFAKLCTIHNPIIYFLLNKQ 0 | |||
0 FKDALIQLFCCGENPFDRDESEHEGRGGRHRHRTAPSATAHIGGRGRASSLPTATSMLDIPQAASTAASSSGKTQNKESLEKGPSTSETTNKRVFELSSKIQKFEISEKNNTPSSSELPGASSLSGALMPPRRAMKNQVGCLPPVDN* 0 | |||
>ENC_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu AAGJ02133080 full G? Sp-opsin2 GLEAN3_03451 terminal exon extended to stop codon | |||
0 MENFTSIVTDGTNEENTDGDAWPGYAHLLAGSFLTLVFIISIIGNSVVLFLFAWDRHLRTPTNMFLLSLTISDWLVTVVGIPFVTASIYAHRWLFAHVGCIS 2 | |||
1 YAFIMTFLGLNSLMSHAVIAVDRYLVITKPHF 1 | |||
2 GIVVTYPKAFLMISIPWVFSFAWAVFPLAGWGEFTYEGTGAWCSVRWDSDQPQIMSYVLAMMFLTFISSIVIMMYCYICIFLTTRRMPRWATSNSIKTHERNRRRR 2 | |||
1 EQKLLKTLIAIAIAFLVAWSPYAITSMIVVFGGSELLSLTATTLPSLFAKSSVMINPIIYAVTSRVFRKSLKK 0 | |||
0 MLTSFFPGCMTYIMTDKSPPSSSRPIQLGLCKYHFLY* 0 | |||
>MEL1_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XR_026330 frag G? Sp-Opsin4 GLEAN3_22851 no cdna losing introns larval postoral arm ancestral intron | |||
0 MNAVTTALPHGLNKPTIEAR 2 | |||
1 WTKSLRTPPNMLIVNLAISDFGMVITNFPLMFASTIYNRWLFGDA 1 | |||
2 GCQFYAFCGALFGIMSIANMTAIALDR 2 | |||
1 YYVICWSLEAVRSVTHRRSMIIIIIVWCYAIFWSIPPFFGVGSYVLEGYGLGCTFDFMTKDLNHYLHV | |||
SFLFASSFVVPVTIIIVCFTRIAITVRAHRHELNKMRTKLTEDKDKKHKSSIRRANKAKTEFQIAKVGFQVTIFYVLSWM | |||
PYSIVAVIGQYFDSDLLTPLGTVVPVIFAKCSAIWNPIIYCLSHEKFNAALKEKLMGMCGIEIPSKHRSMGSQESSVTGR | |||
RGMHRQNSSTLSESSVTSTVDQDAIELKDRKQGPATVKVQQEKVEGGTYRRNPGDVTFSKDAGVEVDEKRRGDQGQRDDR | |||
VRPQGEGQMDQWSQPPPAPASASAPTPGVNDKEYLTKM* 0 | |||
>MEL2_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu full G? Sp-opsin5 GLEAN3_06737 tube feet | |||
0 MPTTLMENSTPGWMADDSQMEETHPAFPLIGGYLLVVVLLGTAGNSLVIYTFLRFKKLHSPINLLIVNLSASDLLVATTG | |||
TPLSMVSSFYGRWLFGTNACAFYGFVNYYCGCISLNSLAAISVFRYIIVVRGQAQNNKLSLRSSIYAILVIHLYTLIFST | |||
PPLYGWNRFVLAGYHTSCDIDFHTKTPLFVSYICYMFFFLFFLPLGLISWSYFKIYQRVSKHSNSMRTSFTGVTKEINSDEKHA 2 | |||
1 FNHRRTASTLFVTIVVFLFAWFPYCIVSLWVLIGDANSISKLSTTIPSLFAKSSVIYNPLIYVVLNSKFRKALIQTLSFLKCLSKHELSESS* 0 | |||
>MEL2_strDro Strongylocentrotus droebachiensis (sea_urchin) Deut.Echi.Eleu DQ285097 full G? Sp-opsin5 GLEAN3_06737 tube feet retained intron WLEKMKTTQILHKPVTFLRLKPSFEPRLKPRFKPR | |||
0 MPTTLMENSTPGWIADEGEMEETHPEFSLIGGYLLVVVLLGTAGNSLVIYTFLRFKKLHSPINLLIVNLSASDLLVATTGTPLSMVSSFYGRWLFGTNACAFYGFVNYYCGCISLNSLAAISV | |||
FRYIIIVRGKAQNNKLSLRSSIYAILVIHLYTLIFSTPPLYGWNRFVLAGYHTSCDIDFHTKTPLFVSYICYMFFFLFFLPLGLISWSYFKIYQRVSQHSNSMRTSFTGVTKETNIDEKHA 2 | |||
1 FNHRRTASTLLVTIVVFLFAWFPYCIVSLWVLIGDANSISKLSTTIPSLFAKSSVIYNPLIYAVLNSKFRKALIQTLSSIKCLYTHELSESS* | |||
>PER2a_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_778236 full Go Sp-Opsin3.1 GLEAN3_27634 RRH peropsin overshoots iMet spread across tandem | |||
0 MAASVTESSATEAISRLEPEYMVPLTRTGYLLTAIYLTIV 1 | |||
2 GSIATVGNITVICVLCRYRTFRKRSINLLLINMAASDLGVSVAGYPLTTVSGYWGRWLFGDVGCQFYAFCVYTLSCSTISTHAAIAVYRYIYIVKTDL 1 | |||
2 RPKLTANFTSGVIVVIWVYAFFWTVTPFVGWSSYIYEPFGTSCSVNWVGRTISDISYMVACTIGVYLLQIFIMLYCYIRVAKK 2 | |||
1 IRGVDPGRTEEKDAGVVVFGRLRKREAKIDTHVTK 0 | |||
0 MCFMMMLTFIVVWAPYAVECLRAAHVHRISALSSVLPTMFAKSSCMVNPIIFLTSSSKFRQDLGKLWSRPSSQDSLQLEER | |||
NKTQRSLYVRHSELGSAHGNDTASVYYEKERIYIGEMRATSIQKEAELLQRDPELLSIASSTNSDVKFVVRDRPKRYTKR | |||
PVKPQGPRGPEMFTASGVTNKGSSTSDSGGQSTSSGTTGSKPKRSGRKASRQYSMKSQSEDTGEIFTLDGSALEMMSLRKL* 0 | |||
>PER2b_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_778236 17067569 full Go Sp-Opsin3.2 GLEAN3_27633 RRH peropsin no cdna inline tandem PER2a_strPur | |||
0 MNSFSEESYVTDPTTTQPTLFLTPLSQTGYLLTALYLTLV 1 | |||
2 GIVSTIGNITVLCVLCRYGTFRKRSVNILLMNMAVSDLGVSVAGYPLTAISGYRGRWVFADIGCQFSGFCVYALSCSTISTHAVVAIYRYIYIVKPYH 1 | |||
2 RPRLSSSTSCLAILCIWTFTLFWTITPFFGWSSYTYEPFGTSCSINWYGKSLGDLTYIICCVVFVFILPIIIMLYCYIGVAKK 2 | |||
1 IKGIDPLRTEERDIAVVFGRLRKHETKIDTRVTK 0 | |||
0 ICFMMMASFIVVWTPYAVGSIWASKIGKISASASVLPTMFAKSSCMINPIIFLTSSSKFRADLGKLWNRPSSLEHTIRVEERSREQRSFF | |||
VRQSALPDAMVSRSASVYYDKERIYIGEMRAASIQKEADLLHRDPEAISIASSTSSSLQFVLKDRQNRYKKKAGEASKKGSNILHFPYDDTE | |||
GSMINNLMRPRSHSVTSDNISRVFAPSLKRPTKKRSMSHPDIPSTSADIFTVSPTTIKNLQKQ* 0 | |||
>NEUR_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_001197837 full G? CX694910 CX690664 no GLEAN3 blastula | |||
0 MDVNAKWWTNETLRTRDQFSDDHYTSVLSYEGDIWAGVYLMFI 1 | |||
2 SLIAFIGNISVIVISLRKREKLKPIDLLTINLAIADFLICVVSYPLPMISAFRHR 0 | |||
0 WSFGKFGCVWYGFTSFLFAVGSMATLMVIALLRYAKLCRENV 1 | |||
2 DQYQSRPFVIKVIVAIWGFAFFTTAPPLFGWS 2 | |||
1 SYVPEPYHLSCTIDFADTSPSGLSYTYFTTIVVFFMPLMIIVLCYVAIARKMIHHNRRINVGHNAGRMLLEIRLLK 0 | |||
0 TACMITMAYTISWTPYAVIAMWVTYIPVNQIPDAFRILPAFCAKTSSVYNPIIYCIFNKSFRQDLSSLICCCACQCYTITINLDINSHAQQQFRRIEERR | |||
DEVGTYKRRPLMICSNPFAWSRDFHETWRQRRIRGIHRNCRNNVRVENINVNFRRDTDMVELNAPTPAEIHRPELNTASTRSGARTKSMATHLPALEEVPSG | |||
APQCSALLHNTPIPRSLQGTPLPYQPQPSTSDLHDEFLNPSVVSRNMCVIVVKPNIEEELSTD* 0 | |||
=== <span style="color: #990099;">Hemichordata: Saccoglossus kowalevskii (acornworm) .. 2 peropsins</span> === | === <span style="color: #990099;">Hemichordata: Saccoglossus kowalevskii (acornworm) .. 2 peropsins</span> === | ||
This surviving member of early branching deuterostomes has excellent genomic and | This surviving member of early branching deuterostomes has excellent genomic and transcript coverage, with diverse full length multi-exon genes often recoverable. Be aware that some transcript data has been <span style="color: #990099;">misplaced</span> by GenBank to reside under Saccoglossus 'other' at the trace archives while [http://www.biolbull.org/cgi/content/full/214/3/284?view=long&pmid=18574105 new transcription factor ESTs], accession numbers FF418995-FF534157 and FF602128-FF677500, are properly located. | ||
[[Image:SaccoKol.jpg|left|]] | [[Image:SaccoKol.jpg|left|]] | ||
| Line 218: | Line 676: | ||
However acorn worm may not illuminate photoreception at its ancestral node with echinoderms (Ambulacraria). Acornworm have isolated photoreceptive cells are [http://webs.lander.edu/rsfox/invertebrates/saccoglossus.html scattered through the epidermis] but no eyes or eye spots even in S. horsti [http://adsabs.harvard.edu/abs/1952RSPTB.236..553B non-phototaxic planktonic larva], as befits an animal that settles in its burrow on day two. Light striking epidermal photoreceptors elicits burrowing behavior. | However acorn worm may not illuminate photoreception at its ancestral node with echinoderms (Ambulacraria). Acornworm have isolated photoreceptive cells are [http://webs.lander.edu/rsfox/invertebrates/saccoglossus.html scattered through the epidermis] but no eyes or eye spots even in S. horsti [http://adsabs.harvard.edu/abs/1952RSPTB.236..553B non-phototaxic planktonic larva], as befits an animal that settles in its burrow on day two. Light striking epidermal photoreceptors elicits burrowing behavior. | ||
In searching for opsins that might underlie epidermal (or other) photoreception, the best queries (for detecting diverged sequences) are sea urchin and amphioxus opsins versus | In searching for opsins that might underlie epidermal (or other) photoreception, the best queries (for detecting diverged sequences) are sea urchin and amphioxus opsins versus Saccoglossus trace and est 'other' because, being transcripts rather than short exons these give longer matches. However opsins expressed in scattered cells may not be represented in transcript collections if rarely transcribed or restricted to a narrow unsampled window of development. | ||
Promising traces must be back-blastxed against the <span style="color: #990099;">Opsin Classifier</span> because the initial query choice may have | Promising traces must be back-blastxed against the <span style="color: #990099;">Opsin Classifier</span> because the initial query choice may have been sub-optimal. Good matches must be intronated using the exact-matching Saccoglossus probe against genomic trace reads because intron patterns are critical adjuncts to low-scoring sequence alignments in establishing opsin orthology classes as surviving synteny at this time depth is rare and not currently determinable. | ||
I report here the very first hemichordate opsins. Although quite diverged, these classify cleanly as peropsins, a class that appears significantly expanded in Saccoglossus. Here again the Schiff base lysine is present but the nature of covalent ligand (if any) is by no means established as retinal. While | I report here the very first hemichordate opsins. Although quite diverged, these classify cleanly as peropsins, a class that appears significantly expanded in Saccoglossus. Here again the Schiff base lysine is present but the nature of covalent ligand (if any) is by no means established as retinal. While peropsins might accomplish the photoreceptive task in this species (and so may represent yet another approach to evolving photosensors), they cannot serve as ancestral sequence to deuterostome ciliary opsins (which are well-represented in sister taxon echinoderms). | ||
Hence ciliary opsins have been lost in Saccoglossus. Whatever its [http://www.ncbi.nlm.nih.gov/pubmed/18192182,18192177 other merits] for understanding body plan or centralized nervous system origins, this species cannot clarify why ciliary opsins were retained in early deuterostomes prior to the evolution of imaging vision. It's not clear whether additional enteropneust or pterobranch hemichordate species would improve this situation -- the 90 known are all benthic marine. | Hence ciliary opsins have been lost in Saccoglossus. Whatever its [http://www.ncbi.nlm.nih.gov/pubmed/18192182,18192177 other merits] for understanding body plan or centralized nervous system origins, this species cannot clarify why ciliary opsins were retained in early deuterostomes prior to the evolution of imaging vision. It's not clear whether additional enteropneust or pterobranch hemichordate species would improve this situation -- the 90 known are all benthic marine. | ||
The process for obtaining accurate gene models in such a remote species prior to assembly is a difficult exercise in bioinformatics and use of the [ | The process for obtaining accurate gene models in such a remote species prior to assembly is a difficult exercise in bioinformatics and use of the [[Opsin_evolution|Opsin Classifier]] -- a detailed procedure is given in the annotation tricks section. A final check consists of back-blast of the final sequence against all of genBank to verify top hits are deuterostome peropsins. This cannot detect chimeric proteins (unbridgeable fragments) however. | ||
A complete set of opsins for this species | A complete set of opsins for this species became feasible with the long-overdue release of the [http://www.genome.gov/Pages/Research/Sequencing/SeqProposals/AcornWorm.pdf 7x assembled contigs] by the Baylor sequencing center on 04 AUG 2009. This enabled some minor fixes in the two previously recovered peropsin sequences (which can now be seen to be identically intronated and 34% identical) but did not result in any additional opsins. | ||
<pre> | <pre> | ||
> | >PER1a_sacKol Saccoglossus kowalevskii (acornworm) 7 ests + ACQM01133041 | ||
0 MVTTDSLANSTDEPVPSILTLQQHYAASVTLLAL 1 | |||
2 | 2 AVIGTVLSSVNFRMLLSNPDYCSKAGNFFLSLAVTDL 1 | ||
0 | 2 CVCIFETPFSAFSHHAGFWIFGDTACQLYAFFGIFFGLVNIFMVTFISLDRYWATCSPVE 1 | ||
2 VELKSKYYTRMTALGWMVALFWAAAPVFGWSRYaMEPSMASCSIDYMTNDFSYVTYITCLTLTCYVVPIVVMVYCYVKASKNIKYTGKVTEWAHENNATK 0 | |||
0 ISRLCVLQLVFCWSLYGFNCMWTVVADDVETLPKMLTVLAPILAKTTPILNSGLYFLHNKKFRGAAVDMFKAKEE* 0 | |||
> | >PER1b_sacKol Saccoglossus kowalevskii (acornworm) ACQM01067921 | ||
2 | 2 aVLSVIGNSVVLEMFRRYKELLSPSAILLISLALADL 1 | ||
0 | 2 GLTIFGMSLSCVSSFAGRWLFGKFGCYFHGFAGMLFGLGSIGNLTVISIDRYIITCKRsL 1 | ||
2 WSYRHYYALLAVAWSNALFWSMMPLFGWSSYALEPEGTSCTIDWMNNDNQYISYVSCVTVTCFILPCAVMTYDYLAAYMKMVKAGYTLSEETEKPNND 0 | |||
0 MCIALVAAFLLSWFPSATVFLWAAFGNPGNIPLSFTGVADAFTKIPAVFNPVIYVALNPEFRKYFGKTIGCRRKRKKPIAVRLNGSEQNVENTI* 0 | |||
</pre> | </pre> | ||
| Line 247: | Line 709: | ||
No genome or major transcript studies are under consideration. A quick check via tblastn of sea urchin opsins against available transcripts does not turn up good opsin candidates as of 28 Nov 2007 (other than a weak melanopsin match in Convoluta, EV602614, that might instead be generic GPCR). No information about photoreception in these species is readily available. While the above two taxa might not be ideal for opsin purposes, extant species are very limited. | No genome or major transcript studies are under consideration. A quick check via tblastn of sea urchin opsins against available transcripts does not turn up good opsin candidates as of 28 Nov 2007 (other than a weak melanopsin match in Convoluta, EV602614, that might instead be generic GPCR). No information about photoreception in these species is readily available. While the above two taxa might not be ideal for opsin purposes, extant species are very limited. | ||
'''See also:''' [[Opsin_evolution|Curated Sequences]] | [[Opsin_evolution:_key_critters_%28cnidaria%29|Cnidaria]] | [[Opsin_evolution:_key_critters_%28ecdysozoa%29|Ecdysozoa]] | [[Opsin_evolution:_key_critters_%28lophotrochozoa%29|Lophotrochozoa]] | [[Opsin_evolution:_update_blog|Update Blog]] | |||
[[Category:Comparative Genomics]] | [[Category:Comparative Genomics]] | ||
Latest revision as of 13:55, 9 June 2011
See also: Curated Sequences | Cnidaria | Ecdysozoa | Lophotrochozoa | Update Blog
Marsupials: Sarcophilus harrisii (tasmanian devil) .. 8 opsins
The 454-based genome of Sarcophilus harrisii (tasmanian devil) has recently become available (with better coverage than the new Macropus eugenii Sanger genome). Sarcophilus provides a fresh check on received wisdom on the mammalian opsin portfolio. Here the entire repertoire of opsin genes are collected for this species along with a summary of what is available for the marsupial clade and its current phylogenetic organization. As expected, Sarcophilus has but 8 of the 20 ancient vertebrate opsin genes.
