Opsin evolution: LWS PhyloSNPs: Difference between revisions
Tomemerald (talk | contribs) |
Tomemerald (talk | contribs) No edit summary |
||
| Line 53: | Line 53: | ||
One speculative scenario: the block tandomly duplicated giving rise to an ancestral SWS1 with its own (soon-to-diverge) upstream copy of regulatory element called here SWS1:LCR. After protein divergence (that explains the closer relation of SWS2 to SWS1 than to LWS), the SWS1 gene but not its regulatory element duplicated tandomly downstream to give SWS1:LCR SWS1 SWS2 LWS:LCR LWS. A later chromosomal rearrangement broke off the now self-sufficient SWS1 from the cluster. This scenario then repeated itself, but with a twist as SWS2 alone duplicated internally, gave rise to upstream RHO2:LCR and RHO2. RHO2 later gave rise to RHO1 which is targeted still to retina but to rod cells. This event may have taken place separately post-lamprey giving confusing RHO2 comparisons. This scenario distinctively predicts regulatory elements upstream for LWS and SWS1 but downstream/shared for SWS2 and upstream of RHO2 and RHO1 coding. | One speculative scenario: the block tandomly duplicated giving rise to an ancestral SWS1 with its own (soon-to-diverge) upstream copy of regulatory element called here SWS1:LCR. After protein divergence (that explains the closer relation of SWS2 to SWS1 than to LWS), the SWS1 gene but not its regulatory element duplicated tandomly downstream to give SWS1:LCR SWS1 SWS2 LWS:LCR LWS. A later chromosomal rearrangement broke off the now self-sufficient SWS1 from the cluster. This scenario then repeated itself, but with a twist as SWS2 alone duplicated internally, gave rise to upstream RHO2:LCR and RHO2. RHO2 later gave rise to RHO1 which is targeted still to retina but to rod cells. This event may have taken place separately post-lamprey giving confusing RHO2 comparisons. This scenario distinctively predicts regulatory elements upstream for LWS and SWS1 but downstream/shared for SWS2 and upstream of RHO2 and RHO1 coding. | ||
Proposed sequence of events giving rise to vertebrate visual opsins explaining opsin gene tree and chromosomal proximities. | |||
Key issues: SWS2 closer to LWS in gene tree yet SWS1 syntenic with LWS and using its control region | |||
Status of chrX locus Other loci Comment | |||
SWS1 | LCR-LWS original gene targeted to cone | ||
SWS1:LCR SWS1 | <span style="color: #0066CC;">LCR-LWS</span> LCR-LWS first duplication: LWS and regulatory element | ||
LCR-SWS1 LCR-LWS SWS1 attributes emerge | |||
LCR-SWS1 <span style="color: #0066CC;">LCR-SWS1</span> LCR-LWS second duplication: just SWS1 and its LCR | |||
LCR-SWS1 LCR-SWS2 LCR-LWS SWS2 attributes emerge in middle gene | |||
LCR-SWS1 LCR-LWS LCR-SWS1 translocation of SWS1 and its LCR to an autosomal chr | |||
LCR-SWS1 <span style="color: #0066CC;">SWS1</span> LCR-LWS LCR-SWS1 third duplication: SWS1 but not its control region | |||
LCR-RH02 SWS1 LCR-LWS LCR-SWS1 RHO2 attributes emerge, SWS1 transcribed by LWS control region | |||
SWS1 LCR-LWS LCR-SWS1 LCR-RH02 translocation of SWS1 and its LCR to new autosomal chr | |||
SWS1 LCR-LWS LCR-SWS1 LCR-RH02 <span style="color: #0066CC;">LCR-RH01</span> fourth duplication:RHO2, emergence and translocation of RHO1 | |||
SWS1 LCR-LWS <span style="color: #0066CC;">MWS</span> LCR-SWS1 LCR-RH02 LCR-RH01 fifth duplication: LWS in OW primates but not control region | |||
</ | |||
In this scenario, the regulatory elements diverge paralogously from their common ancestor just like the corresponding opsin genes. Note the history of descent differs slightly from that of the gene tree in that SWS2:LCR descends with RHO2 as SWS2 has no regulatory element of its own. Testing this would require finding these paralogous regulatory elements and establishing comparative genomics nesting. | In this scenario, the regulatory elements diverge paralogously from their common ancestor just like the corresponding opsin genes. Note the history of descent differs slightly from that of the gene tree in that SWS2:LCR descends with RHO2 as SWS2 has no regulatory element of its own. Testing this would require finding these paralogous regulatory elements and establishing comparative genomics nesting. | ||
Revision as of 16:16, 17 August 2008
Comparative Genomics of LWS opsins
The LWS (long wave sensitive) cone opsin gene is especially favorable for comparative genomics because numerous pre-existing studies on specific species and the happenstance of NISC targeting have greatly expanded the depth of phylogenetic coverage available from the 50 genomic projects and accidental EST coverage, bringing the total coverage close to 100 species of vertebrates. This affords the opportunity to examine the robustness of phyloSNPs deduced just from genomic projects to a doubling of sampling density.
It is quickly apparent however that GenBank sequences cannot be used at face value; they are error-ridden, with more than a handful of entries having spurious (disabling) indels and highly implausible regional bursts of radical amino acid substitutions. In some cases, multiple independent sequences allow confident correction of error, though it must be noted that transcript and genomic projects rarely reference the same individual animal (meaning some polymorphism can be expected). Because no central physical depository for dna exists, it may not be possible to ever resequence many of the anomalous entries.
GenBank entries for LWS genes display a bizarre range of haphzard descriptors (green opsin, red opsin, red-green opsin, red-sensitive pigment, red-green sensitive, medium-wave-sensitive, long-wave-sensitive, iodopsin, LWSA, M/L, MW, LW, M/LWS, KFH-R , cp-L, AYU-R, LWS_QUEm5_L06, color blindness deutan, color blindness protan), with less than half the entries including appropriate terms such LWS or OPN1LW that would allow simple database retrieval. GenBank has been roundly criticized by 256 signatories for its refusal to allow corrections.
Less than 5% of the entries mention the peak adsorption (lambda max) even when that was experimentally determined and indeed the whole purpose for sequencing. In some cases, the abstract provides the critical data; in other cases, access to proprietary text is needed; in still others, no article was ever published.
Consequently, the GenBank annotation -- if not the sequence itself -- is often worthless. It is necessary to begin from scratch with a fixed 'type specimen' and expand that out to a core set of LWS genes with syntenically verified orthology. That was done here earlier in building the original opsin classifier. Each putative new LWS ortholog must then unequivocally cluster with the core set distinctly from all other (cone) opsin gene family members. There are various complexities at the level of the individual species that are not the focus here, such as triallelic species. LWS has experienced a fair number of lineage-specific tandem and whole-genome duplications (notably in fish), pseudogenization, and gene losses (new here: Xenarthra).