RHO1: 5 marsupials SWS2: 10 marsupials LWS: 10 marsupials ENCEPH: 3 marsupials TMT: 3 marsupials PER: 3 marsupials NEUR1: 3 marsupials MEL1: 4 marsupials
The optimal wavelength for scotopic (dim light) vision of Sarcophilus is easily predictable provided key tuning residues are covered by the assembly. The 97% match to Sminthopsis and agreement at tuning residues suggests this aspect of vision will be nearly identical between the two species.
Cone rhodopsin RHO2 has been lost in all mammals and no debris from this gene is expected in Sarcophilus. The short wavelength cone opsin SWS2, while still present in platypus, has also been lost in all theran opsin too long ago to leave detectable remnants in syntenic position. Cone opsin SWS1 has this turned around, being present in theran mammals but only as debris in platypus. A nearly full length gene, most similar to Sminthopsis, can be recovered from Sarcophilus read coverage.
The basal long wavelength LWS imaging opsin is available from 97 vertebrates and has already been analyzed for phyloSNPs and rare genomic events. The Didelphimorphia experienced a 3-4 residue insert in exon 1 that separates them from all other marsupials. Note this region has quite a complicated indel history. The extra residues have repeat character DVNE DDND suggesting replication slippage. The gene is present and intact in Sarcophilus though two exons are not currently available. LWS so far in tasmanian devil is identical to its Sminthopsis ortholog.
LWS_loxAfr MAQQWGPHRLTGARLQDASE---DSTQASIFVYTNTNT elephant LWS_echTel MAQRWGAHRLTGGQLQDTYE---GSTRTSIFVYTNSTS tenrec LWS_monDom MTQAWDPAGFLARRRDVNEDDNDETTRSSLFVYTNSNN Didelphimorphia LWS_didAur MTQAWDPVGFLARRRDENEDDHDDTTRASLFVYTNSNN Didelphimorphia LWS_tarRos MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia LWS_macEug MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia LWS_smiCra MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Dasyuromorphia LWS_sacHar MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Dasyuromorphia LWS_setBra MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Diprotodontia LWS_cerCon MTQAWDPAGFLAWQEDENE----ETTRASLFVYTNSNN Diprotodontia LWS_myrFas MTQAWDPAGFLAWRREENE----ETTRASLFTYTNSNN Dasyuromorphia LWS_isoObe MTQAWDPAGFLAWRRDENE----ETTRASLFVYTNSNN Peramelemorphia LWS_ornAna MTPAWNSGVYAARRRFEDEE---DTTRTSVFVYTNSNN platupus LWS_tacAcu MTQAWDPAGFLAWRRDENEE---TTRASLFVYTNSNNT echidna
Pinopsin, parapinopsin, parietopsin and VA opsin terminate in sauropods so are missing in all mammals including marsupials. Encephalopsin has a very peculiar history of gene loss in tetrapods, requiring some seven independent and asynchronous events including platypus. While this limits the phylogenetic utility of any gene loss within marsupials, the status of the gene within Sarcophilus is still informative. A full length gene can be recovered with 94% identity to opossum, strongly indicating that encephalopsin is fully functional within Sarcophilus.
TMT is an ancient locus that is present in monotremes and marsupials but lost in all placentals.
RGR has apparently been lost specifically in the marsupial clade, though support for that is only provided by the Monodelphis and Macropus genome projects. It would be of considerable interest to find the gene or a fragment thereof in syntenic position in Sarcophilus. However nothing can be found with tblastn of current reads.
Sarcophillus can be expected to have this gene and it does. The protein sequence substantiates the 4 previously defined phyloSNPs characteristic of the marsupial/placental transition.
Here Sarcophilus can be predicted to contain only NEUR1 because the ancient vertebrate genes NEUR2 and NEUR3 appear to terminate in sauropods and NEUR4 in platypus.
Similarly Sarcophilus can be expected to have the main melanopsin MEL1 but not the paralog MEL2 which terminates in sauropods.
Marsupial phylogenetic relationships have been reviewed in a 2009 paper that established the mitochondrial genome sequences of the Tasmanian tiger (Thylacinus cynocephalus) and numbat (Myrmecobius fasciatus). It emerges from this that thylacine is basal among Dasyuromorpha with numbat diverging next: (thyCyn,(myrFas,(smiCra, sarHar))). Dunnart and tasmanian devil are very similar at the protein level, in some cases 100% identical.
Newick tree that generates the marsupial-centric vertebrate phylogenetic tree: ((((((((((((sarHar,smiCra),myrFas),thyCyn),(macEug,triVul)),monDom), ((((loxAfr,proCap),echTel),(dasNov,choHof)), ((((((bosTau,turTru),susScr),vicPac),((equCab,(felCat,canFam)),(myoLuc,pteVam))),(eriEur,sorAra)), (((((((((homSap,panTro),gorGor),ponPyg),macMul),calJac),tarSyr),(micMur,otoGar)),tupBel), (((((musMus,ratNor),dipOrd),cavPor),speTri),(oryCun,ochPri)))))), (ornAna,tacAcu)), ((galGal,taeGut),anoCar)), xenTro), (((tetNig,takRub),(gasAcu,oryLap)),danRer)), calMil), petMar);
>RHO1_sacHar Sarcophilus harrisii (tasmanian_devil) 97% identity Sminthopsis crassicaudata 0 MNGTEGPNFYVPHSNKTGVVRSPYEEPQYYLAEPWMFSCLAAYMFMLIVLGFPINFLTLYVTIQHKKLRTPLNYILLNLAVADLFMVICGFTTTLVTSLNGYFVFGTTGCQIEGFFATTG 1 2 GEVALWALVVLAIERYIVVCKPMSNFRFGENHAIMGVVFTWIMALACSVPPLFGWSR 2 1 YIPEGMQCSCGIDYYTLNPEFNNESFVIYMFVVHFTIPLTVIFFCYGQLVFTVKE 0 0 AAAQQQESATTQKAEKEVTRMVIIMVIAFLICWVPYASVAFYIFTHQGSDFGPIFMTLPAFFAKSSSIYNPVIYIMMNKQ 0 0 FRTCMITTLCCGKNPLGDDEASATVSKTETSQVAPA* 0 >SWS1_sacHar Sarcophilus harrisii (tasmanian_devil) part of last exon missing 96% identity Sminthopsis crassicaudata 0 MSGDEEFYLFKNISPVGPWDGPQYHIAPAWAFHLQTAFMGFVFFAGTPLNGVVLIATLRYKKLRQPLNYILVNISLAGFIFCVFSVFTVFVSSSQGYFVFGRHVCAMEGFLGSVA 1 2 SGLVTGWSLAFLAFERFIVICKPFGNFRFHSKHATMVVLATWVIGIGVSIPPFFGWSR 2 1 YIPEGLQCSCGPDWYTVGTKYRSEYYTWFLFIFCFIVPLSLICFSYSQLLGALRAVS 0 0 VAAQQQESATTQKAEREVSRMVVVMVGSFCLCYVPYAALAMYMVNNRNHGLDLRLVTIPAFFSKSACVYNPIIYCFMNQ 0 0 KPMTDDSETTSSQKTEVSTVSSSQVGPS* 0 >LWS_sacHar Sarcophilus harrisii (tasmanian_devil) half of exon 2, all of exon 4 missing frag 100% identical to Sminthopsis 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTK 1 2 FKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMSTCCILPLSIIILCYIQVWLAIRA 0 0 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >ENCEPH_sacHar Sarcophilus harrisii (tasmanian_devil) 94% identity monDom 0 MYSGNSSDDAGGGYWGSGGTGGAGGTGVAGEPAPEGSPRPAPLFSPGTYELLALLIATIGLLGLCNNLLVLVLYYKFQRLRTPTHLFLVNISFSDLLVSLFGVTFTFVSCLRSGWVWDSVGCAWDGFSNTLF 1 2 GIVSIMTLTVLAYERYNRIVHAKVINFSWAWRAITYIWLYSLIWTGAPLLGWNRYTLEIHGLGCSVDWKSKDPNDSSFVLFLFLGCLVLPVGVMAYCYGHILYAIRM 0 0 FRCVEELQTLQVIKILRYEKKVAKMCFLMIATFLFCWMPHAVICFLVANGYGSLVTPTVAIIPSLFAKSSTAYNPIIYIFMSRK 0 0 FRRCLLQLLcFRQLKFQQPKKDRAIIRTEKQIRPIVMSQKVGDRPKKKVTFSSSSIIFIITSDETQMIDDNDKNSETKVNVIQVRPL* 0 >TMT_sacHar Sarcophilus harrisii (tasmanian_devil) FP5MBH101BETOZ needed to finish 0 MSINLTTNLSFGPLLIDSEEKPRSGLSRTGHTVVAVFLGIILILGFINNFIVLILFCKFKVLRNPVNMLLLNISISDMLVCLSGTTLSFASSIRGRWIGGYHGCRWYGFANSCF 1 2 GIVSLISLAILSYERYRTLTLCPRRGADYQKALLAVAGSWLYSLIWTVPPLIGWSSYGTEGAGTSCSVHWTSKSVQSVSYIMCLFIFCLVIPILIMIYFYGRLLYTVKQ 0 0 VGKIRKTAARKREYHVLFMVVTAVICYLICWVPYGLIALVATFGPPGVVSPVANIVPSILAKSSTVCNPIIYILMNKQ 0 >PER_sarHar Sarcophilus harrisii (tasmanian_devil) 5.5 of 7 exons 0 MFKNDSFRSLEPEKEGHSVFSPAEHNIVAAYLITA 1 2 SILSNVIVLGIFVKFKELRTATNAIIINLA 0 0 1 2 GRRMTSFNYTIMILTAWVNGFFWALMPIVGWASYAPDPTGA 2 1 SFVSYTVTVIAINFVMPLVVMIYCYYNVSQKIKQYTPSNCPEYINRDWSNEVAVTK 0 0 MSVIMILMFLLAWSPYSVVCLWASFGDPKEIPPAMAIIAPLFAKSSTFYNPCIYVAANKK 2 1 FRrAISAMIQCQTHQSMSVSKALPMN* 0 >NEUR1_sarHar Sarcophilus harrisii (tasmanian_devil) 4 of 6 exons 0 1 2 GVLSTLGNGYVIYMSSKRKKKLRPAEIMTVNLAVCDLGIS 1 2 VVGKPFTIISCFSHRWVFGWVGCRWYGWAGFFFGCGSLITMTAVSLDRYLKICHLSY 1 2 GTWLKRHHAYICLVIIWAYATFWATMPLAGLGNYAPEPFGTSCTLDWWLAQASVTGQTFILNILFFCLLLPTAVIVFSYVKIIAKVKSSTKEVAHFDSRIQNSHSHVLEMKLTK 0 0 VAMLICAGFLIAWIPYAVVSVWSAFGQPDSIPVQFSVVPTLLAKSAAMYNPIIYQVIDCKFACCQSGGQKAAKKESLRDYR 2 1 * 0 >MEL1_sarHar Sarcophilus harrisii (tasmanian_devil)96% identity smiCra last exon missing FKUJDAX01C1KMN needed 0 MNPSPMLRHLSCSAQDTNCTKIMASISEWNNTEVDAYHLVDLPPITPT 0 0 AVVLPPYSQNVFPTADVPDYAHYTIGATILVVGFTGVLGNLLVIYTFCR 2 1 SRSLRTPANMFIINLAISDFFMSFTQAPVFFASSLYKRWIFGEK 2 1 GCEFYAFCGALFGITSMITLMVIALDRYFVITRPLASIGMISKKKTGLILLGVWLYSLAWSLPPFFGW 1 2 sAYVPEGLLTSCSWDYTTFTPSVRAYTILLFCFVFFIPLIVIIYCYIFIFRAIKDTNK 2 1 AVQNIGSRASTPSPRHFQRMKNEWKMAKIALVVILLFVLSWAPYSTVALVAFAG 2 1 YSHVLTPYMNSVPAIIAKASAIHNPIIYAISHPKYR 2 1 MAIAQNFPCLRAVLGIRHPRTQSFSSYRFTHRSTTASQASDISWQSRGRRQLSLGSESEA 0 0 GWNNIEAGIEGLTLRSLEGYCGMDEETMETREPSASAKAKGQ 0 0 * 0
Snakes: Python molarus (python) .. 11 opsins
The python genome became available in Feb 2011. Snakes fill an important data niche -- while still amniotes, they are sister to the squamate lizard Anolis and together these are sister to the three birds with determined genomes. These determine the diapsid node; when a turtle genome becomes available, that will allow reconstruction of the sauropsid node of mammalian divergence.
In terms of opsins, ten are completely missing from the python assembly: RHO2, SWS2, PIN, PPIN, PARIE, TMTa, TMT2, TMT3, MEL1, and NEUR3. Here TMT2 is definitely expected whereas its paralogs have perhaps already headed out the door, phylogenetically speaking. While this could reflect inadequate coverage, the opsins that are present seem to have good coverage (nearly all their exons). However it should be noted that contigs are very small, often 1-2 kbp, barely an assembly at all.
The 11 opsins from this assembly are shown below. Missing exons are shown in blue.
>RHO1_pytMol Python molarus (python) 0 MNGTEGLNFYVPMSNKTGIVRSPFEYPQYYLAEPWKYSALGAYMFLLILLGFPINFLTLYVTIQHKKLRTPLNYILLNLAIANLFMVLVGFTTTMYTSMNGYFVFGTVGCNVEGFFATLG 1 2 GEMALWSLVVLAIERYVVVCKPMSNFRFTETHAIMGLCFTWIMALACAGPPLVGWSR 2 1 YIPEGMQCSCGVDYYTPTPEVHNESFVIYMFIVHFVIPLAVIFFCYGRLVCTVKE 0 0 AAAQQQESATTQKAEKEVTRMVIIMVIAFLICWVPYASVAFYIFTHQGSDFGPVFMTIPAFFAKSSAIYNPVIYIVLNKQ 0 0 FRNCMITTLCCGKNPLAEDDTSAGTKTETSTVSTSQVSPA* 0 >SWS1_pytMol Python molarus (python) 0 MSGEEDFYLFENISSVGPWDGPQYHIAPMWAFHFQTLFMGLVFFAGTPLNAIILIVTIKYKKLRQPLNYILVNISFAGLLFCVFAVFTVFLASSQGYFFFGRHVCALEAFLGSVA 1 2 GLVTGWSLAFLAFERYVVICKPLGNFRFNSKHALLVVVATWVIGVGVSVPPFFGWSR 2 1 FIPEGLQCSCGPDWYTVGTKYKSEYYSWFLFVFCFIVPLSLIVFSYLRLLGALRA 0 0 VAAQQQESATTQKAEREVSRMVVVMVGSFCLCYVPYATLAMYMVNHPKHGLDLRLVTIPAFFSKSSCVYNPIIYCFMNKQ 0 0 FRACIMQTVCGKPLTDESDAGSSAQKTEVSSVSSSQVSPSSGAGATGSPRS* 0 >LWS_pytMol Python molarus (python) 0 MQTRGKEVAKAMTEAWNVAVFAARRRNDDDTTRESVFIYTNSNNTR 1 2 GFEGPNYHIAPRWVYNLTSLWMVFVVIASVFTNGLVLVATAKFKKLRHPLNWILVNLAIA DLGETVIASTISVINQFFGYFILGHPLCVVEGYTVSVC 1 2 GITGLWSLAIISWERWVVVCKPFGNLKFDAKLALAGILFSWVWSATWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGNEDPGVQSYMIVLMVTCCIIPLSIIILCYLQVWMAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMILAYIFCWGPYTTFACFAAANPGYAFHPMTASLPAFFAKSATIYNPIIYVFMNRQ 0 0 QFRNCIMQLFGKKVDDGSEVSSTSRTEVSSVSNSSVSPA* 0 >VAOP_pytMol Python molarus (python) 0 MDGARRESENGSLLLDASAGSSLVDPFSQPLDSIESWNFHLLSALMFLVTSLSLFENFTVILVTLKFRQLRQPLNYVIVNLSVADFLVSLVGGTISFLTNLKGYFYMGRWACVLEGFAVTFF 1 2 GIVALWSLALLAFERYVVICRPLGNVHFKGKDATLGIAFVWIFSFIWTIPPTMGWSSYTTSKIGTTCEPNW 2 1 YSGEYSDHTFIITFFTTCFIMPLLIILVSYGKLMRKLRK 0 0 vsntqgRLGSTRKPERQVTRMVVIMILAFLICWTPYAAFSILVTTCPSLELDPRLAAIPAFFSKTATVYNPVIYVFMNNQ 0 0 FRKCLVQLFRCSRQDIADSNMNQISKRAVLTPTKKYGEVSTAAARVTIFNQRNEDERSSsQSFALLSVLENWLCQFLRG* 0 >ENC_pytMol Python molarus (python) 0 MYSGNGSGGSLEPTAGKQDLERRRQQPAGDSGTPFSASTYELLALLIAAIGFVGLCNNLLVLVVYTKFKRLRTPTNLFLVNISLSDLLVSLFGVSFTFLSCLRNHWAWDAAGCVWDGFSNSLFG 1 2 GIVSLMTLTVLAYERYIRVVHARVVDFSWSWRAITYIWLYSLAWTGAPLLGWNHYTLELHGLGCSVDWSSREPGDTSFVFFFFLGCLVAPLGIVVYCYGHILHAVRM 0 0 FRRVEELQTVHVIKILRYEKKVAKMCFLMITTFLICWMPYAVVSLLIAYGYGHLITPTVAIIPSFFAKSNTAYNPVIFIFMSRK 0 0 FRRCLVQLFCVQFLRFKRTLKEQPAIESNKPIRPIVMSQKVGDRPKKKVtfssssiifiitsDDTEQIDVSTKCSDTKINVIQVKPL* 0 >MEL2_pytMol Python molarus (python) 0 MATAHPTKVDAPDYVLYAVGSCVLVIGCIGIIGNLLVLYAFYS 2 1 NKRLRTPPNYFIMNLAVSDFLMCATQAPVCFFNSIHKEWVLGDT 1 2 GCNFYAFCGALFGITSMMTLLAISVDRYCVITKPLQSIKRSSKKRSCLIIMFVWLYSLGWSVCPLFGW 1 2 EKDIPEGLMISCTWDYVTYSPANRSYTMLLCCCVFFIPLIIIFHCYLCMFLAIRNTGR 2 1 RKHSASQNIKSEWKLAKIAFVAIVVYVVSWSPYACVTLIAWAG 2 1 YARILTPYSKSVPAIIAKASAIYNPIIYAIIHPSYR 2 1 RTIRSAVPCLRFLIPISKSDLSTSSMSESSFRASVSSRHSFSYRNKSTYISSISAKET 0 0 TWNNVELDPVESVHTKLQPPQSNSFSTNAEEKSELPMKAPGCDVPTEEKVKAVFSNAPIHFTNKSSCVQAQSGVLPPSTVVVVMFIQ 0 >RGR1_pytMol Python molarus (python) 0 MTVSDSLPEGFTELEMFSFGTVMLAE 1 2 ALFGFSLNVLTIVSFWKITELQTPGNFLIFNLALSDCGICINALIAAFSTFLR 2 1 RYWPYGSDGCQIHGFQGFLTALTSINCAAAIAWDRYHQYCT 1 2 RSKLQWNSVFSMGLCAWCFAGFWSAMPLLGWGTYDYEPLRTCCTLDYTKGDR 2 1 NYIMFLIPLVLFNFVIPIFIMLMSYQSIDNKFRKTAQVK 0 0 FNTGLPVKSLVICWGPYSLLSFYAAVENVAFVSPKILM 0 0 IPALMAKTSPTMNAFIYALGNENYRGGMWQFLTGEKIEKAKIDDKTN* 0 >PER1_pytMol Python molarus (python) 0 MRRNDSANLLESEHHDRSAFSQTDHNIVAAYLITA 1 2 GVISLLSNIVVLSIFIKFKELRTPSNTIIIHLAFTDIGVSSIGYPMSAASDLHGSWKFGYMGCQ 0 0 IYAGLNIFFGMSSIGLLTVVAVDRYLTICRPAI 1 2 GNRLTAHNYIALIFAAWTNAVFWASMPVVGWASYAPDPTGATCTINWRENDM 2 1 SFISYTMTVIAVNFAVPLIVMFYCYYNVSKAMRQYPASRVLENLNIDWSEQVDVTK 0 0 MSVIMILMFLLAWSPYSIVCLWSSFGDPKKIPPTMAIIAPLFAKSSTFYNPCIYVIANK 2 1 FRRAMLSMVQCQTHREITITDVLPMNRSRSPH* 0 >NEUR1_pytMol Python molarus (python) 0 MYLRQNASSQDINLPHYQRDGDPFASKLSKEADIIAGIYLLII 1 2 GIMSTLGNGYVIYMSIQRKRKLRPAEIMTVNLAVCDLGIS 1 2 VTGKPFSVIAFLSHRWIFGWSGCRWYGWAGFFFGVGSLITMTAVSLDRYFKICYLAY 1 2 GIWLKRHHAFICLGIIWSYAVFWATIPFAGLGNYAPEPFGTSCTLDWWLAQGSVAGQAFILNILFFCLVFPTAVIVFSYIKIIAKVKSSSKEVAHYDNRFQNSHELEMKLTK 0 0 VAVLICAGFLIAWIPYAAVSVWSAFGKPDSVPIKVSVVPTLLAKSAAMYNPVIYQVINCKSVCCQPEALRPLQKKNSLNKSR 2 1 VYTISTFRKSTTSAR* 0 >NEUR2_pytMol Python molarus (python) 0 MDPSFANSTFQSKITEAADIVVGTCYMVF 1 2 GVVSLFGNSLLLWVAYRKRAILKPAEFFIVNLAVSDIGMTVILFPLATPSFFAHR 2 1 WLYGKHICLFYAFCGLFFGICSLSTLTLLSVVCCLKICFPAY 1 2 GNKFSPPYAGVLLVCVWIYAFIFAAAPLADWGSYGPEPYGTACCIKWKASTREAKFYIMALFVFCYIIPCILILISYSLILWTVKVSRRAVRQHMSPQSKHNSVHSLIVK 0 0 LSISVCIGFLAAWTPYAIVAMWAAFGDPSKVPILAFALSAVFAKSSATYNPLVYLLFKPNFQKFLSKDLSFFQAVCAVCCCSRSRVITLQSFHTRDGRASMRFSTA FTDHRGSCRNCSDTFECFSNYPRCYRLTQKSDPASKTNRLAILTDGRACRPSFKRTVQVMVLMTRKKTGIGTMNVAGDVLPSNIVRDLM* 0 >NEUR4_pytMol Python molarus (python) 0 MSFQVSVQAPWRNNNMTFLNKDHPVSEQGETIIGFYLLTL 1 2 GWMSWIGNSIVIFVLYKQRAVLQPTDYLTFNLAVSDGSIAIFGYSRGIIDIFNVFQDDGFLVTSIWTCQV 0 0 VDGFLTLLFGLSSINTLTVISVTRYIKGCHPNR 1 2 GHCISTSSISVAIFLIWTAALFWSLAPFLGWGSYr 1 2 DRMYGTCEIDWTKANFSTTYKSYIVSIFICCFFMPVMVMVFSYMSIINTVKSSHALSGMGDPTDRQRRMEQSVTR 0 0 VSLVVCTCFIAAWSPYAVISMWSASGCPVPNLTSIFASLFAKSASFYNPIIYFGMSSKFRKDIAVLFHFAKEIKDPVKLKQFKILKQKLDSSPSHGEEKGAIDAQPALHSDSGVGSHPNTPPPVNRKGYFVVFDNWPQNPDFECDRL* 0
Chondrichthyes: Callorhinchus milii (elephantshark) .. 17 opsins
Six ray-finned fish genomes and massive transcript studies in yet other Clupeomorpha are available but the usefulness of this data is complicated by lineage-specific expansions and very rapid evolution of protein sequences. Little data is available from lobed-finned fish; whereas coelacanth genome has been proposed, that has stalled at 169,000 traces as of May 2008.
This makes the preliminary genome assembly of the much earlier diverging Callorhinchus (oft-misspelled) and numerous skate transcripts very special because they are the "last stop" before the very difficult lamprey genome (currently assembled but with contigs seldom larger than 1-2 exons and only rarely containing syntenic information).
This large-eyed cartilaginous fish lives to depths to 200m on the continental shelf of southern Australia and New Zealand but migrates into coastal estuaries to lay egg cases in sand and muddy substrates. The distinctively-shaped egg cases are sometimes found washed ashore after storms. They are up to 25cm long, 10cm wide, and take up to eight months to hatch. The one member of the genus studied has a vitamin A1-based photopigment with maximum absorbance at 499 nm presumably adapted to the overall photic environment at that depth.
Sequencing of the Callorhinchus genome resumed in the fall of 2008 with 454 technology; those reads are released at GenBank short read archive but are not blastable online. New Sanger 'finishing' reads from WUGSC became available at NCBI trace archives in mid-May 2009. From an exhaustive search of elephantfish data in WGS and Trace divisions of GenBank on 5 Nov 2007, many opsin exons but few complete genes were recovered and posted here. The opsin classifier reliably assigns these fragments to their ortholog class.
In Jan 2009, Davies et al reported sequencing full length RHO1, RHO2 and duplicated LWS1 and LWS2 genes in Callorhinchus, further establishing that SWS1 and SWS2 are missing in this basal chondrichthyes. That reflects a lineage-specific loss because Geotria lamprey has both.(These sequences have not yet been released from hold by GenBank but can be recovered from Fig. 3.)
Overall, Callorhinchus has good complement of vertebrate opsin genes. Parietopsin is also missing to date. Two encephalopsin- and two melanopsin-class opsins were found. The RGR, peropsin, and neuropsin genes will prove important in better determining their unresolved overall gene tree placement (which an October 2007 opsin phylogeny paper placed deeply within rhabdopsins) but more commonly classify as very basal ciliary opsins.
There are 17 Callorhinchus opsins currently available in the reference collection: RHO1, RHO2, LWS1, LWS2, PIN, PPIN, RGR1, TMTa1, TMTa2, VAOP, ENCEPH, PER1, NEUR1, NEUR2, NEUR4, MEL1b, and MEL1.