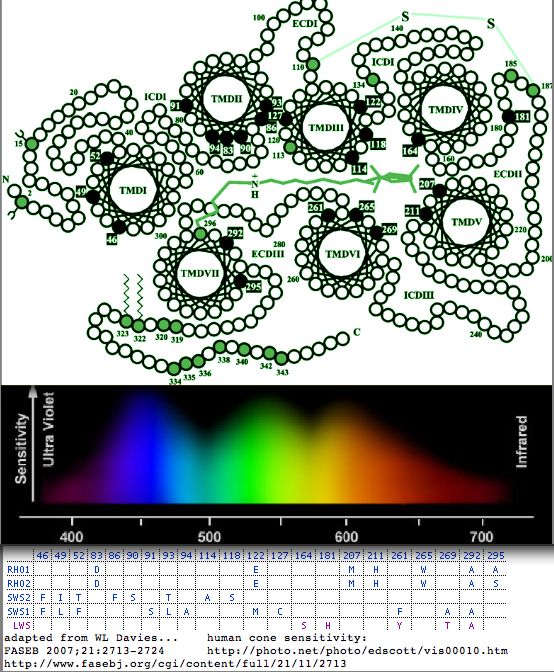
Then, using the "five sites rule" of Yokoyama, as subsequently amended and refined, the peak adsorption can be rather reliably predicted using S180A, H197Y, Y277F, T285A, and A308S as shifting toward blue by 7, 28, 8, 15, and 27 nm from a base level of P560. These are not quite additive as the double change S180A/H197Y shifts lambda max by 11 nm.
This could be revisited today in view of a vastly expanded data set. Using the above offsets, lambda max in the curated set of LWS genes below range from 505 in eriEur (shrew) to 560 in sciCar (squirrel). These tuning residues potentialy explain the highly restricted reduced alphabet at these positions (with an alternatives observed only at S180P in lampreys and two fish), despite the greatly expanded the species set. These 5 residues are shown in the alignment markup track below which are easily extracted for spreadsheet analysis without distraction from extraneous residues. Spectra for all lamprey cone opsins other than LWS (P560 predicted) have been experimentally measured.
It would be necessary to comb the literature for all available experimentally determined wavelengths prior to re-parameterization. So many species have been directly measured that predictions for residual species scarcely add value. Here it must be noted that chromophores such as vitamin A2 are used in some species (perhaps at some mixing level) and impact actual lambda max whereas reconstituted proteins can be measured with reference to vitamin A1. Ancestral spectra need recalculation at each divergence node with today's much higher species sampling density and experimental testing in full length ancestral protein reconstitutions.
The diagram shows the location of residues tuning lambda max in the five imaging opsin gene classes. It could be improved by considering the full reduced alphabets seen in high density sampling and instituting a cutoff for significant change as ultimately nearly every substitution at nearly every central position will have some effect however small on the chromophore environment. Further, a shift in one opsin should not be considered in isolation but rather in the context of the overall visionary system together with the photic environment experienced by the animal.
More ambitiously, all opsins might be considered for a comprehensive prediction program. The paralogous reduced alphabets at the five sites can vary quite considerably in non-cone opsins, for example in peropsins, neuropsins, and rgropsins. Here it is not so clear whether these five residues are still the principal tuning residues (as might be expected from structural conservation) nor what the functional significance of lambda max shifts might be in non-imaging opsins.
In the LWS alignment below, it can be seen that variations the five tuning positions are phylogenetically incoherent (not phyloSNPs). Note though however coherence within Glires (rodents + rabbits) of the tyrosine at position 197. It's not clear what the specific adaptive value of this 28 nm blueshift might be; perhaps further sampling would dissipate the apparent coherence.
Locus control region (LCR) between SWS2 and LWS
SWS2 and LWS occur as a tandem pair ancestrally, as implied by a conserved association from teleost fish through amniotes and monotremes (note SWS2 has been deleted in the stem to marsupials and placentals). A cis-acting locus control regulatory region, analogous to that of the beta hemoglobin cluster, sits between the two genes and regulates both. It was not affected by the SWS2 deletion event; indeed, as LWS duplicated downstream in old world primates, the LWS:LCR region extended its reach to regulate this gene as well. Color imaging opsin genes must be regulated so that they are expressed in the retina only and only in cone cells there often in a certain gradient patter.
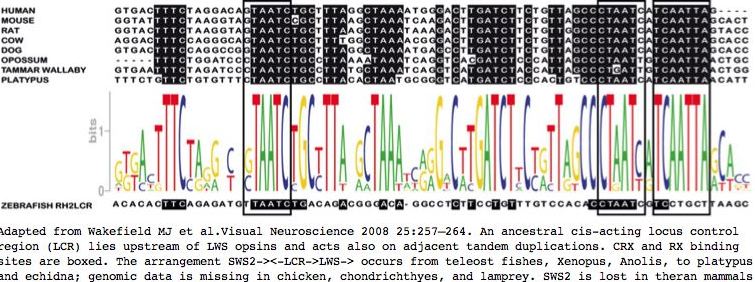
Here, the LWS:LCR region is recovered from 35 species ranging from human to lamprey, expanding on recent work of MJ Wakefield et al. The group of mammalian LWS:LCR sequences are clearly orthologous, based on experimental evidence, high percentage identity, syntenic location and orientation with respect to LWS strand, uniqueness as Blat matches within whole genomes, and genomic alignment provided in the UCSC 28-species alignment (where LWS:LCR is a significant phastCons element). By the same criteria, the sequences from teleost fish, frog, and lizard are also orthologous.
Lamprey also is 80% identical to the consensus sequence from this group and retains conserved nucleotides, but the genome assembly has gaps in this region and the anticipated intermediate position in lamprey SWS2-LWS cannot be validated. This could be better pursued in the Geotria genome which has a full complement of opsins.
The evidence is weaker that the two groups of LWS:LCR sequences are orthologous. Ordinary alignment is not convincing in part because the sequences are only 70 bp to begin with and the time of group divergence is 310 myr and more. However both sets of sequences are positioned similarly as recognized phastCons elements downstream of SWS2 and upstream of LWS. Experimental data in human, mouse, frog, and zfish support similar cis-acting roles. Further, the lizard comparative genomics track method recovers the platypus LWS:LCR sequence at the expected place relative to lizard, frog, and fish LWS:LCR elements.
Thus it appears the LWS:LCR regulatory element arose prior to lamprey divergence and has retained its position upstream of LWS in all descendent clades, yet signicant evolutionary changes occured in the earliest mammals (conceivably associated with Walls' postulated nocturnal era in conjunction with loss of SWS2 in therans). Additional sequences relevent to this transitional era (eg more early amniotes, echidna, more marsupials) may illuminate these changes but more likely insufficient extant species exist.
LWS occupies a special basal position in the vertebrate imaging opsin gene tree -- a subsequent duplication cascade gave rise to the other opsins. This raises the question of whether the ancestral LWS:LCR regulatory element also propagated to these other opsins which, except for SWS1, are at unrelated chromosomal positions. Since these opsins retained the fine details of exon structure, the duplication events were not retropositioning and reintronation, ie were some form of block duplication that may well have brought the LWS:LCR regulatory element along because otherwise the new gene could not be plausibly targeted to retinal cone cells prior to acquiring disabling mutations.
Note that the opsin gene tree (as deduced independently in dozens of publications) directly conflicts with the sistering tree topology expected from supposed whole genome duplications. That is, the chain of duplication is LWS -> SWS1 -> SWS2 -> RHO2 -> RHO1. Why then is SWS2 rather than SWS1 adjacent to LWS?
A key experimental observation here is tandem duplications of LWS still fall under the control of the LWS:LCR regulatory element even though that element itself may not have been duplicated. This provides a temporal window of retained functionality during which the spectal properties of the new gene can diverge with selective advantage, for example to peak sensitivity at shorter wavelengths. We know such events have been fixed at least twice (the SWS2 event pre-lamprey and the MLS event in primates).
One speculative scenario: the block tandomly duplicated giving rise to an ancestral SWS1 with its own (soon-to-diverge) upstream copy of regulatory element called here SWS1:LCR. After protein divergence (that explains the closer relation of SWS2 to SWS1 than to LWS), the SWS1 gene but not its regulatory element duplicated tandomly downstream to give SWS1:LCR SWS1 SWS2 LWS:LCR LWS. A later chromosomal rearrangement broke off the now self-sufficient SWS1 from the cluster. This scenario then repeated itself, but with a twist as SWS2 alone duplicated internally, gave rise to upstream RHO2:LCR and RHO2. RHO2 later gave rise to RHO1 which is targeted still to retina but to rod cells. This event may have taken place separately post-lamprey giving confusing RHO2 comparisons. This scenario distinctively predicts regulatory elements upstream for LWS and SWS1 but downstream/shared for SWS2 and upstream of RHO2 and RHO1 coding.
Proposed sequence of events giving rise to vertebrate visual opsins explaining opsin gene tree and chromosomal proximities.
Key issues: SWS2 closer to LWS in gene tree yet SWS1 syntenic with LWS and using its control region
Status of chrX locus Other loci Comment
LCR-LWS original gene targeted to cone
LCR-LWS LCR-LWS first duplication: LWS and regulatory element
LCR-SWS1 LCR-LWS SWS1 attributes emerge
LCR-SWS1 LCR-SWS1 LCR-LWS second duplication: just SWS1 and its LCR
LCR-SWS1 LCR-SWS2 LCR-LWS SWS2 attributes emerge in middle gene
LCR-SWS1 LCR-LWS LCR-SWS1 translocation of SWS1 and its LCR to an autosomal chr
LCR-SWS1 SWS1 LCR-LWS LCR-SWS1 third duplication: SWS1 but not its control region
LCR-RH02 SWS1 LCR-LWS LCR-SWS1 RHO2 attributes emerge, SWS1 transcribed by LWS control region
SWS1 LCR-LWS LCR-SWS1 LCR-RH02 translocation of SWS1 and its LCR to new autosomal chr
SWS1 LCR-LWS LCR-SWS1 LCR-RH02 LCR-RH01 fourth duplication:RHO2, emergence and translocation of RHO1
SWS1 LCR-LWS MWS LCR-SWS1 LCR-RH02 LCR-RH01 fifth duplication: LWS in OW primates but not control region
In this scenario, the regulatory elements diverge paralogously from their common ancestor just like the corresponding opsin genes. Note the history of descent differs slightly from that of the gene tree in that SWS2:LCR descends with RHO2 as SWS2 has no regulatory element of its own. Testing this would require finding these paralogous regulatory elements and establishing comparative genomics nesting.
This raises the question of where the regulatory element first arose in progenitors to LWS, namely pinopsin (a gene restricted to frog through bird expressed in pineal), VAOP (fish through bird), parapinopsin (tunicates to bird), and parietopsin. None of these genes were retained in mammal, so the bioinformatics of finding homologs to LCR are unfavorable. As the retina arose prior to lamprey divergence, gene duplicates destined to provide imaging opsins needed to be targeted to emerging cone cells, even as parent genes retained expression in pineal. Thus the regulatory circuits here must already have been well-established 540 myr ago in the Cambrian.
A potential paralog RHO2:LCR has been studied experimentally in the zebrafish tetrameric tandem. Here an upstream RH2:LCR element controls the four downstream opsin genes consistent with the scenario non-vacuously in that regulatory elements can occur separated, upstream, or downstream of their targeted genes. Unfortunately, that element has no bioinformatically detectable homologous counterparts even in other species of fish with determined genomes; the opsin itself is lost in mammals.
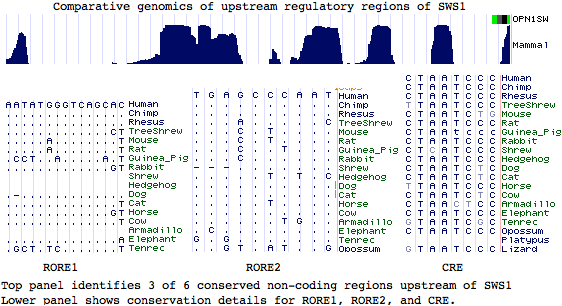
Similarly, while upstream regulatory regions of SWS1 have been experimentally studied, including a putative CRX binding site (LCR), there is no basis for infering homology to LWS control regions. Indeed sites of best sequence conservation, in the form of phastCons features from the 28-species alignment, correlate most imperfectly with the short CRX and orphan nuclear receptor RORB binding motifs (consensus CTAATC and AGGTCA respectively). The 3 unassigned peaks of conservation could correspond to as yet undescribed transcription regulatory proteins.
It is highly implausible that a small block translocation of a coding gene would evolve a de novo regulatory element from adjacent junk dna (even the small CRX and RX motifs together with 39 bp dimer spacing are a formidable barrier) or land near a pre-existing appropriate regulatory element (that of an adjacent unrelated gene targeted to retina cones at appropriate developmental window) even though element sharing can occur.