Agnatha: Petromyzon marinus (lamprey) .. 9 opsins + 2 pseudogenes
Lamprey (and less-studied hagfish) are the closest surviving outgroups to jawed vertebrates and thus central to reconstructing that last common ancestor. This importance is accentuated by the considerable temporal and evolutionary distance to the preceding two divergence nodes, cephalochordate (Branchiostoma) and urochordate (Ciona), especially for opsins because imaging eyes and advanced color vision emerged within a fairly compressed time frame within the lamprey stem.
Among extant lamprey, the genera Geotria and Petromyzon split 280-220 myr ago (helpful in breaking up the 500 myr long branch) whereas Lethenteron/Petromyzon split much later at 20 myr (helpful in locating orthologs from the former in the sequenced genome of the latter). There are no opsin sequences available for the third lamprey family, represented by Mordacia praecox, but it too is observed to have multifocal crystalline lenses that compensate for longitudinal chromatic aberration.
Photoreceptor systems have been studied extensively in both larva and adults. Geotria australis has a full complement of sequenced imaging porphyropsins LWS, SWS1, SWS2, RHO2, and RHO1 (RHO2 not so clearly a straightforward ortholog to jawed vertebrate counterparts), though its non-pineal opsin classes are not as well characterized as Petromyzon. This implies that the ancestral vertebrate possessed already possessed full photopic (bright light) cone-based color vision with the potential for pentachromacy, multi-focal lenses, pigment filtration, and likely circadian rhythm, pupillary reflex, and pineal and parapineal functions. Photoreceptor morphology and spectral sensitivity can change dramatically during various phases of the lamprey lifecycle, presumably adaptively; lineage-specific issues are not a central focus here.
An April 2009 paper directly characterizes Petromyzon imaging opsins. RHO1 and LWS, regenerated with 11-cis retinal rather than 3-dehydroretinal porphyropsin, give λmax values of 501 nm and 536 nm, respectively. Bidirectional experimental substitution show the S164P substitution accounts for the LWS blue-shift of 19 nm relative to Geotria.
This site normally waffles between serine and alanine but proline also occurs in pufferfish, turbot and japanese lamprey (4% of 94 available LWS sequences). Consequently the ancestral lamprey state was proline in the Lethentron/Petromyzon ancestor at 20 myr and S/P at the ancestral node with Geotria with serine favored because the two available chondrichthyian LWS sequences have serine. The timing is consistent with, but not proof of, a compensatory shift in Petromyzon that accompanied loss of SWS2 and/or SWS1. Six of eight lamprey GenBank genera remain unsequenced. If LWS is absent or lost from hagfish, then no earlier diverging outgroup can ever be found. It is better to view the value of each ancestral residue as a compositional vector that accommodates natural population polymorphism abundances and uncertainties in reconstruction.
The authors confirm the August 2008 report here that RHO2 is completely lost, SWS2 a very deteriorated pseudogene, and SWS1 a fairly recent pseudogene, ie the pentachromat Geotria better represents the ancestral lamprey state. Pseudogenes presumably evolve at the neutral mutation rate subsequent to the initial inactivation event, yet this could be influenced by chromosomal context and base composition issues.
However dating may work out quantitatively, it seems clear that Petromyzon passed through tetrachromat and trichromat post-Cretaceous stages separated by perhaps tens of millions of years. The loss of these opsins may not be adaptive so much as reflect 'use it or lose it'. Some advantage might accrue if the number of cones remained fixed and SWS vacancies were occupied by additional LWS, yet it is not so clear how LWS would displace suboptimal SWS in the early stages of pseudogenization.
Should lampreys be considered living fossils? The 360 mya fossil Priscomyzon riniensis closely resembles living lampreys in external morphology, large oral disc, circum-oral teeth and branchial basket. However many aspects of soft tissue anatomy are not preserved, leaving their constancy a matter of speculation. Lamprey larva being filter feeders, the question arises of what ancestral adult lamprey ate prior to the evolution of fish.
Lamprey are definitely not living fossils from the molecular point of view. Lamprey proteins evolve rapidly -- it is the norm for a cephalochordate protein to match human ortholog better than lamprey despite greatly increased branch length. This divergence of lamprey proteins is not just pointwise residue change but also involves indels and seemingly radical changes in protein character. This change could be mostly neutral relative to the ancestral lamprey state (function molecular fossil), but the fact is, Branchiostoma (sequence molecular fossil) sheds far more light on its ancestral divergence node than does Petryomyzon. At some point the ability to reconstruct ancestral genomes will be limiting for our understanding of craniate evolution.
Opsins are by no means the only fully developed gene system to have emerged in vertebrates by the time of lamprey divergence. The rod and cone cyclic nucleotide phosphodiesterases of Petromyzon consist of a single PDE6 catalytic subunit EF432251 but two inhibitory Pgamma subunits expressed in the long and short photoreceptors, EF427669 and EF470978 respectively. These are not in tandem position in the current assembly. Recently it has also become clear that the androgen receptor, GABA receptor network, and many other systems need to be backdated to agnathans.
The larvae (ammocoetes or 'sand dweller') are often said blind. It is more accurate to say larva have developmentally arrested eyes but with a small region of differentiated retina. That potentially allows for limited imaging vision -- opsin expression has not been examined. The larval stage, which can last up to 17 years, was considered a separate species by early workers. The anatomical resemblance of lamprey larva to (adult) amphioxus is quite remarkable, a case perhaps of ontogeny recapitulating phylogeny.
The metamorphosis into adult lamprey is every bit as profound as that of amphibians, involving a radical rearrangement of internal organs, continued development of eyes and transformation from sediment-dwelling filter feeder into pelagic marine vertebrate predator. Adult lampreys, like many early-diverging vertebrates, have three eyes: a pair of lateral eyes on the sides of the head and a single, unpaired median eye on top of the head. Each lateral eye is a spherical black structure lying beside the posterior prosencephalon that marks its boundary to the mesencephalon.
According to a 1936 study, larva respond to light in the green-blue by swimming avoidance, perhaps via lateral line photoreceptors. More recently Dickson and Collard, Am J Anat. 1979 Mar;154(3):321-36 write:
"Development of the retina of the ammocoete begins early in embryogenesis, with the formation of the optic vesicle, but development of the rudimentary eye is suspended and remains arrested during larval life. Prior to the onset of metamorphosis, the retina of the ammocoete is completely undifferentiated, with the exception of a small area (Zone II) surrounding the optic nerve head, where all of the adult retinal layers are found. The photoreceptors in this area have developed to include synaptic contacts as well as inner and outer segments. The pigment epithelium in this area, too, has differentiated to include well-formed melanin granules, myeloid bodies and endoplasmic reticulum and is closely associated with the receptor cell outer segments. "
"With the approach of metamorphosis, differentiation of the remainder of the retina (Zone I) begins, taking place in a radial fashion from the optic nerve head. Differentiating pigment epithelial cells adjacent to the differentiated retinal zone begin to accumulate melanin granules. In the neural retina, junctional complexes are established in the form of an external limiting membrane, and connecting cilia project into the optic ventricle. Photoreceptor differentiation begins with the formation of a mitochondria-filled ellipsoid within the inner segment."
The lamprey genome project has resulted in an anemic assembly despite accruing 19 million traces. It seems a 16 kbp retroposon has expanded enormously, which in conjunction with very high GC content and high levels of heterozygosity makes assembly of the 2.4 Gbp genome quite difficult. However the blast page at WUSTL allows a Petromyzon "3.0 supercontig" tblastn option. UCSC provided a lamprey browser In January 2008. Gene name searches work better than Blat because of very considerable divergence of protein sequences.
New sequencing technologies make affordable an immense collection of cDNA a potential solution in lamprey. It would permit a pseudo-assembly of exons flanked by some at least genomic dna. That is, transcripts are aligned into the trace archives to obtain non-coding context for exons. This would allow restricted topics to be studied (intron retention, invariant non-coding) but not others (upstream regulatory, chromosomal gene order). Labeling techniques such as FISH could provide some linkages but not gene-level order; perhaps this could be done at the level of BACs using exons from all possible gene pairs. Alternatively, the genome of Geotria might present fewer issues; it would certainly be better for the study of opsins.
Petromyzon sequences are very disorganized at GenBank. Some 147,000 conventional ESTs are stored inappropriately at the trace archives. These are attributed to WUGSC genome sequencing center but that site makes no mention of the project. Another 120,731 short ESTs from a MSU group are stored in the conventional location but the individual lamprey used may be only distantly related to that used in the genome project. Some 32,000 454-technology cDNA reads are also available but stored in the trace archives but not where they belong (short read archive SRA) but rather in the Petromyzon_marinus_OTHER database.
GenBank opsin data for Geotria and Lethenteron can be mapped into Petromyzon orthologs using its traces to obtain a supplemental set of opsin sequences parsed for intron breaks and phases. Opsin classes not available from these other lamprey can be queried using chondrichthyes counterparts, with most genes only recoverable as fragments because of gaps in coverage (rather than divergence). In all cases, intron placement remains perfectly conserved from lamprey to mammal, indeed to amphioxus with some complications.
In evolutionary accounts, imaging opsins are often claimed to have appeared abruptly in lamprey as a complete color vision set orthologous to fish and amniotes (actually far exceeding the more restricted mammalian set). It is fair to say that the lamprey opsin complement establishes that the ancestral node animal had full-blown color vision and auxiliary photoreception (ie the gene family had finished expanding on its way to seriously contracting in mammalian clades).
Had one key species of lamprey happened to have gone extinct, namely Geotria, this conclusion would be seriously weakened because RHO2 orthologs are absent to date from the Petromyzon and other lamprey; SWS1 could almost be added to this list as it is known outside of Geotria only as a recent and rapidly disappearing pseudogene, an ironic parallel to later loss of SWS1 in monotremes. Additionally, SWS2 has a much older pseudogene in Petromyzon as discussed below; its loss may have predated the divergence from Lethenteron.
It's also accurate to say complete genomes of living urochordates, cephalochordates and hemichordates (Strongylocentrotus, Saccoglossus) lack imaging opsins among their various opsin-based photoreception systems. However, parapinopsin-class opsins in Ciona today establish a much older ancestral presence of near-imaging opsins, meaning the chain of gene duplication was well along prior to lamprey stem, somewhat dissipating the supposed abruptness. The encephalopsin/TMT opsins are older still, having an unmistakable representative in the imaging eye system of cnidaria.
Branchiostoma photoreception today revolves about an expanded set of TMT/encephalopsins, melanopsins, and neuropsin-class opsins; we know nothing about what other (imaging) opsins might have been lost during its 500 myr long branch. Imagine the inferential nonsense that would emerge from using humans -- which have lost 9 of 16 genes from the vertebrate opsin complement including the descendent of the cnidarian imaging opsin -- to pontificate about photoreceptor capacities in ancestral amniotes.
The new cnidarian opsin establishes that imaging opsins have arisen at least twice out of the descending ciliary opsin gene clade of light-activated receptors. That is, TMT-class opsins are not imaging in deuterostomes as far as we know across their broad phylogenetic range in extant deuterostomes (Branchiostoma to marsupial). Evidently TMT-class opsins in cnidaria retained their potential for recruitment to imaging at least in the derived complex cubomezoan cnidaria, though here we do not know at all how basal that recruitment was and whether other cnidarian lines have lost imaging.
In summary, differentiation of deuterostome ciliary opsins (TMT/encephalopsin --> parapinopsins --> cone opsins) was dispersed over a very considerable evolutionary time frame. Note the vast majority of major deuterostome clades have gotten along just fine without cone opsins, undercutting the victim-to-predation argument with 500 myr of successful evolution. However, further intermediate details in the sequential gene expansion of cone opsins may elude us if informative additional species prove unavailable. The situation is not nearly as bad as the internode temporal gap between bird-lizard and platypus divergences, yet lies much further back in the past.
Care must be taken in discussing "vision" not to confuse low resolution with no resolution. No lensing does not equate to no image of the outside world but instead a blurry view (as reported after cataract surgery gone astray). Up/down, forward/back are already resolution seen in sponge larva. Today we view images on a color computer monitor with 3 million pixel resolution; ten years ago that was a tenth the pixels in rod opsin monotone, yet that primitive monitor image was already a big step forward from linux command lines, not to mention teletypewriter return output (no monitor at all). Newton/Gauss/Einstein lacked even those -- were they primitive evolutionary dead-ends, are we better at math today with big color monitors?
It's sometimes argued that oxygen levels in the sea rose markedly in the early Cambrian. This terminal electron acceptor allows 38 ATP to be produced from a single glucose molecule, a big energy gain over 2 ATP possible with fermentative glycolysis under anaerobic conditions. Since eukaryotes demonstrably developed aerobic pathway enzymes and mitochondria much earlier, way prior to the emergence of sponges, the discussion is really about intermediate oxygenation levels in the pre-Cambrian and the rate of oxygen consumption that it enabled for a given caloric intake relative to higher levels in Cambrian seas.
Energy-intensive processes such as swimming muscles and imaging vision (with its intensive consumption of ATP via cyclic GMP, nerve conduction, and CNS processing) may have first become feasible at some threshold of oxygen partial pressure. While accompanying caloric intake must sustain the energy drawdown, that is readily provided without predation (eg cows). We don't have an energy budget for photoreception in non-cone species such as Branchiostoma to assess whether its 7 opsins use more or less ATP than 3 retinal opsins in mammals. We don't have an energy budget for jellyfish bell-flexing swimming to compare cnidarian oxygen consumption with myotome swimming -- recall the predation direction here is often cnidarian-on-vertebrate.
A fairly recent SWS1 pseudogene is readily locatable in Petromyzon maritimus. Internal stop codons and frameshifts are not artifacts of assembly because each has multiple supporting raw trace reads but none supporting a functional reading frame. A not wholly satisfactory EST sequence EB722598 suggests, not unusually, that transcription of the pseudogene still occurs. Using Geotria as template, the coding sequence can be corrected from fragments to give a better idea of what Petromyzon SWS1 looked like prior to loss of function, though recovery of the spectrum seems like a dubious proposition.
>SWS1_petMar Petromyzon marinus (lamprey) pseudogene, sequence from genome: 7 frameshifts (^) and 2 stop codons 0 MSGDEEFYLFKNISKVGPLDGPHFHIATKWAFDFQAAFMGFVFLCG^TPLN^AIVLIVTVKCKKLRQPLTYMLVNISAAGLVFCLFSISTVFLFSTQGYFVFGPTVCALESLFGSMA 1 2 GLVTGWSLAFLAAERYIVICKPFGNFRFGSIHSLFAFCLTWVLGLGVALPPFFGWSR 2 1 YIP*GLQCSCSPDWNTVGTKYESEYCTYFLF^VFCFFVQLSIIIFSYGKLLNTL^ra^ 0 0 VAVQ^QESSLSSTQKAEREMSRMVIVMVGSFCTCYV^AALALYVVTNRDHNIDLRFVTVPAFFSKASCVYNPLIYSFMNKQ 0 0 FRARIMETVCGKFITDESETSSSRTAVSSVSTSQVSPG* 0 >SWS1_petMar Petromyzon marinus (lamprey) pseudogene, sequenced corrected for frameshifts using SWS1_geoAus template 0 MSGDEEFYLFKNISKVGPLDGPHFHIATKWAFDFQAAFMGFVFLCGTPLNAIVLIVTVKCKKLRQPLTYMLVNISAAGLVFCLFSISTVFLFSTQGYFVFGPTVCALESLFGSMA 1 2 GLVTGWSLAFLAAERYIVICKPFGNFRFGSIHSLFAFCLTWVLGLGVALPPFFGWSR 2 1 YIPeGLQCSCSPDWNTVGTKYESEYCTYFLFVFCFFVQLSIIIFSYGKLLNTLra 0 0 VAVQqQESSLSSTQKAEREMSRMVIVMVGSFCTCYVAALALYVVTNRDHNIDLRFVTVPAFFSKASCVYNPLIYSFMNKQ 0 0 FRARIMETVCGKFITDESETSSSRTAVSSVSTSQVSPG*
A very old SWS2 pseudogene can still be detected in Petromyzon maritimus either by keyword search in the lamprey genome browser or by trace archive blastn using SWS2_geoAus dna as query, in both methods verified by blastx against reference opsin sequences. Only exon 1 and exon 5 are still detectable; these lie in the same 16 kbp Contig 31930 and still retain their splice donor and acceptor locations and phases (indicating this is not processed pseudogene debris). Two further pieces of adjacent tandem debris related to exon 1 also occur in this contig. These exon fragments, when assembled onto a SWS2_geoAus template and corrected for frameshifts below, have greatly higher blastp matches to SWS2 sequences from a variety of species than to any other opsin class.
>exon1:5_SWS2_petMar Petromyzon marinus (lamprey) Contig31930:6639-6953 + very old pseudogene frameshifted assembly 0 PEDFYIPIPLNVKNLTELAPFLVPQTHLGgSGLFHAMSAFMLILITAGFPLNFLTIFLAFQYKKFRSHLNYILVNLAIANLVVVCFGSTFSFDSFINTYFVCGPLFCKMEGISATLG 1 0 TRFCMMKTTFCEKIPLVDD TRSTTMQVSCVSTSQVAPT* Top blastp matches against reference collection of 700 opsins: SWS2_geoAus Geotria australis (lamprey) Gt 0...2.1.0.0 i... 407 1.2e-41 SWS2_ornAna Ornithorhynchus anatinus (platypus) Gt 0...2... 369 1.3e-37 SWS2_utaSta Uta stansburiana (lizard) Gt 0...2.1.0.0 ind... 365 3.4e-37 SWS2_xenTro Xenopus tropicalis (frog) Gt 0...2.1.0.0 ind... 354 5.0e-36 SWS2_taeGut Taeniopygia guttata (finch) Gt 0...2.1.0.0 i... 353 6.4e-36 SWS2_neoFor Neoceratodus forsteri (lungfish) Gt 0...2.1.... 345 4.5e-35 SWS2_galGal Gallus gallus (chicken) Gt 0...2.1.0.0 indel... 337 3.2e-34 SWS2_takRub Takifugu rubripes (pufferfish) Gt 0...2.1.0.... 308 3.7e-31 SWS2_gasAcu Gasterosteus aculeatus (stickleback) Gt 0.2.... 266 1.1e-26 RHO1_petMar Petromyzon marinus (lamprey) Gt 0...2.1.0.0 ... 240 6.0e-24 RHO1_geoAus Geotria australis (lamprey) Gt 0...2.1.0.0 i... 236 1.6e-23 RHO1_letJap Lethenteron japonicum (lamprey) Gt 0...2.1.0... 231 6.5e-23
The available lamprey opsin sequence data as of August 2008 consists of 17 lamprey sequences from 3 species. Petromyzon notably has a SWS1 intronic transcribed pseudogene, apparently its only copy (Geotria has a functional copy). RHO2 and SWS2 are also missing at least at this level of coverage but again were located in Geotria by PCR. A number of opsin family members are newly reported here for Petromyzon, including neuropsin. No trace of RGR opsin or peropsin can be found in this species, despite an earlier allusion to them in L. japonicum that provided no data.
Summary of lamprey sequence data and lambda max available in the opsin reference collection: RHO1_geoAus Geotria australis full P497 AY366493 PMed:17463225 RHO1_letJap Lethenteron japonicum full P--- AB116382 PMed:15096614 RHO1_petMar Petromyzon marinus full P501 AB116382 PMed: RHO2_geoAus Geotria australis full P492 AY366494 PMed:17463225 SWS2_geoAus Geotria australis full P439 AY366492 PMed:17463225 SWS2_petMar Petromyzon maritimus pseu ---- unprocessed pseudogene SWS1_petMar Petromyzon maritimus pseu ---- transcribed unprocessed pseudogene EB722598 SWS1_geoAus Geotria australis full P359 AY366495 PMed:17463225 LWS_geoAus Geotria australis full P560 AY366491 PMed:17463225 LWS_letJap Lethenteron japonicum full P--- AB116381 PMed:15096614 LWS_petMar Petromyzon maritimus full P536 genome PMed:no_ref PPIN_petMar Petromyzon maritimus full P370 genome PMed:15096614 PPIN_letJap Lethenteron japonicum full P370 AB116380 PMed:14981504 VAOP_petMar Petromyzon marinus full P--- U90671 PMed:9427550 PARI_petMar Petromyzon marinus frag1:4 P--- genome PMed:no_ref ENCE_petMar Petromyzon marinus frag2:4 P--- genome PMed:no_ref MEL1_petMar Petromyzon marinus frag5:10 P--- genome PMed:no_ref NEUR_petMar Petromyzon marinus frag1:6 P--- genome PMed:no_ref
Chordate: Eptatretus burgeri (hagfish) .. 0 opsins
Hagfish, after decades of back-and-forth, are often sistered with lamprey, news not accepted yet by the taxonomy division of GenBank nor numerous researchers and paleontologists. Given that common phylogenetic algorithms already grossly misplace mouse within the mammalian tree, we cannot expect nuclear coding genes to perform better at 500 myr back (five times the distance). Mitochondrial genome analysis has a similar history, though hagfish was found basal to lamprey + jawed vertebrates according to a 2001 analysis.
Branchiostoma could serve as an outgroup (provided the first assembly is not used) but including radically evolving tunicates would skew results. The unfortunate lamprey assembly itself raise serious obstacles to determining topology. At this distance 28S, 18S rRNA and mitochondrial 16S rRNA may work better yet secondary and tertiary co-evolutionary considerations have not yet been incorporated. Hagfish experienced an extra round of HOX gene expansion, undercutting both HOX cluster copy number as a hallmark of supposed vertebrate body plan innovation and relentless speculation on 1R and 2R whole genome duplication in the vertebrate lineage.
Smallest lamprey genome: 1.29 Mbp, Lampetra fluviatilis Largest lamprey genome: 2.50 Mbp, Petromyzon spp. Smallest hagfish genome: 1.29 Mbp, Eptatretus cirrhatus Largest hagfish genome: 4.59 Mbp, Myxine garmani
None of the 660 nucleotide entries at GenBank as of May 2009 pertain to any component of vision, frustrating given that hagfish eye anatomy has been studied since 1886 (Krause, Die Retina der Fische. Cyclostomata. Internationale Monatschrift fur Anatomie und Histologie v3 p8-21) and opsin antibody labeling was demonstrated in Myxine glutinosa in 1984 by Vigh-Teichmann.
Jawless fish first appear 358 myr back in the Late Devonian fossil record]. Hagfish and lamprey split well after the Cambrian, roughly 430 myr ago according to much-questioned molecular clocks. That's a time span comparable to divergence of human from shark. The oldest fossil hagfishes are Late Carboniferous (330 myr). The two surviving hagfish groups split some 75 myr ago (similar to human/mouse). Only recently has it been possible to obtain hagfish eggs and embryos and revisit the neural crest issue (establishing the conventional vertebrate series of events).
Hagfish are nocturnal in aquaria and deep-sea in their natural habitat -- a new basal Eptatretus species was recently captured at a deep hydrothermal vent. Such habits do not suggest that the ancestral imaging opsin portfolio will be retained (even as recognizable pseudogenes), yet hagfish still have circadian rhythm (based in the preoptic nucleus) and dermal photoreceptors (despite no pineal gland) as well as eyes.
These non-imaging but paired eyes lack cornea, lens, vitreous body, and extrinsic eye muscle but nonetheless the retina and optic nerve react with RHO1 antibody. The eyes are larger in Eptatretus than in Myxine, where they are partly covered by the trunk musculature. However 1.3 mm is still quite small for an eye.
After comparison of all extant genera, Fernholm and Holmberg concluded in 1975 that the hagfish eyes are secondarily degenerated from more conventional eyes adapted for shallow water (for example an early lens placode disappears). However bipolar neurons are never present and photoreceptors connect directly to ganglion cells, calling homologization to protostome wiring into question. The comparative anatomy of hagfish eyes has an excellent 1977 review in The Biology of Hagfishes (JM Jorgensen), pages 542-555 available by google book search.
However this view is disputed in a wide-ranging 2008 proposal for imaging eye evolution:
"It is often suggested that the hagfish eye is degenerate, having regressed from a more lamprey-like eye that existed in the common ancestor of hagfish and lampreys. We find this view implausible. Given that the hagfish eye has survived for hundreds of millions of years, in comparison with the degeneration that has occurred in just thousands of years in cavefish. The hagfish eye must have had considerable survival value. Degeneration cannot explain how it was advantageous to hagfish for: (a) a three-layered retina to revert to two-layered; (b) the processing power of the bipolar cells to be lost; (c) reversion to a more rudimentary photoreceptor structure; (d) the disappearance of the lens, cornea, iris, and ocular muscles, all without trace; (e) re-covering of the eye by skin; and (f) re-projection of the retinal ganglion cells from tectum to the hypothalamus. It seems more parsimonious that the hagfish eye would simply have been lost..."
These authors argue further that the hagfish eye is more like a pineal gland (which hagfish lack but lamprey have) and functions in circadian rhythm rather than vision. Here we might wonder why bilateral eyes are needed with pigmented epithelial backstop giving forward and side directional capability, whereas mere light photoperiod detection would be more sensitive without it. At a minimum, paired hagfish eyes determine orientation with respect to light source (sky) and enable consistency of forward motion.
No hagfish opsins have been sequenced; no genome project is scheduled (even as the price drops in 2009 to $3,000). Even if hagfish imaging opsins are mostly gone, residual ciliary and rhabdomeric opsins could be quite informative. If so, hagfish have information about a critical stage in vertebrate imaging opsin evolution.
How will we interpret hagfish opsins as sequence data becomes available? That data ideally will come from multiple hagfish genera because gene loss can take different patterns in individual species causing the ancestral state to be underestimated. The critical issue is how eye opsins classify (root among ciliary opsins) -- that's just a quick blastp at the opsin classifier. Hagfish may well use a melanopsin in the preoptic nucleus for circadian rhythm but cannot plausibly utilize rhadomeric opsins in the eye as do protostomes.
- If hagfish diverged very early, only earlier opsins classifying to PIN, VAOP or PPIN (the likeliest of these to localize to the eye) will be found. While still ciliary in terms of their signaling transduction cascade, these are perhaps more likely in the dermal photoreceptors, which highlights the need for in situ hybridization should the data come from whole genome determination or whole animal transcripts. If confirmed as primary photoreceptor in hagfish eye, this would rule out Cyclostoma since regression from LWS etc (cone opsins) to PPIN is an unacceptable scenario.
- If hagfish diverged early, LWS is most plausible among imaging opsins given its basal position in the sequential duplication cascade of cone opsins (already finalized prior to last common ancestor of lamprey and gnathostomes). However its lambda-max may be adaptively shifted by now to a shorter wavelength better suited to dim light at depth. This outcome does not favor sistering with lamprey but does not quite rule it out either (other cone opsins could be lost). It does however require appropriately localized genes of the cis-retinal regenerative cycle such as RPE65.
- If hagfish diverged late and experienced opsin gene loss after a larger complement of opsins had evolved in a common stem ancestor, then any of LWS, SWS2, SWS1, RHO2 or RHO1 could still be present. As an example, suppose only a single opsin is found, classifying as SWS1. This implies, given the gene history, a 'lower bound' of LWS and SWS2 (ie trichromaticity at some point) having been lost but says nothing about whether RHO2 or RHO1 had ever been present in a hagfish ancestor.
- If hagfish diverged very late, the residual opsin repertoire could be either large or mostly lost. The three species of lamprey studied establish a full complement of opsins at agnathan ancestor with gnathans. These opsins include 5 imaging opsins, parietopsin, pinopsins VAOP, encephalopsin, melanopsin, and neuropsins. Lamprey TMT, RGR and peropsin appear to have been lost but may merely be too diverged to recognize. The surviving opsins in hagfish then provide clues to what opsins were lost.
It's worth reviewing the opsin classes found in the two earlier diverging clades Ciona and Branchiostoma and their tissue localization. Between extensive genome, transcript projects and expression experiments, we can be sure of having complete opsin sets and expression assignments.
- Tunicate opsins (below) include an advanced ciliary opsin classifying to PPIN expressed appropriately in its ocellus and containing the terminal VAPA* cilium targeting sequence. That implies hagfish at one point in its history had a similar opsin classifying to PPIN VAOP PIN. A second PPIN, two RGR and a highly diverged melanopsin complete the repertoire of both Ciona intestinalis and the distantly related 'congeneric' Ciona savignyi. Here all GPCR containing a lysine at Schiff base homologous position (a K-rhodopsin) have been evaluated as potential photoreceptors. Ciona has not retained the neuropsin and peropsin opsin classes.
- Branchiostoma has only basal ciliary opsins classifying to the TMT/ENCEPH group (in addition to melanopsins, neuropsins and peropsins). Its highly diverged encephalopsin lacks the fixed conserved length and VAPA targeting sequence also characteristic of the C-terminus of encephalopsins (at least from Callorhinchus on; lamprey coverage is incomplete here). While it is implausible that a PPIN opsin could have been overlooked given the extensive trace coverage reflected in the second assembly and the intense experimental targeting of opsins in B. belcheri, other cephalochordates should also be sequenced.