35 LCR control region sequences and 2 consensus sequences >LWS:LCR_homSap Homo sapiens (human) tttctaggacagtaatctgctttaggctaaaatgggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_ponPyg Pongo pygmaeus (orang_sumatran) tttctaggacagtaatctgctttaggctaaaatgggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_macMul Macaca mulatta (rhesus) tttccaggacagtaatctgctttaggctaaaatgggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_calJac Callithrix jacchus (marmoset) tttctaggacagtaatctgctttaggctaaaatgggacttgatcttctgttagccctaatcatcaGttagc >LWS:LCR_otoGar Otolemur garnettii (bushbaby) tttctagaacagtaatctgcttgaggctaaaatgggacttgatcttctgttagccccaatccttgattag >LWS:LCR_tupBel Tupaia belangeri (tree_shrew) ttttgagggcagtaatctgctttgggctaaaaggggagctgatctgtggttagccctaatcatcaattagc >LWS:LCR_musMus Mus musculus (mouse) tttctaaggtagtaatccgctttaagctaaatcaagacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_ratNor Rattus norvegicus (rat) tttctaaggtagtaatctgctttaagctaaataaagacttgatcttctgttggccctaatcatcaattagt >LWS:LCR_speTri Spermophilus tridecemlineatus (squirrel) gttttaggacaataatctgctttaggctaaatcaggatttgatcttctgttagccctaatcatcgattagc >LWS:LCR_dipOrd Dipodomys ordii (kangaroo_rat) ttccagcacagtaatctgctttgagctaaattgggacttgatcttctgctagccataatcatctcttaac >LWS:LCR_oryCun Oryctolagus cuniculus (rabbit) tttctagggcagtaatctgctttaggctaaaatgggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_ochPri Ochotona princeps (pika) tttctagggcagtaatctgctttgggctaaaatgggacttgagcttgtgttagccctaatcatcaattagc >LWS:LCR_canFam Canis familiaris (dog) tttccaggccggtaatctgctttaggctaaaatgagccttgatcttctgttagccctaatcatcaattagc >LWS:LCR_felCat Felis catus (cat) ttccagggcagtaatctgctttaggctaaaatgagacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_bosTau Bos taurus (cow) ttccagggcagtaatctgcttttggctaaaacgggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_equCab Equus caballus (horse) ttccagggcagtaatctgctttcagctaaaacgggacctgatcttctgttagccctaatcatcaattagc >LWS:LCR_myoLuc Myotis lucifugus (microbat) tttccagggcagtaatctgctttctgctaaaacaggacttgatcttctgttagccctaatcatcaattagc >LWS:LCR_pteVam Pteropus vampyrus (macrobat) 4 bp del tttacagagcagtaatctgctttctgttaag gacttgatcttttgttagccctaatcttcaattagc >LWS:LCR_turTru Tursiops truncatus (dolphin) tttccagggcagtaatctgcttttggctaaaactggacatgatcttctgttagccctaatcatcaattagc >LWS:LCR_eriEur Erinaceus europaeus (hedgehog) ttcca ggcagtaatctgctttcagctaaaacaggacttgatctttttctagccctaatcatcaattagc >LWS:LCR_sorAra Sorex araneus (shrew) tttct gctgtaatctgctttcggctaaaacgggacttgatcttcggttaaccctaatcatcaattagg >LWS:LCR_loxAfr Loxodonta africana (elephant) tttctagggcagtaatctgctttaggctaaactgggacttgatcttctgttagccctaatcatcaatta >LWS:LCR_proCap Procavia capensis (hyrax) tttccaggacagtaatctgctttaggctaaactgggacttgatcttctgttagccctaatcatcaatta >LWS:LCR_echTel Echinops telfairi (tenrec) tgtctagggcagtaatctacttttggctaaaccgggacttgatcttctgttagccctaatcatcgattatc >LWS:LCR_monDom Monodelphis domestica (opossum) tttctggatccctaatctgccttaaaataaatcaggtcacgatctcccattagccctaattgtcaattaac >LWS:LCR_macEug Macropus eugenii (wallaby) tttctagatccctaatctgccttatgctaaatcaggtcatgatctaccattagccctgattgtcaattaac >LWS:LCR_ornAna Ornithorhynchus anatinus (platypus) gttctgtgtttctaatctgccttacactaatgcgggtcatgatctcccactgtccctaatcatcaattaac >LWS:LCR_mamCon mammalian consenus sequence tTTctaggacagTAATCTGCtTTaggcTAAaatggGactTGATCTtctgtTagCCCTAATCaTCaATTAgc >LWS:LCR_taeGut Taeniopygia guttata (finch) 86% identical; from anoCar probe gggcgcgcggccgagttctgccagtagtgagctttt ttccaggaaaaagttcagaactagcggaattcc >LWS:LCR_anoCar Anolis carolinensis (lizard) gagcgcgcggccgagttctgctagtagtgaactttttctaactgtaaatagttcagaactagcggaattcctgac >LWS:LCR_xenTro Xenopus tropicalis (frog) catg removed ccgagttctgctagtagtgaacttttacttactaa gaatagttcagaactagcggaattcctgacaagcac >LWS:LCR_danRer Danio rerio (zebrafish) attcagttgtatatccgagttctgctagtagtgaacttttgccaatcacgaatagttcagaactagcggaattccaggcatacgactgact >LWS:LCR_tetNig Tetraodon nigroviridis (pufferfish) ctcgacggt agtcgggagcaaatctgagttctgctagtagtgaacttttaccaaacatgaatagttcagaactagcggaattccagacctgctctcgaaa >LWS:LCR_takRub Takifugu rubripes (fugu) ctcgacggt agtcgggagcaaatctgagttctgctagtagtgaacttttaccaaacatgaatagttcagaactagcggaattccagacctgccctcgaaa >LWS:LCR_gasAcu Gasterosteus aculeatus (stickleback) ctcggcggtagtcgggagcaaatctgagttctgctagtagtgaacttttaccaatcccgaatagttcagaactagcggaattccagacctaccctcgaaa >LWS:LCR_oryLap Oryzias latipes (medaka) cccgtcggc agtcgggagcaaatccgagttctgctagtagtgaactttttccaagcatgaatagttcagaactagcggaattccagacctactctcgaaa >LWS:LCR_petMar Petromyzon marinus (lamprey) traces 1.2e-06 80% gcgacgtaactaCGCGACTGAGTTCTGCTCGTAGTGAACTTGTACGGAGTGGAAA GTTCAAAACTCGCGGAATTCCTGCCGGGCA >LWS:LCR_telCon fish-lizard-bird consenus sequence CcGAGTTCTGCtAGTAGTGAaCTTTTactaactatgAAtAGTTCAGAACTAGCGGAATTCCagac
PhyloSNPs in LWS opsins: alignment of 94 LWS vertebrate opsins
.position ...................................................................................................1.........1.........1.........1.........1.........1.........1.........1.........1..1 .position .........1.........2.........3.........4.........5.........6.........7.........8.........9.........0.........1.........2.........3.........4.........5.........6.........7.........8..8 .position 123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123 .....exon 111111111111111111111111111111111111122222222222222222222222222222222222222222222222222222222222222222222222222222222222222222222222222233333333333333333333333333333333333333333333333 ..markers .................................gly.......................................................................................diS.cI....................ERY...........................P-7. 01.homSap MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTRGPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLCGITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWIWAAVW 02.panTro ...................................................................................................................S....................................L.................I........S... 03.gorGor ....................................................................................................................................................................................... 04.ponPyg ................................................................T..................................................S.........................................................V...V..... 05.nomLeu ................................................................T..................................................S....................................L.................I........S... 06.macMul ...................................................................................................................S.........................................................V...V..... 07.macFas .............................................................................................................................................................................V...V..... 08.papHam ........................................................................................................................................................L..........................S... 09.cal539 .................N..........I...............................L...V.................................................I.....................................L.............................. 10.cal553 .................N..........I...............................L...V.......................................................................................L.............................. 11.cal561 .................N..........................................L...V...............................I..................H....-...............................L...................V......S... 12.ceb530 .................N..........................................V...F..........V......................................I.....................................L...................V.......... 13.ceb545 .................N..........................................V...V..........V......................-...........I...I.....................................L........M..........V.......... 14.ceb560 .................N..........................................V...F...............................I..................H....................................L...................V......S... 15.ate560 ..........................V.....N.A.........................V...T.................................