- While this observation could reflect loss of PPIN in amphioxus, that implies regressive replacement of PPIN by ENCEPH in the eye, far more complicated genetically than simply losing a cone opsin or two (eg Petromyzon vs Geotria). Thus the overall opsin picture strongly favors the phylogenetic tree with urochordates closer to lamprey than cephalochordates. We may be better off with such considerations because published large-scale molecular studies to date have not been definitive for chordate tree topology:
- the sea urchin and hemichordate genomes are too diverged to work as outgroups, drosophila is too far back and its proteome too derived
- Ciona assemblies have gotten worse, the KH assembly must be used
- the half-baked Oikopleura genome project indicates a very rapidly evolving and immensely diverged species
- the first Branchiostoma assembly had mediocre gene models, the second remains unannotated
- lamprey genome is a mess due to retroposons, poor exon coverage, and lack of gene models
- chondrichthyes genome is not far enough along
- teleost fish genomes are highly diverged, unfinished, and confounded by lineage-specific duplication
- orthologs are difficult to establish without synteny, complete genomes, and extensive prior gene family annotation
- mutational models used in ML are extremely dubious over this time scale and diversity of population genetics
- immense long branches have been used throughout, a risky strategy
Hagfish auxiliary genes are an important further consideration because ciliary opsins appear incapable of replenishing themselves, unlike rhabdomeric opsins which can regenerate cis-retinals in situ. RPE65, RBP3 (IRBP), and Galpha signaling factors are three interesting gene families to consider here. The first should localize to retinal pigmented epithelia, the second to interstitial matrix. and the third should cluster with vertebrate cone transducins.
- IRBP presents a timing opportunity because of the rare genomic event raising the domain count from one in Branchiostoma to four in lamprey (and all gnathans) with a concomitant radical change in exon structure and probable improvement in retinal shuttling. Unfortunately the gene seems lost in those tunicates for which there is data. No expression data is available for cephalochordate IRBP. No data is available for hagfish but if IRBP here proved structured like amphioxus rather than lamprey, that would unequivocally resolve hagfish position on the phylogenetic tree.
- RPE65 is more favorable in terms of data availability but is complicated by two additional ancestral paralogs and lacks the 'smoking gun' of IRBP non-convergent evolution. No data is available for hagfish. The amphioxus genome shows an inconvenient expansion of this gene family, not all of which can be attributed to high-polymorphism assembly artifacts. Lamprey genome is woefully incomplete in its coverage. Ciona has two paralogs with the expected intronation.
- Only GNAI sequence data (from leukocytes) is available for hagfish as of May 2009. Here hagfish eye needs to be specifically screened for imaging transducins as absence of evidence is hardly evidence of absence. This inhibitory Galpha is unsuitable for imaging opsin transduction but might work for parapinopsin; it clusters equally among vertebrate GNAi1/i2/i3 (analogously to its GATA3-like and BTK-like genes) whereas lamprey has the contemporary complement of clearly separated GNAT1 and GNAT2. Such data has been cited for wholesale gene duplication between hagfish and lamprey; observe the Galpha gene tree is, like that for imaging opsins, inconsistent with whole genome duplication and a better fit to simple tandem gene expansions during a very rapid adaptive era in eye evolution.
Urochordata: Ciona intestinalis (tunicate) .. 6 opsins
Tunicates occupy a strategic urochordate position in the phylogenetic tree as immediate outgroup to vertebrates. Three tunicate genomes have been sequenced but these proved disappointing for comparative genomics due to their derived nature, which adversely impacts coding sequence divergence, gain and loss of genes, overwriting of ancestral introns, almost total loss of gene order, and high positional heterozygosity. The situation in the seven sequenced tunicate mitochondrial genomes is also extreme.
It may not be possible to find more conservatively evolving tunicates if rapid generation time and free spawning are characteristics of all extant urochordates. Yet in other aspects, such as reconstructing the evolutionary trajectory of the vertebrate eye, contemporary tunicates may have retained critical information.
The most useful of these rogue genomes to is Ciona intestinalis -- Ciona savignyi and Oikopleura dioica have meagre transcript data and see low annual use as model organisms. PubMed abstracts mention these species in the publication ratio 886:86:46 overall and 62:8:7 for calendar year 2008.
Halocynthia roretzi has many cdna but has not been evaluated for genomic characters whereas Ciona intestinalis has been developed extensively as an experimental system; its massive cdna coverage allow better contig joining and recovery of complete coding gene models which would be nearly impossible (because of divergence and intron gain/loss) from mere homological alignment to other deuterostome proteins.
The first two assemblies of Ciona intestinalis were highly defective, having some 5,109 missing genes, faux duplications and truncated gene models (a third of the 15,254 total) with the second JGI assembly ironically worse than the first. This raises serious questions about recent papers in comparative genomics that relied on highly defective early gene sets, notably those revising urochordate taxonomic position and comparing rates of protein evolution to Branchiostoma (whose initial assembly also required serious revision).
Note however that the new Ciona assembly KH is available on 7 Nov 08 only as raw download (though a blast server with username = guest, password = welcome is available) and that the June 2008 Branchiostoma genome paper refers to v1.0, whereas the much-revised release 2.0 became available at its public blast site in Nov 2008 (gene models are not yet recalculated but simply lifted from 1.0). Obviously Ciona KH and Branchiostoma 2.0 are what need to be compared (and to a better lamprey assembly which is not even underway).
A complete set of Ciona opsin genes cannot necessarily be recovered even from the KH assembly because the article notes "it is still possible that a minor fraction of genes, such as genes expressed only under particular environmental conditions, are not covered by these ESTs. A fraction of previous models not supported by paired ESTs were excluded from the KH model set. A part of them may be real genes or unannotated fragments of genes represented by the KH models, because the encoded protein shows sequence similarity to proteins known in other species (approximately 1,641 loci with <1E-05 blast hits in the human proteome), These are provided as a supplemental model set (see Materials and methods) along with other unsupported or incompletely supported models.... probable that a minority of additional genes reside within gaps in the current assembly (48 EST-supported loci)... 47,511 ESTs (4%) were not mapped anywhere in the KH assembly... Moreover, we estimate that at least 84% of the KH transcript models contain the complete protein-coding ORF..."
Fortunately, both larval and adult photoreception have been thoroughly studied. Ciona lacks imaging eyes and thus any counterparts to rod and cone opsins (as with the cephalochordate Branchiostoma). The relative topology of these two with respect to the vertebrates has tilted in recent years towards amphioxus as immediate outgroup but the lack anything beyond encephalopsins in Branchiostoma yet a parapinopsin in tunicates argues strongly against this.
The tadpole larva CNS contains 335 cells of 13 types. These include 30 retinal photoreceptor cells in an unpaired ocellus and 5 accessory cells -- 3 for a ocellus lens-like structure, 1 for the pigment cup, and 1 pigment cell in the otolith (inconsistently with a hydrostatic sensing role for its 19 receptors). The pigment cells of the ocellus and otolith form an equipotent developmental equivalence group -- a bilateral pair of cells in the blastula gives rise to the otolith and ocellus melanocytes whereas the retina arises from both left and right cell lineages. The observed genomic complement of opsins may largely come into play in the larva because the adult is sessile with little resemblance to vertebrates. The larva are non-feeding which scarcely fits a super-predator role for early deuterostomes opsins.
An evidently ciliary opsin called Ci-opsin1 is expressed in the larval ocellus (stored here as PPINa_cioInt). The opsin classifier places this in the PPIN/PIN/VAOP group with best match 44% identity, quite respectable given a billion years of roundtrip evolutionary time. As noted initially by Kusakabe et al in 2001, this opsin shares 3 identical introns with the vertebrate group.
Today there are 25x as many opsin sequences available with much greater phylogenetic dispersion. It appears Ci-opsin has 2 new intron insertions relative to the ancestral Gq ciliary opsin 4-intron pattern 0.2.0.0. This pattern is specific (not shared by Gt melanopsins nor Go retinal isomerases) and diagnostic (disregarding a few lineage-specific gains and losses) -- see documentation.
Three ancient ciliary introns were already established at the time of amphioxus and tunicate encephalopsin divergences. Indeed they already occur in sea urchin, ragworm, mosquito, moth, and beetle ciliary opsins. Consequently they were present in the parent ciliary opsin of Ur-bilatera and no doubt Cnidaria. There's nothing surprising about this because the vast majority of (human) coding introns originated far earlier in unicellular eukaryotes and have been conserved ever since. Outside of rogue lineages such as drosophila, nematode, and tunicates, event rates for intron gain and loss are perhaps 1-2 per five billion years of branch length. Convergence is not favored because 333 aa sites x 3 intron phases = 1000 distinct possibilities in an opsin-sized protein -- for an already very rare event to happen twice in the available branch length requires predisposing factors.
We will use these deep intron characters later to supplement -- and even trump -- maximal likelihood inference from primary sequence divergence which captures the broad picture but fails to resolve the issues of most interest. With opsins, alignment (at these time depths and rates of change) hits the wall of generic rhodopsin superfamily and indeed generic GPCR proteins, which numbered many hundreds at the time of Ur-bilatera. There are already many constraints on proteins which must have seven transmembrane helical segments, covalently bind retinal with a lysine and counterion, and interact with heterotrimeric signaling protein.
With the genome in hand, we can see Ci-opsin1 has an unstudied paralog (here called PPINb_cioInt) of 58% identity and identical introns (other than a new phase 21 intron breaking exon 4). There is no expression data for the paralog in the UCSC browser track but it cannot plausibly be a pseudogene due to the conserved nature of amino acid replacements, so we wonder about subfunctionalization. The hybridization experiment will have to be repeated at various life stages. Paralog lambda max might be computable or measurable in a construct. The 1999 experiment (which measured speed-up in swimming after light decreased, reminiscent of the pineal-mediated frog tadpole response) deduced a lambda max of 505 nm -- perhaps that was a composite action spectrum. The new paralog in fact conserves the key lysine and counterion.
We can hope that photoreception in Ciona retains ancestral characteristics that descended intact, at the same time knowing evolution of protein sequences and development have not stood still for 600 million years. Ciona photoreception may have both degenerative and innovative aspects. It is premature to homologize ocellus with pineal (or amacrine or horizontal retinal cells etc) until the role of all the opsins in the Ciona genome have assigned roles (not to mention dozens of other genes). I suggest from evo-devo equipotency that the paralog opsin functions as a photoreceptor in the otolith.
Here neural integration of hydrostatic pressure signaling with brightness directionality could advantageously inform the larva of its position and orientation even in a murky water column and help with dispersal and settlement. A pigment cell is hardly needed for hydrostatic pressure sensing -- what functionality would maintain it over evolutionary timescales? The function of pigment cells is blocking light from the back, here so the larva knows up from down. Curiously, a crystallin of definite homology to refracting vertebrate lens crystallins is expressed in the otolith but not ocellus lens-like accessory cells. The statocyte itself is sprung by its footpiece and two fibrous structures, all synaptotagmin-positive. Movement of the statocyte would be detected by these three structures and thus sense gravitational orientation.
We're left wondering if the speculative otolith/photoreception connection in urochordates has any connection to the balance sensory system (vestibular apparatus) in the vertebrate brain. The otolithic organs (utricle and saccule) detect inertial movement using tiny calcium stones (otoconia) coupled to hair cells. The Allen Brain Atlas could be explored on vestibular sections for extremely detailed expression of most opsin genes. The vestibular system coordinates extensively with the visual system via the vestibulo-ocular reflex. If true, this could radically affect homologization.
Possibly this ciliary paralog pair descended from a gene duplication already present in the last common ancestor, leading after still more gene duplications to the current portfolio of vertebrate ciliary opsins. This would account for its ambivalent behavior in the Opsin Classifier with respect to the PPIN/PIN/VAOP group. Alternately the pair might represent a tunicate-specific duplication of secondary interest. Ciona savignyi has a clear ortholog (88% identity) to PPINa_cioInt but a lesser match at 59% to PPINb_cioInt, in both cases with identical introns (not an unusual pattern in gene duplications assuming PPINa_cioInt continues the original function). C. savignyi -- which is only in the same genus from a severe anthropocentric perspective -- helps gauge the rate of evolution of C. intestinalis opsins.
Photoreception in the adult ascidian, which might seem gratuitous in a sessile filter-feeder, has not been studied in quite such detail. However several non-opsin expression studies suggest that adult photoreceptors may develop about pigmented spots around oral and atrial siphons, epithelial cells of sperm duct and cerebral ganglia, involving behaviors such as siphon contraction, phototropism, and gamete release. The anterior photoreceptor of the oral siphon has even been homologized to vertebrate lateral eyes.
We'll see below that exactly the same problem as above (undocumented paralog) may affect interpretation of a comprehensive experimental study of Ciona Ci-opsin3 (RGR1_cioInt at the Opsin Classifier). Here too I was able to recover a related second gene in both C. intestinalis and C. savignyi. This illustrates the power of genomics -- provided coverage is complete, a full complement of bioinformatically extracted opsins can guide experimental design from the beginning. A full set of opsin classes should be sought in the genome, even if their degree of sequence divergence and lack of transcripts makes this difficult.
Kusakabe,Tsuda and coworkers have studied the overall visual cycle -- a much better approach than considering opsins in isolation for purposes of homologization. Recall incident photon absorption by rhodopsin isomerizes 11-cis-retinal to all-trans. Without recycling or fresh cis-retinal, this would soon exhaust vision. In mammals replenishment of the visual cycle (retinal isomerase, RGR) takes place in retinal pigment epithelial cells which are distinct from the photoreceptor cells, unlike lophotrochozoa where the cycle is completed within the photoreceptor cell. What about Ciona? We might expect a mixed system since the deuterostome divide preceded the deuterostome photoreception divide with Ciona occupying a strategic phylogenetic position.
If life were simple, Ciona would have strict 1:1 orthologs to the 4 components of the mammalian visual cycle protein, RGR (Ci-opsin3), cellular retinaldehyde-binding protein CRALBP, β-carotene monooxygenase BCO, and retinal pigment epithelium RPE65. At this phylogenetic depth, we can expect a certain degree of non-parallelism between photoreceptor systems and complications from lineage-specific duplication and subfunctionalization, not to mention lack of exact mammalian counterparts to Ciona larval and adult stages.
It turns out (using closest homologs) that Ci-BCO is predominantly expressed in larval ocellus photoreceptor cells, whereas Ci-RPE65 is not significantly expressed there nor in larval brain vesicle but rather in photoreceptor cells of the neural complex (a photoreceptor organ of the adult) right along with Ci-opsin3 and Ci-CRALBP (ie, like cephalopod). It appears the larval visual cycle uses Ci-opsin3 as restorative photoisomerase whereas the adult visual cycle Ci-RPE65. The remote paralog RGR2_cioInt was not studied and its role remains speculative. Given its degree of divergence yet persistence in a second ascidian, it is an old gene duplication maintained somewhat by selective pressure.
What about rhabdomeric opsins in Ciona? We know that melanopsin persisted into vertebrates so it must have been present at the common ancestor with ascidians. Rhabdomeres themselves as a subcellular opsin housing specialization did not persist so their apparent absence in Ciona does not imply the absence of melanopsin.
A Ciona melanopsin could be very diverged. The best possible search involves tblastn of the Ciona assembly and GenBank est_others with a variety of queries (since the best query is not known in advance; after the fact it is provided by the Opsin Classifier). Reconstructed ancestral melanopsins can improve on specific species queries by eliminated half of the roundtrip divergence.
However overly sensitive queries have the risk of merely returning generic rhodopsin-superfamily members (notably ADRA1A adrenergic receptor). While these won't receive clean approval from the Opsin Classifier, any putative melanopsin must be secondarily validated by retention of intron pattern, synteny with vertebrate melanopsins (unlikely in Ciona), and internal amino acid signatures of authentic melanopsin-type photoreceptors.