-...............F.....................................L...................V......S... 16.ate552 ..........................V.....N.A.........................L..............V........................T..I..........F.....................................L...................V......S... 17.tarSyr ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,.....L...........V..I.........T...........R.......................I..........I.........L...........,,,,,,,,,,,,,,,,,,,,,,,,,,,,......A............ 18.otoGar .....GP...T..Q...TH.....G.........T......................A.........I...........R.......................I..........I.........L...........................L........................V..... 19.otoCra .....GP...T..Q...TH.....G.........T......................A.........I...........R.......................I..........I.........L...........................L........................V..... 20.micMur .....GPH.F..GQ....H.....A.........A......................A.................V...R................I......I..........I.........L...........................L........M........M..T...V..... 21.tupBel ...R.GPHK...GQ..................T.....................I..M.....................R................I.................IF...I................................L.................A............ 22.musMus ...,,,,,,.T.EQTL.H.....HA...........K....................T...L..V..............R.......................I..........I.........L..I...I....................L................T...V...V...I. 23.ratNor ...,,,,,,.T.EQTL.H......A................................T...L.................R.......................I..........I.........L..I...I....................L................T...V...V..... 24.nanEhr .....AP.....GQT.........A.L.............................TT...L..V..I.......V...R.......................I..........I.........L..I.........................................TM..T.A.V..... 25.speTri ...R.DP.....GQ....H...............A...................I..I...I.............V....................I.................FF........L..V......V.................L......................A.T.S... 26.sciCar ,..R.DP.....GQ....H...............A...................I..T...I.............V....................I.................L.........L..V......V.................L........M.................S... 27.dipOrd ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,...................TI..M..............V...R.......................I..........FN........L.IV........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 28.cavPor ...R.GPHA.S.VQA..A......A.L.......N......................A..TI.....I.......V...R............................................L..V........................L....................V...V.S... 29.oryCun .T.P.GP.M...GQ.PE.H.....A...................F............A...L.............V...R..................................F.........L..V........................L.................A............ 30.canFam .T.R.GP.....GQ..AGL.E...A.........A..D...................A.................V...R................I......I..........I.........L..V........................L.................A............ 31.phoVit ...T.GP....DGR..PG......A.........A......................T.....................R.......................I......I...I.........L..V.................................................V.S.L. 32.lepWed ,,,,,,,,,,,,,,,,,,,,,,..A.........A...................F..I.....................R.......................I......I...IH........L..V.................................................V.S.L. 33.mirAng ,,,,,,,,,,,,,,,,,,,,,,..A.........A...A..................I.....................R.......................I......I...IH........L..V.................................................V.S.L. 34.eriBar ,,,,,,,,,,,,,,,,,,,,,,..A.........A......................T.....................R.......................I..........IH........L..V.................................................V.S.L. 35.zalCal ,,,,,,,,,,,,,,,,,,,,,,..A.........A......................I.....................R.......................I..........I.........L..V........................L.................A......V.S.L. 36.odoRos ,,,,,,,,,,,,,,,,,,,,,,..A.........A......................T.....................R.......................I..........I.........L..V........................L.................A..V...V.S.L. 37.ursMar ,,,,,,,,,,,,,,,,,,,,,,..A.........A......................A.....................R..................................I.........L..V........................L.................A......V..... 38.enhLut ,,,,,,,,,,,,,,,,,,,,,,..A.........A......................T......F..............R.......................I..........I.........L.............................................A........S... 39.felCat .T.R.GP.....GQ.HAGL....RA.........A...................V..A.............................................I..........I.....................................L.................A............ 40.bosTau ..HA.GP.....GQ..ANF.E...G............D...................A..V..................R................I......I..........M.........L..V..........................................T............ 41.odoVir ..HE.GP.....GQ..ANF.E...G............D...................A..V..................R................I...V..I..........M.........L..V..........................................A........-... 42.turTru .....GP..F..GQ..T.F.....G.V..........D..D...................V..L...I....P.................M.......................M.........L.IV..F.......................................A............ 43.delDel ...T.GP..F..GQ..T.F.....G.V..........D..D...................V..L...I....P.................M.......................M.........L.IV..F.......................................A........-... 44.gloMel .....GP..F..GQ..T.F.....G.V..........D..D...................V..L...I....P.................M.......................M.........L.IV..F.................................................... 45.susScr .....GPR....GQ..A.F.....G.......A....D................I..A..V..................R................I......I..........M.........L..V..........................................A............ 46.equCab ...R.GP.K...GQ..AGF.....A.......N.A..D................V..A.....................R.......................I..........I............V.........................................VA............ 47.myoLuc ...R.GP.....GQL.A.F....LA.........A......................A..V..............V...R.......................L..........I.........L..V........................L........M........A..T...V.S... 48.myoVel ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,-----.V........................L.................A..T...V.S... 49.pteVam ...S.VP....EGQ..A.F.E...A...I.....V......................A.................V...R.......................I.......A..I.........L..V........................L........M.......T.......V..... 50.pteDas ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,..V........................L........M.......T.......V..... 51.hapFis ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,..V........................L............T...T.......V..... 52.sorAra ....,,,,,,,,,,,,RGP.....H....F....A...................V..A......F..............R.......................L....................L...........................L.......SM...............V.S.I. 53.eriEur ...REGP.....VQ...GL.S..LG.T...SG....I..........T...AF..A.A..V...F..............R..............................L...MF......Y.V....C......................L.......SL.....M..L........S... 54.loxAfr .....GPH..T.ARL..AS.....A...V...T.T......................A..L...L..........V...R..................................I....I....L..V........................L.................A..........I. 55.triMan .....GTH..T.GQ...A......A...V...T.T.........H............S..L...M..LL.........VR....................L.............IH...I....L...........................L..........................S.I. 56.echTel ...R.GAH..T.GQL..T..G..RT...V....T...............P.......T......F..........V...R.......................I..........I.....F...L...........................L....................T...V.S.I. 57.proCap ..EP.GTH..T..QLP.AS.....A...V.....A......................A......L..........V...R..................................I....I................................L.................A............ 58.monDom .T.A.DPAGFLA.RRD.DNDET.R..L.V.....N..................N...L..V......I.......V........................G..........I..I....I....L...........................V.........K......M...I...V..... 59.macEug .T.A.DPAGFLAWR-R.EN.ET.RA.L.V.....N.K...............FN...L.........I.......V........................G..L.......I..I....I................................V.........K......M...V...V..... 60.setBra .T.A.DPAGFLAWR-R.EN.ET.RA.L.V.....N.K...............FN...L.........I.......V........................G..M.......I..I....I................................V.........K......M...V...V..... 61.cerCon .T.A.DPAGFLAWQ-E.EN.ET.RA.L.V.....N.K...............FN...L..V......I.......V....................I...G..I.......I..I....I................................V.........K......M...V...V...I. 62.tarRos .T.A.DPAGFLAWR-R.EN.ET.RA.L.V.....N.................FN...L..V......I.......V........................G..I.......I..I....I................................V.........K......M...V...V.-.I. 63.isoObe .T.A.DPAGFLAWR-R.EN.ET.RA.L.V.....N..................N...F..-..............V........................G..I.......I..I....I................................V.........K......M...V...V..... 64.myrFas .T.A.DPAGFLAWRREEN-.ET.RA.L.......N.K................N...F.................V........................G..I.......I..I....I................................V.........K......M...V...V..... 65.ornAna .TPA.NSGVY.A.RRFEDE..T.RT.V.V.....N..D.............A.NV..L.................V........................G..L.......I..IF...I........................S.......I.........K......M...V...V..... 66.tacAcu .T.A.DPAGFLAWR-R.EN.ET.RA.L.V.....N.................FN...L..V......I.......V........................G..I.......I..I....I....L...........................V.........K......M...I...V.-... 67.galGal .-AA.EAA-F.A.RRHEE-..T.RD.V.......N..................N...L......A..........V..W.....................G..........I..IS...I.......V......A....A............F........IK..G...VA..L...L.SCA. 68.taeGut .-AT.DGAVF.A.RRH.D-..T.RD.........N..................N...L......V..........V..A.....................G.............IF...I.......I......A....A............F........IK..G...VA.VL.....SCA. 69.serCar ,,,,,,,,,,,,,,,,,,,,,,,,,,,.......N..................N...L......V..........V..A..............K.S..E.G.............IF...I.......I......A....A............F........IK..G...VA.VL.....SCA. 70.anoCar ..EA.DVAVF.A.RRN.E-D.T.RD.L.......N..................NI............I.......V..A.................I...G..........I..IS...I..............T...SA.....V......V.........K......VA..V...V.S... 71.gecGec .TEA.NVAVF.A.RSR.D-D.T.RG.V.....T.N..................N.V.FF..I.....C.......V..A.................FV..V..LV......F..IF...I....L..I...V..S.................F........IK..S....I..V...V..WG. 72.pheMad .TEA.NVAVF.A.RSR.DDD.T.RG.V....D..N.K...D............NF..F..VI.....T.......V..V.................FV.MV......G...I..IF...I....L..I...L..A.................F........IK..S....L.........WG. 73.utaSta ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,...A............V.........K......MA..V...V.SC.. 74.ambTig ..HS.NSGAY.A.RRY.D-..T.R..........N..................N..TL......F..........V....................I...G..........I..MF...I....L..I......V...SA....T..A....F........IK..G...AA..I...V.S.G. 75.cynPyr ..YS.NSGAY.A.RRY.D-..T.R..V.V.....N..................N..TL..F...A..........V....................I..I...L.......I..IF...I....L..I......V.........T..A....F........IK..G...AA..I...A...F. 76.xenTro ..SH.NEAVF.A.RRN.D-D.T.R..V.......N..................NIS.L......L..........V..L...............M.I...G..........C..IF..........I.......V...AA....TV.A....F........IK..G...AT..I...V...G. 77.xenLae ..S.LNEAIF.A.RRN.D-D.T.R..V.......N..................N...I......F.........IV..L...............M.I...G..........F..IF...I..........F...T....A....TV.A....F........IK..E...AT..I...V.S.G. 78.neoFor ..EP.D-AV..A.RRHQD-.ET.R.T..V.....N..................N...L......F..C.......M..Y.................I...G..L.......T..IF...I......M...F..AT.........T..A....V........IK..G.W.AG..I...V.S.F. 79.danRer ..EH.GDAIY.A.RKG---DET.REAM.......N.KD...............NVAT...F...V..T.......V..A.................I...G..LF......I..FF...I......IF......V...AA....TV......V.........K....W.SA..I...V...A. 80.cypCar ..E..GDAIF.A.RRG---DET.RETM.V.....N..D...............N.AT...F......T.......V..A.................I......LL......I..IF...I......IF......V...A.....TV......V.........K....W.SA..I...V.S.F. 81.salSal ..ER.GSAAY.A.RQN---Q.T.RE.S..F....N.KD...............N.STL..VI...L.........V..A.....Q...........I..IG..LL......C..FF...I.......F......V...AA.....V......V.......S.K....W.MG..I...V...F. 82.pleAlt .TDE.GNAVF.A.RRN---..T.RE.S.......N.KD...............NISTM.................V..A.....Q...........I...G...L......C..TF...I.......F......T...AA....TV......V.........K....W.TA..V...V.S.A. 83.tetNig ..EE.GK.SF.A.RYH---....RG.A.V.....H..D...............N.ATL..F...VL.........V..A.....................G..LF......C..FF...I......IF...V..V...A.....T.......I.........K....W.TA..V.....PIC. 84.takRub ..EE.GK.SF.A.RYH---..T.RG.A.V.....H..D...............NVAT...FI..VL.........V..A.................I...G...F......C..FF...I.......F......T...AA....T.......V.........K....W.TG..V...V..... 85.gasAcu ..EE.GK.AF.A.RYN---..T.RG.M.V.....N.KD...............N.STL..FI..AL.........V..A.....Q...........I...G...F......C..FF...I.......F...V..V....A....T.......I.........K....W.TA..V.....S... 86.oryLat ..E..GK.VF.A.RQN---..T.RG.A.......H..D...............N.ATL..F...VL.........V..A.............S...I...G...F......C..FF...I.......F...V..T...AA....T.......V.........K....W..G..V...V.S... 87.poeRet ..EE.GK.VF.A.R-H---..T.RGAA.......H.KD...............NVSTL..CI..VL.D.......V..A.................I...G...F......C..FF...I.......F...V..T...AA....T.......I.........K....W.TA..V...A..... 88.lucGoo ..E..EK.AF.A.RYN---.ET.RG.V.......H..D...............NVST...F...VL.........V..A..E..............I...G...F......C..FF...I.......F..FI..T...AA....T.......I.........K....W.TA..V...V.S... 89.ophVen ..EE.GK.SF.A.RYH---....RG.A.A.....N..D.............I.NVATL..FV..VL.........V....................I...G...F......C..FF...I......IF...V..V...AA....T.......I.........K....W.TA..V...V.S... 90.pseMax ..ED.GKPAF.A.RYH---..T.RG.A.M.....H.KD.............I.N.ATL..FV..V..........V..A.................I...G...F......C..FF...I.......F......V...A.....T.......I........IK....W.TG..L.....PI.. 91.calMil ,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,D.............A.N......VG..V..........V..VR..............M.L...G...L...V..A..FF...I....L..F..FV.......A....T..A....V.........K..G.W.AF..I...V.SIG. 92.petMar .TAS.QGAMF.A.RR..D-..T.ME.L.R...E.N.KD..............FN.......I..VL.L.S.....V..V............I....I..IL..IF......C.......I.......F...V..T...A......................IK..G.I.TIL.V...V.P.S. 93.letJap .TAS.HGAVF.A.RRN.D-..T.KD...R...E.N..D.............MFN.......I..VL.L.......V....................I..IL..IF......C...F...I.......F...V..T...A......................IK..G.I..IL.V...V.P.C. 94.geoAus ...S.ERAMF.A.RR..--..T.KGDL.R...E.N..D.............M.N...F...I...L.L.......V..L.................I..IG..IF...V.....IF...I....L..F..F...V....A........F............LK..G.V...L.I...A.S.G. .position 1.....1.........2.........2.........2.........2.........2.........2.........2.........2.........2.........2.........3........3..........3.........3.........3.........3.........3....3 .position 8.....9.........0.........1.........2.........3.........4.........5.........6.........7.........8.........9.........0........1..........2.........3.........4.........5.........6....5 .position 45678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345678901234567890123456789012345 .....exon 33333333334444444444444444444444444444444444444444444444444444444555555555555555555555555555555555555555555555555555555555555555555555555555555556666666666666666666666666666666666666 .topology NNNNNNNNNeeeeeeeeeeeeeeeeeeeeeeeeeeMMMMMMMMMMMMMMMMMMMMMMMMMMMMccccccccccccccccccccccNNNNNNNNNNNNNNNNNNNNNNNNeeeeeeeeMMMMMMMMMMMMMMMMMMMMMMMMMccccccccccccccccccccccccccccccccccccccc* ..markers .............P28.diS.........................................................................P-8.....P-15...............P27.....K..................................................... 01.homSap TAPPIFGWSRYWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLSIIVLCYLQVWLAIRAVAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQFRNCILQLFGKKVDDGSELSSASKTEVSSVS--SVSPA 02.panTro ..............................................I..A..............................................................................................................................--..... 03.gorGor ................................................................................................................................................................................--..... 04.ponPyg ................................................................................................................................................................................