A more powerful search technique evaluates K-rhodopsins, defined as any GPCR with lysine in Schiff base homologous position (ie the lysine continues past a NAxxY motif to the YR motif at the deeply invariant length of 19 residues). In May 2009, two GPCR classifying as melanopsins were recovered here from Ciona intestinalis and Ciona savignyi. Those sequences are still rough despite three available transcripts because of divergence and a scrambled assembly in this region. The transcripts originated in blood cells and juvenile whole animal according to their annotation, leaving their role in photoreception undemonstrated.
Opsin sequences available from urochordates in Nov 2009. Renamed from original literature using the opsin classifier to place. PPINa_cioInt PPINa_cioSav ciliary NM_001032555 Ci-opsin1 ERY KTATIYNPLIYIGLNRQFR PPINb_cioInt PPINb_cioSav ciliary XM_002119927 --- ERY KTANIYNPLIYIGLNKQFR MEL1_cioInt MEL1_cioSav melanopsin --- --- DRH KASCVHSSFAYITNAHFR RGRa_cioInt RGRa_cioSav other NM_001032468 Ci-opsin3 DKY KVFVGSNPFIYIYFDPELR RGRb1_cioInt --- other NM_001032464 CiNut DRY KVISVVNPYLYMRSDPELLA RGRb2_cioInt RGRb2_cioSav other XM_002121277 --- DRY KFISVMNPYMYMRSDPELLR
Cephalochordata: Branchiostoma (amphioxus) .. 12 opsins
The phylogenetic tree of cephalochordates, based on mitochrondrial dna, shows 9 living species coalescing at 162 myr whereas the last common ancestor with vertebrates is some 400 myr earlier, thus limiting our ultimate ability to ever accurately reconstruct this node. Cephalochordates species are anthropocentrically grouped into 'genera' at divergences that applied evenhandedly to placentals would require lumping them all into a single genus.
Cephalochordate opsin data is entirely concentrated in detailed studies of Branchiostoma belcheri and bioinformatic efforts on the genomic species Branchiostoma floridae. The seven full-length B. belcheri opsin genes can be mapped into 'best-blast orthologs' in B. floridae where intron structure can be determined assuming GT-AG junctions, hopefully with accurate transfer of experimental annotation to these unstudied genes. B. floridae has 4 additional opsins whose counterparts in B. belcheri are unknown; the total of 12 opsins exceeds human by 4.
Every study to date must be repeated using the full genomic complement of 12 opsins (rather than just the initial 7). For example, it cannot be concluded when two melanopsin paralogs exist that rhabdomeric Joseph cells and pigmented dorsal ocelli (Hesse organ) photoreceptors must use one melanopsin but not the other, simply from rough action spectrum consistency. (The other melanopsin could not have endured 500 myr without contributing a function.)
The Allen Brain Atlas of mouse has exposed the great fallacy of assuming photoreceptor genes are functionally expressed only in anatomically obvious cell types. Thus frontal eye and lamellar body (pineal) for the ciliary opsins and Joseph cells and Hesse ocelli for microvillary opsins may not be the entire photoreceptor repertoire. To understand photoreception in amphioxus, it will prove necessary to revisit all life stages of all amphioxus cell types with hybridization with all 12 opsins to see what else expresses them.
Four B. floridae opsins are ciliary (same count as human), three are rhabdomeric melanopsins, and the rest peropsins and neuropsins. Lancelet ciliary opsins all classify to the basal TMT/Enceph type; no counterparts to mid-opsins (PPIN, VAOP, PIN, PARIE) or cone/rod imaging opsins occur even allowing for immense divergence. Conceivably this is attributable to genome incompleteness or lineage-specific loss; far more likely, these specialized opsin classes never evolved in cepholochordates in view of photoreceptor anatomy and phylogenetic position. Observe Ciona has an opsin classifying to PPIN, consistent with later divergence of urochordates.
The 12 cephalochordate opsins classified, with signaling class, synonymous terminology, and C2-TM7 switch; full sequences here. TMT5_braFlo Branchiostoma floridae Gt Amphiop5 ERY KSSTCYNPLVYFAMNNQFR TMTx_braFlo Branchiostoma floridae Gt ..new .. ERY KSSVVYNAAIYVAMNNQFR TMTy_braFlo Branchiostoma floridae Gt ..new .. SRY KTSYIVNTIIYLVMEKEFR ENCEPH4_braFlo Branchiostoma floridae Gt Amphiop4 ERY KSSTAYNPIIYVLMNNQFR MEL6_braFlo Branchiostoma floridae Gq Amphiop6 ERY KLSVLFNPVAYVLSIPSFR MELmop_braFlo Branchiostoma floridae Gq Amphi-mop DRY KSSAVYNPIVYAITHPKFR MELx_braFlo Branchiostoma floridae Gq ..new.. ERF KLSVLINPVAYVFSIPSFR NEUR1a_braFlo Branchiostoma floridae .. ..new .. MRF KSNSLWNPIIYLGMNERFR NEUR1b_braFlo Branchiostoma floridae .. ..new .. MRF KSSSLWNPIIYLGMNDRFS PER1_braFlo Branchiostoma floridae Go Amphiop1 YRY KSSCMMNPIIYSCCNGKFR PER2_braFlo Branchiostoma floridae Go Amphiop2 HRY KTHCAFNPILYMLMSEVYR PER3_braFlo Branchiostoma floridae Go Amphiop3 DRY KSSALYNPIIYIIANRRFR
The bioinformatics of B. floridae is very difficult, first because the individual sequenced was highly polymorphic (not inbred). This caused misassembly on a vast scale, with thousands of genes duplicated either in tandem (due to mate pair logic) or to unanchored contigs. These extra copies -- really just alleles -- can differ by several percent at the amino acid level and more within corresponding introns (due to different retroposon and indel histories). It is very difficult to identify recent true tandem duplications in this environment. The second assembly discarded one copy arbitrarily (causing cDNAs to sometimes mismatch) whereas what is needed is a diploid assembly.
Assembly 2.0 is available for blast search but its gene content had not been re-analyzed as of May 2009. The published gene count of 23,000 surely needs major downward revision. Synteny relations should still hold, though that was loosely defined under twin dubious assumptions allowing unlimited 'local' inversions and admitting weak uncurated partial homology to establish orthology. Branchiostoma genes remain uncurated; nearly all pipeline gene models are wrong in significant details. However the above set of 12 probably reflects a complete set of all K-rhodopsins in this species.
Considerable continuity can be seen to protostome invertebrates. The best ciliary opsin matches to lophotrochozoan and arthropod opsins have 38% identity, whereas melanopsins do better at 46%. To call these orthologs is not helpful (in the absence of synteny) when each clade has experienced lineage-specific gains and losses.
The TMT sequences in Branchiostoma contain an extra phase 00 intron relative to exon 2 of vertebrate counterparts; the encephalopsin has additionally an extra phase 12 intron in exon 1. Neither of these introns is seen from lamprey on, nor do they occur in lophotrochozoan TMT (which have their own novel phase 12 intron in exon 2). It is parsimonious to take vertebrate intronation as ancestral and invoke two intron gain events in cephalochordate.
None of the six available Branchiostoma ciliary opsins contain the VAPA* targeting signal. This may have evolved later -- it is first seen in the Ciona PPIN homolog and, in the gene tree, encephalopsins. Note the specialized ciliary membranes housing opsins arose far earlier as it is seen already in the protostome Platynereis (indeed its opsins are close to the signal with VAAA* in TMT1_plaDum and VAAT* in TMT2_plaDum).
Thus ciliary opsins in amphioxus seem not to have this specialization (in addition to great divergence at the sequence level). It's also unclear if any member of the expanded TMT/ENCEPH gene set has a special homology relation to the PPIN opsin in tunicates (their only ciliary opsin). Here the best match is amphioxus TMTx to Ciona PPIN but the percent identity is only 36%. Perhaps it is better to look backwards to echinoderm, yet no special affinities are observed there either (TMTPIN_stoPur and ENCEPH_strPur are its only two ciliary opsins).
A May 2009 [PNAS article provides refreshingly direct experimental evidence that the Joseph laminar body and Hesse ocelli function as photoreceptors. It provided possible to isolate individual cellular units and make observations on them. In conjunction with three previous cdna hybridization and ultrastructural detailed studies, the evidence seems overwhelming. However that seemed the case to comparative anatomists already in 1904. The issue today: if MELmop_braFlo (Amphi-mop) provides the primary photoreceptive opsin in both organs, what then is the role of the other melanopsic opsin MEL6_braFlo (Amphiop6)?
(to be continued)
Echinodermata: Strongylocentrotus purpuratus (sea urchin) .. 7 opsins
The sea urchin genome carried a big surprise: the long-belittled echinoderm has a large set of genes for sensory and signaling capability (comparable in number to human). These include seven opsins considered here.
Adult sea urchins exhibit a variety of responses to light intensity: predator avoidance, shelter seeking, covering reactions, diurnal migrations, testis response to light, and spine defense reaction to shadowing. Various pedicellariae (jaw-like appendages around the base of spines) keep the body surface clear of encrusting organisms and aid in food capture. Larva exhibit negative phototaxis and photosensitive vertical migration apparently mediated by a encephalopsin-class opsin.
Because sea urchin proteins are quite diverged from those of other deuterostomes (including sister hemichordates), it is difficult to recover accurate full-length sequences by homology alone, especially poorly conserved or short exons, in view of the so-so genome assembly. As of June 2011, only three of seven urchin opsins have any transcript support and two of these are apparent orthologs in related species, Hemicentrotus pulcherrimus and Strongylocentrotus droebachiensis. The single S. purpuratus opsin with experimental transcripts (3 fragmentary reads, no publication) is a neuropsin entirely missed by the urchin annotation effort.
The H. pulcherrimus opsin 224809166 is 95% identical to an unstudied S. purpuratus opsin TMT1_strPur (represented at GenBank by two poor quality pipeline models XM_778209 and XM_001177470, the largely correct GLEAN3_05569 stored elsewhere), enabling the experimental picture to be consolidated over in S. purpuratus.
The initial methionine remains somewhat uncertain and the DRY region is anomalous (QRC) but probably retains most motif functionality. Glycosylation occurs as in other ciliary opsins. The intron pattern 0.2.2.0.0 is a perfect match in position and phase to pinopsins. There is no detectable counterpart within the one hemichordate genome available (Saccoglossus). The best Opsin Classifier match 41% identity is to frog, TMTa_xenTro, which has a conventional ERY motif.
Expression of MEL2_strPur begins during larval development at the 14 hours postfertilization swimming blastula stage initially in cells around the tip of the archenteron in early gastrulae. By pluteus stage, cells expressing the encephalopsin are found in the aboral ectoderm but by eight-arm pluteus stage are restricted to the tips of the larval arms and posterior body. Pigment cells are present. Knockdown inhibits vertical swimming of the larvae, suggesting the opsin has a role in photosensitive larval swimming vertical migration (a functionality commonly reported in earlier diverging species). In adult urchins, the gene is expressed exclusively in tube feet.
The sea urchin genome contains a second potential ciliary opsin (ENC_strPur below). It aligns best to Branchiostoma and Platynereis ciliary opsins but only at 33% identity and not that distant to certain melanopsins. This opsin too is likely to be involved in some aspect of photoreception, though that won't advance vertebrate photoreceptor understanding.
The two peropsin-class Go urchin sequences are adjacent in parallel tandem configuration with identical intron pattern but have only 64% amino acid identity, consistent with a moderately old tandem duplication and inconsistent with assembly stutter. Despite weak percentage identity among members of this group, these are K-296 opsins quite distinct from miscellaneous 'rhodopsin' superfamily GPCRs which have (979 genes in 70 families annotated in urchin).
The two remaining opsins classify as rhabdomeric melanopsins. One of them, MEL2_strPur, has a GenBank transcript DQ285097 alluding to an unpublished expression study concerning tube feet photoreceptors. The other melanopsin is expressed post-oral arm of two-week-old larvae. Since the ancestral melanopsin has four exons, this evidently arose as a retroposed gene that retained functionality and displaced the now-vanished parental gene, an uncommon event but with numerous precedents in bilateran genomics.