--..... 05.nomLeu ..............................................I..A..............................................................................................................................--..... 06.macMul .................................................T........................................F..Y.V.....T............A..........Y..........I.......................................--..... 07.macFas .................................................T.....................................M..IF.Y.V.....T............A..........Y..........I.......................................--..... 08.papHam .................................................T........................................F.......................A..........Y..........I.......................................--..... 09.cal539 .........................................I....L...........................................IA.Y.V.....T............A..........Y..........I.......................................--..... 10.cal553 .........................................I...FL...........................................IA.Y.V.....T............A..........Y..........I.......................................--..... 11.cal561 .........................................I...FL..G........................................IV.Y.V.....T............A..........Y..........I.......................................--..... 12.ceb530 .........................................I....L...........................................IVT..V..................A..........Y..........I.......................................--..... 13.ceb545 .........................................I....L...........................................I.T..V.....T............A..........Y..........I.......................................--..... 14.ceb560 .........................................I...FL..G........................................IM.Y.V.....T............A..........Y..........I.......................................--..... 15.ate560 .............................D...K...........FL..G........................................IM.Y.V.....T............A..........Y..................................................--..... 16.ate552 .............................D................L...........................................IM...I.....T........K...A..........Y..................................................--..... 17.tarSyr .................................................T.........................................F.Y.L.....T........H...A....V.....Y..........I..........................V............--..... 18.otoGar .....................................M.......VI.....M........................................Y.V.............SH...A....V.....Y..........I.......................S.....T.........--..... 19.otoCra .....................................M.......VI.....M........................................Y.V.............SH...A....V.....Y..........I.......................S.....T.........--..... 20.micMur ......................................M.......I..............................................Y.V..R..T........H...A..........Y..........I.............................T.........--..... 21.tupBel ..............................................I............................................C.Y.V.....T........H...A....L.....Y..........I.......................S.......R..A....--..... 22.musMus .............Y.............T.........M........F............................................F.Y.L.....T......T.H...A....V.S..SY..........I..............H........S.....T.........--..... 23.ratNor .............Y.............T.........M........F............................................F.Y.L.....T......T.H...A....V.S..SY..........I.......................S.....T.........--..... 24.nanEhr ....V........Y.............T.........V...I....I.....L...........T...................H......F.Y.L.....T..V...T.H...A....V.....Y..........I.........Q.............S.....T....A....--..... 25.speTri .............Y.............N.........M........I.....I........................................Y.L......S.....T.....A....V..V..Y..................................T.......R..A....--..... 26.sciCar .............Y.............T....M....M........I.....I......................................F.Y.L.....T......T.....A....V.....Y..........I,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 27.dipOrd ,,,,,,,,,,...Y.............T.........M........I.....................L......................F.Y.V............T.H...A....V....SY..........I.......................S.....T....A....--..... 28.cavPor .............Y.............T.........M...................H...................................Y.L............T.....S....V.....Y..........I.....................E.S.....T.R..A....--..... 29.oryCun .............Y.............T.........M........I...V.........M..P...........................F.Y.L.....T......T.H...S....V..I.SY..........I.....................E.S.......R..A....--..... 30.canFam .........................................T....I...V.I........................................Y.L.....T........H...A....V.....Y..........I.......................S.......R..A....--..... 31.phoVit .........................................T....I..GV.I......................................F...V.....T........H...A....V.....Y..........I.......................S..........A....--..... 32.lepWed .........................................T....I..GV.I.....................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 33.mirAng .........................................T....I..GV.I..................D,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 34.eriBar .........................................T....I..GV.I.....................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 35.zalCal .........................................T....I...V.I....................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 36.odoRos .........................................T....I...V.I.......................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 37.ursMar .........................................T....I...V.I.....................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 38.enhLut .........................................I....I.....I.......................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 39.felCat .........................................I....I...V....................................M..IF.Y.V.....T........H...A....V.....Y..........I.............M.................R..A....--..... 40.bosTau .........................................I...FI...V.I..................................M..IF.Y.L.....T........H...A....V.....Y..........I.......................S.....V....A....--..... 41.odoVir .........................................I...FI...V....................................M..IF...L........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 42.turTru ......................................T..I...FI...V.........................R..........M..IF.Y.L.....T........H...A....V....SY..........I.......................S.....V....A....--..... 43.delDel ......................................T..I...FI...VT........................R.V........M..IF.Y.L.....T........H...A....V....SY..........I.......................S.....V....A....--..... 44.gloMel ......................................T..I...FI...V.........................R..........M..IF.Y.L.....T........H...A....V....SY..........I.......................S.....V....A....--..T.. 45.susScr .........................................I...FI..GV....................................M..IF.Y.L.....T........H...A....V.....Y..........I.......................S.....V....A....--..... 46.equCab .........................................I....I...V....................................M...F...L.....T........H...A....V.....Y..........I.......................S.....V....A....--..... 47.myoLuc .........................................T....I...V.......................................I..Y.L.....T........H...A....V.....Y..........I...................R...S.....T.R..A....--..... 48.myoVel .........................................T....I...V.......................................I..Y.L.....T........H...A....V.....Y..........I........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 49.pteVam ..............................................I...V.LF.......................................Y.L.....T........H...A....V.....Y..........I.......................S.....S....A....--..... 50.pteDas ..............................................I...V.LF.......................................Y.L.....T........H...A....V.....Y..........I........