cDNA Name Model Class DRY K246 Motif PubMed AB458218 ......... ............ TMT1_hemPul QRC KLCTIHNPIIYFLLNKQFK Ooka ........ Sp-Opsin1 GLEAN3_05569* TMT1_strPur QRC KLCTIHNPIIYFLLNKQFK ........ Sp-opsin2 GLEAN3_03451** ENC_strPur DRY KSSVMINPIIYAVTSRVFR ........ Sp-Opsin4 GLEAN3_22851 MEL1_strPur DRY KCSAIWNPIIYCLSHEKFN ........ Sp-opsin5 GLEAN3_06737** MEL2_strPur FRY KSSVIYNPLIYVVLNSKFR Ullrich DQ285097 ......... ............ MEL2_strDro FRY KSSVIYNPLIYAVLNSKFR Lesser ........ Sp-Opsin3.1 GLEAN3_27634 PER2a_strPur YRY KSSCMVNPIIFLTSSSKFR ........ Sp-Opsin3.2 GLEAN3_27633* PER2b_strPur YRY KSSCMINPIIFLTSSSKFR CX690664 ......... ............ NEUR_strPur LRY KTSSVYNPIIYCIFNKSFR * discrepancies vis-a-vis curated gene models provided below (typically different iMet or extraneous exons >TMT1_hemPul Hemicentrotus pulcherrimus (sea_urchin) Deut.Echi.Eleu AB458218 20067495 full G? Hp-encephalop larval vertical movement exons by homology 0 MNYSTPVMTPTASFSGSWTSTIESTAMSNLMMNIVTNVNALSGIGNETPTTLGPSSLVVPVSRSTYNYLTVYTGFLTIFGILNNGIVMVLFARFPSLRHPINSFLFNVSLSDLIISCLASPFTFASNFAGRWLFGDLGCTIYAFLVFVA 1 2 GTEQIVILAALSIQRCMLVVRPFTAQKMTHRWALFFISLTWIYSLIICLPPLFGWNHYTYEGPGT 1 2 ACSVAWNSPLPGDTSYIIFIFVMVLVIPFGIIIFCYGLLVYAVKK 0 0 ISRTQAALSSEAKADRKVSKMIFIMILFFLIAWTPYTGFSLYVTFKKNVVITPLAGTFPPFFAKLCTIHNPIIYFLLNKQ 0 0 FKDALIQLLCCGENPFDRDESEHEGRGGRHRHRTAPSATAHIGGRGRASSLPTATSMLDIPQAASTTVSSSGKTQNKENLEKGPSTSETTNERVFQLSSKVQKFEISDKNKMPSSSELPGASSLSGALMPPRRAMKNQVGCLPPVDN* 0 >TMT1_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu AAGJ03036958 full G? encephalopsin-type introns Sp-Opsin1 GLEAN3_05569 no sacKow 0 MNYSTPVMTSTASVSGPSPWTSTLESKAMSNLMTGLVTNVNALSGIGNETPTTIGLSSLVVPVSRTTYNYLTVYTGFLTIFGILNNGIVMILFARFPSLRHPINSFLFNVSLSDLIISCLASPFTFASNFAGRWLFGDLGCTLYAFLVFVA 1 2 GTEQIVILAALSIQRCMLVVRPFTAQKMTHRWALFFISLTWIYSLIICVPPLFGWNRYTYEGPGT 1 2 ACSVAWNSPSPGDTSYIIFIFVLVLVIPFGIIIFCYGLLVYAVKK 0 0 ISRTQAALSSEAKADRKVSKMIFIMILFFLIAWTPYTGFSLYVTFGKNVVITPLAGTFPPFFAKLCTIHNPIIYFLLNKQ 0 0 FKDALIQLFCCGENPFDRDESEHEGRGGRHRHRTAPSATAHIGGRGRASSLPTATSMLDIPQAASTAASSSGKTQNKESLEKGPSTSETTNKRVFELSSKIQKFEISEKNNTPSSSELPGASSLSGALMPPRRAMKNQVGCLPPVDN* 0 >ENC_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu AAGJ02133080 full G? Sp-opsin2 GLEAN3_03451 terminal exon extended to stop codon 0 MENFTSIVTDGTNEENTDGDAWPGYAHLLAGSFLTLVFIISIIGNSVVLFLFAWDRHLRTPTNMFLLSLTISDWLVTVVGIPFVTASIYAHRWLFAHVGCIS 2 1 YAFIMTFLGLNSLMSHAVIAVDRYLVITKPHF 1 2 GIVVTYPKAFLMISIPWVFSFAWAVFPLAGWGEFTYEGTGAWCSVRWDSDQPQIMSYVLAMMFLTFISSIVIMMYCYICIFLTTRRMPRWATSNSIKTHERNRRRR 2 1 EQKLLKTLIAIAIAFLVAWSPYAITSMIVVFGGSELLSLTATTLPSLFAKSSVMINPIIYAVTSRVFRKSLKK 0 0 MLTSFFPGCMTYIMTDKSPPSSSRPIQLGLCKYHFLY* 0 >MEL1_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XR_026330 frag G? Sp-Opsin4 GLEAN3_22851 no cdna losing introns larval postoral arm ancestral intron 0 MNAVTTALPHGLNKPTIEAR 2 1 WTKSLRTPPNMLIVNLAISDFGMVITNFPLMFASTIYNRWLFGDA 1 2 GCQFYAFCGALFGIMSIANMTAIALDR 2 1 YYVICWSLEAVRSVTHRRSMIIIIIVWCYAIFWSIPPFFGVGSYVLEGYGLGCTFDFMTKDLNHYLHV SFLFASSFVVPVTIIIVCFTRIAITVRAHRHELNKMRTKLTEDKDKKHKSSIRRANKAKTEFQIAKVGFQVTIFYVLSWM PYSIVAVIGQYFDSDLLTPLGTVVPVIFAKCSAIWNPIIYCLSHEKFNAALKEKLMGMCGIEIPSKHRSMGSQESSVTGR RGMHRQNSSTLSESSVTSTVDQDAIELKDRKQGPATVKVQQEKVEGGTYRRNPGDVTFSKDAGVEVDEKRRGDQGQRDDR VRPQGEGQMDQWSQPPPAPASASAPTPGVNDKEYLTKM* 0 >MEL2_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu full G? Sp-opsin5 GLEAN3_06737 tube feet 0 MPTTLMENSTPGWMADDSQMEETHPAFPLIGGYLLVVVLLGTAGNSLVIYTFLRFKKLHSPINLLIVNLSASDLLVATTG TPLSMVSSFYGRWLFGTNACAFYGFVNYYCGCISLNSLAAISVFRYIIVVRGQAQNNKLSLRSSIYAILVIHLYTLIFST PPLYGWNRFVLAGYHTSCDIDFHTKTPLFVSYICYMFFFLFFLPLGLISWSYFKIYQRVSKHSNSMRTSFTGVTKEINSDEKHA 2 1 FNHRRTASTLFVTIVVFLFAWFPYCIVSLWVLIGDANSISKLSTTIPSLFAKSSVIYNPLIYVVLNSKFRKALIQTLSFLKCLSKHELSESS* 0 >MEL2_strDro Strongylocentrotus droebachiensis (sea_urchin) Deut.Echi.Eleu DQ285097 full G? Sp-opsin5 GLEAN3_06737 tube feet retained intron WLEKMKTTQILHKPVTFLRLKPSFEPRLKPRFKPR 0 MPTTLMENSTPGWIADEGEMEETHPEFSLIGGYLLVVVLLGTAGNSLVIYTFLRFKKLHSPINLLIVNLSASDLLVATTGTPLSMVSSFYGRWLFGTNACAFYGFVNYYCGCISLNSLAAISV FRYIIIVRGKAQNNKLSLRSSIYAILVIHLYTLIFSTPPLYGWNRFVLAGYHTSCDIDFHTKTPLFVSYICYMFFFLFFLPLGLISWSYFKIYQRVSQHSNSMRTSFTGVTKETNIDEKHA 2 1 FNHRRTASTLLVTIVVFLFAWFPYCIVSLWVLIGDANSISKLSTTIPSLFAKSSVIYNPLIYAVLNSKFRKALIQTLSSIKCLYTHELSESS* >PER2a_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_778236 full Go Sp-Opsin3.1 GLEAN3_27634 RRH peropsin overshoots iMet spread across tandem 0 MAASVTESSATEAISRLEPEYMVPLTRTGYLLTAIYLTIV 1 2 GSIATVGNITVICVLCRYRTFRKRSINLLLINMAASDLGVSVAGYPLTTVSGYWGRWLFGDVGCQFYAFCVYTLSCSTISTHAAIAVYRYIYIVKTDL 1 2 RPKLTANFTSGVIVVIWVYAFFWTVTPFVGWSSYIYEPFGTSCSVNWVGRTISDISYMVACTIGVYLLQIFIMLYCYIRVAKK 2 1 IRGVDPGRTEEKDAGVVVFGRLRKREAKIDTHVTK 0 0 MCFMMMLTFIVVWAPYAVECLRAAHVHRISALSSVLPTMFAKSSCMVNPIIFLTSSSKFRQDLGKLWSRPSSQDSLQLEER NKTQRSLYVRHSELGSAHGNDTASVYYEKERIYIGEMRATSIQKEAELLQRDPELLSIASSTNSDVKFVVRDRPKRYTKR PVKPQGPRGPEMFTASGVTNKGSSTSDSGGQSTSSGTTGSKPKRSGRKASRQYSMKSQSEDTGEIFTLDGSALEMMSLRKL* 0 >PER2b_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_778236 17067569 full Go Sp-Opsin3.2 GLEAN3_27633 RRH peropsin no cdna inline tandem PER2a_strPur 0 MNSFSEESYVTDPTTTQPTLFLTPLSQTGYLLTALYLTLV 1 2 GIVSTIGNITVLCVLCRYGTFRKRSVNILLMNMAVSDLGVSVAGYPLTAISGYRGRWVFADIGCQFSGFCVYALSCSTISTHAVVAIYRYIYIVKPYH 1 2 RPRLSSSTSCLAILCIWTFTLFWTITPFFGWSSYTYEPFGTSCSINWYGKSLGDLTYIICCVVFVFILPIIIMLYCYIGVAKK 2 1 IKGIDPLRTEERDIAVVFGRLRKHETKIDTRVTK 0 0 ICFMMMASFIVVWTPYAVGSIWASKIGKISASASVLPTMFAKSSCMINPIIFLTSSSKFRADLGKLWNRPSSLEHTIRVEERSREQRSFF VRQSALPDAMVSRSASVYYDKERIYIGEMRAASIQKEADLLHRDPEAISIASSTSSSLQFVLKDRQNRYKKKAGEASKKGSNILHFPYDDTE GSMINNLMRPRSHSVTSDNISRVFAPSLKRPTKKRSMSHPDIPSTSADIFTVSPTTIKNLQKQ* 0 >NEUR_strPur Strongylocentrotus purpuratus (sea_urchin) Deut.Echi.Eleu XM_001197837 full G? CX694910 CX690664 no GLEAN3 blastula 0 MDVNAKWWTNETLRTRDQFSDDHYTSVLSYEGDIWAGVYLMFI 1 2 SLIAFIGNISVIVISLRKREKLKPIDLLTINLAIADFLICVVSYPLPMISAFRHR 0 0 WSFGKFGCVWYGFTSFLFAVGSMATLMVIALLRYAKLCRENV 1 2 DQYQSRPFVIKVIVAIWGFAFFTTAPPLFGWS 2 1 SYVPEPYHLSCTIDFADTSPSGLSYTYFTTIVVFFMPLMIIVLCYVAIARKMIHHNRRINVGHNAGRMLLEIRLLK 0 0 TACMITMAYTISWTPYAVIAMWVTYIPVNQIPDAFRILPAFCAKTSSVYNPIIYCIFNKSFRQDLSSLICCCACQCYTITINLDINSHAQQQFRRIEERR DEVGTYKRRPLMICSNPFAWSRDFHETWRQRRIRGIHRNCRNNVRVENINVNFRRDTDMVELNAPTPAEIHRPELNTASTRSGARTKSMATHLPALEEVPSG APQCSALLHNTPIPRSLQGTPLPYQPQPSTSDLHDEFLNPSVVSRNMCVIVVKPNIEEELSTD* 0
Hemichordata: Saccoglossus kowalevskii (acornworm) .. 2 peropsins
This surviving member of early branching deuterostomes has excellent genomic and transcript coverage, with diverse full length multi-exon genes often recoverable. Be aware that some transcript data has been misplaced by GenBank to reside under Saccoglossus 'other' at the trace archives while new transcription factor ESTs, accession numbers FF418995-FF534157 and FF602128-FF677500, are properly located.
However acorn worm may not illuminate photoreception at its ancestral node with echinoderms (Ambulacraria). Acornworm have isolated photoreceptive cells are scattered through the epidermis but no eyes or eye spots even in S. horsti non-phototaxic planktonic larva, as befits an animal that settles in its burrow on day two. Light striking epidermal photoreceptors elicits burrowing behavior.
In searching for opsins that might underlie epidermal (or other) photoreception, the best queries (for detecting diverged sequences) are sea urchin and amphioxus opsins versus Saccoglossus trace and est 'other' because, being transcripts rather than short exons these give longer matches. However opsins expressed in scattered cells may not be represented in transcript collections if rarely transcribed or restricted to a narrow unsampled window of development.
Promising traces must be back-blastxed against the Opsin Classifier because the initial query choice may have been sub-optimal. Good matches must be intronated using the exact-matching Saccoglossus probe against genomic trace reads because intron patterns are critical adjuncts to low-scoring sequence alignments in establishing opsin orthology classes as surviving synteny at this time depth is rare and not currently determinable.
I report here the very first hemichordate opsins. Although quite diverged, these classify cleanly as peropsins, a class that appears significantly expanded in Saccoglossus. Here again the Schiff base lysine is present but the nature of covalent ligand (if any) is by no means established as retinal. While peropsins might accomplish the photoreceptive task in this species (and so may represent yet another approach to evolving photosensors), they cannot serve as ancestral sequence to deuterostome ciliary opsins (which are well-represented in sister taxon echinoderms).
Hence ciliary opsins have been lost in Saccoglossus. Whatever its other merits for understanding body plan or centralized nervous system origins, this species cannot clarify why ciliary opsins were retained in early deuterostomes prior to the evolution of imaging vision. It's not clear whether additional enteropneust or pterobranch hemichordate species would improve this situation -- the 90 known are all benthic marine.
The process for obtaining accurate gene models in such a remote species prior to assembly is a difficult exercise in bioinformatics and use of the Opsin Classifier -- a detailed procedure is given in the annotation tricks section. A final check consists of back-blast of the final sequence against all of genBank to verify top hits are deuterostome peropsins. This cannot detect chimeric proteins (unbridgeable fragments) however.
A complete set of opsins for this species became feasible with the long-overdue release of the 7x assembled contigs by the Baylor sequencing center on 04 AUG 2009. This enabled some minor fixes in the two previously recovered peropsin sequences (which can now be seen to be identically intronated and 34% identical) but did not result in any additional opsins.
>PER1a_sacKol Saccoglossus kowalevskii (acornworm) 7 ests + ACQM01133041 0 MVTTDSLANSTDEPVPSILTLQQHYAASVTLLAL 1 2 AVIGTVLSSVNFRMLLSNPDYCSKAGNFFLSLAVTDL 1 2 CVCIFETPFSAFSHHAGFWIFGDTACQLYAFFGIFFGLVNIFMVTFISLDRYWATCSPVE 1 2 VELKSKYYTRMTALGWMVALFWAAAPVFGWSRYaMEPSMASCSIDYMTNDFSYVTYITCLTLTCYVVPIVVMVYCYVKASKNIKYTGKVTEWAHENNATK 0 0 ISRLCVLQLVFCWSLYGFNCMWTVVADDVETLPKMLTVLAPILAKTTPILNSGLYFLHNKKFRGAAVDMFKAKEE* 0 >PER1b_sacKol Saccoglossus kowalevskii (acornworm) ACQM01067921 2 aVLSVIGNSVVLEMFRRYKELLSPSAILLISLALADL 1 2 GLTIFGMSLSCVSSFAGRWLFGKFGCYFHGFAGMLFGLGSIGNLTVISIDRYIITCKRsL 1 2 WSYRHYYALLAVAWSNALFWSMMPLFGWSSYALEPEGTSCTIDWMNNDNQYISYVSCVTVTCFILPCAVMTYDYLAAYMKMVKAGYTLSEETEKPNND 0 0 MCIALVAAFLLSWFPSATVFLWAAFGNPGNIPLSFTGVADAFTKIPAVFNPVIYVALNPEFRKYFGKTIGCRRKRKKPIAVRLNGSEQNVENTI* 0
Deuterostomia: Xenoturbella bocki + Convoluta pulchra .. 0 opsins
These two taxa have recently been put forward as new phyla of basal deuterostomes, the former as outgroup to echinoderms plus hemichordates, the latter acoel flatworms as more basal still. However sequence data is extremely sparse with 3,127 sequences for all of Acoelomorpha, and Convoluta pulchra evolving far too fast for practical use, with its tree position controversial.
No genome or major transcript studies are under consideration. A quick check via tblastn of sea urchin opsins against available transcripts does not turn up good opsin candidates as of 28 Nov 2007 (other than a weak melanopsin match in Convoluta, EV602614, that might instead be generic GPCR). No information about photoreception in these species is readily available. While the above two taxa might not be ideal for opsin purposes, extant species are very limited.
See also: Curated Sequences | Cnidaria | Ecdysozoa | Lophotrochozoa | Update Blog