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 51.hapFis ..............................................I...V..F.......................................Y.L.....T........H...A....V.....Y..........I........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 52.sorAra ...........................D.............I....I..GV.LF.................................M.....Y.V.....T........H...A....V.....Y..........I.............................V..S.AT...--..... 53.eriEur .............................D.......M........I......F...H...................................Y.V.....T.S.S....H...T....L.T..SY...................,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 54.loxAfr .........................................T....I..........H................................I..Y.L.....T......T.H...A..........Y..........I.......................S..........A....--..... 55.triMan .....................................M...I....I..G.....-.H.............................L..I..Y.......T........H...A....V.....Y..........I..................................A....--..... 56.echTel .................................R.......I....I........................................M..IF.Y.L.....TI.......H...S..........Y..................................S.....T.R..A....--..... 57.proCap ................................A........I....I..........H.............................M..IF.Y.V.....T........H...A..........Y..........I.......................S.....T..M.A..A.--..... 58.monDom ....L........................D...........A....F.....L...V..........................S......I..Y.......TL...........S....T.S...Y..........I..........T...............V..T.R.......--..A.. 59.macEug ....L......................N.D...........S....L...V.F...I.......S..................S......I...........I...........A....T.S...Y..........I..........T...............V..T.R.......--..A.. 60.setBra ....L......................N.D...........S....L...V.L...I..........................S......I..Y........I...........A....T.S...Y..........I..........T...............V..T.R.......--..PA. 61.cerCon .S..L......................N.D..I........S....L.....L...I..........................S......I..Y.......T............A....T.S...Y..........I..........T...............V..T.R.......--..A.. 62.tarRos .S..L......................N.D..I........S....L.....L...V...R......................S......I..Y.......TL...........A....T.S...Y..........I..........T...............V..T.R.......--..A.. 63.isoObe .............................D...........T....L.....L...V..............D...........S......IR.Y.......TL...........A....T.S...Y..........I..........T...............V.GT.R.......--.APA 64.myrFas .............................D...........S....L...V.L...I..........................S......I..Y........I...........A....T.S...Y..........I..........T...............V..T.R.......--..A.. 65.ornAna .............................D...........S....L....................................S......I..Y.......TI...........A....A.....Y..........I.............M...............T.R.......--..... 66.tacAcu .S..L........................D...........A....F.....L...I..........................S......I..Y.......TL...........A....T.S...Y..........I..........T...............V..T.........--..A.. 67.galGal .............................D.......V.......FF..A..I......S.......A...............S......IV.Y.......T............A....A.....Y..........I..........................V.-T.R.......NS..... 68.taeGut ............................TD....-..V.......FF..AV.IF.............A...............S......I..Y.......TI...........A....T................I..........................V.-T.R.......NS..... 69.serCar ...........................TTD.......V.......FF..AV.IF.............A...............S......I..Y.......TI..S......-.A....T................I........,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,, 70.anoCar ....V.......................DD...L.......I...FI..AV.L..............A...............S......II.Y.......TV...........A....A.....Y..........I.............M...............T.R.......NS..... 71.gecGec S...........................VEL.C..F.LT..I...FL..F..IV......M......A............R..S......IV...I......S.VS........A....A.....Y........................M............A.TT.R.......NS..A.. 72.pheMad S..........................NNEL.C....LA...S..FF...V.I.......M......A......................L....V.....TI.VG....H...A....A.....Y.......W.....I....................A.DV.TT.R.......NS..... 73.utaSta ............................DD...........I...FI..AV.I.......M......A............R..S......II.Y.......TI...........A....A.....Y..........I.............................T.R.......NS..... 74.ambTig C............................D.......M...I....L.....II..I...W...Q..M............R..S......II.YI......T............A....A.S..............I.............Y............M....R.......NS..... 75.cynPyr C..........................N.D.....F..T..S....L.....I...V...W...Q..M............R..S......IM.YI......T..V.........S....A.S...Y..........I.............Y...............S.R.......NS..... 76.xenTro C............................D.......L...I....I..A......MH...T..Q..Q............R..S......II.YI......T.......F....N....A..M..Y..........I.............Y............V..T.R.......NS..... 77.xenLae C...M.....F..................D.......L...I....I..A..I....H..WT..Q..Q............R..S......IV.YI......T.......FS...S....A.....Y..........I.............Y.M..........V..T.R.......NS..... 78.neoFor C.M.L.....F................EDKY.TR.F..A..I....I..GV.I...I...W...T..................S......IF.Y.......T.M...G..Y...A....A.....Y..........I.............Y..L............T.........NS..... 79.danRer C...........................ED.......V...I....I..A..I...IA.Y...H...Q...D...........S......IF.Y.......T............A....A..M..Y.....................V..M............V.-T.......-----.A.. 80.cypCar C.........F.................ED...........I....I..A..I...IA.........Q...D...........S......IF.Y.......T............A....A..M..Y..........I..........V..M............V.-T.......-----.A.. 81.salSal C........................G.NED...K....T..I...FF..FV.IF..IF.........A...D...........S......II.Y.V.....TC...........A....A..I..Y.....................T..M.....AE...T.V.-T.......-----.A.. 82.pleAlt C...........................DD...K.......I...FL..A..I...IA.........Q...D...........S......I..YIV.....TV...........A....A.....Y.....................V..M............V.-T.......-----.A.. 83.tetNig C...........................ED...........I....I..A.......A..M......M............R..S......I..Y.V.....T............A....A..M..Y..........I..........V..MK....E......V.-T.......-----.A.. 84.takRub C...........................ED...........I....I..A..I....A......S..M...............S......IV.Y.V.....T............A....A..M..Y.....................V..MK....E......V.-T.......-----.A.. 85.gasAcu C...........................ED...........I...LI..A..I....A.........M............RD.S......IV.YIV.....TT...........A....A..M..Y.....................S..M.....E......V.-T.......-----.A.. 86.oryLat C...V.......................DD...........I....I..A..I....A.........M............R..S......IV.Y.V.....T............A....A..M..Y.....................T..M.....Q......V.-T.......-----.A.. 87.poeRet C...........................DD...L.......I....I..A..I....A.........M............R..S......II.Y.V.....T.....P......A....A..M..Y.....................T..M.....Q....F.V.-T.......-----.A.. 88.lucGoo C...........................ED.......V...I....I..A..I....A.........M............R.GS......I..Y.V.....T............A....A..M..Y...................S.T..M.....Q......V.-T.......-----.A.. 89.ophVen C...........................ED...........I....L..AV.I....A.........M............R..S......IV.Y.V.....T............A....A..M..Y..........I..........T..M.....Q......V.-T.......-----.A.. 90.pseMax C...........................ED...............FL..A..I....A..W..HS..L.............D.S......I..Y.V.....T............A....A..M..Y..........I..........T..M.....E......V.-T.......-----.A.. 91.calMil CL..V.....,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,,.....S.......A...L......C..M.S.L....A....V.SI.SY....S.....I.............................T...D.....NS..... 92.petMar CSL.........................TD.......V...I...FL.....I..........HS..Q......T.....RD.S......I..YV......T............S...IA.....Y...G......I..........................V..S.R.......NS..... 93.letJap CSL..........................D.......V.......FL...V.I..........HS..Q......T.....RD.S......I..YI......T....Y.......A....T.....Y........................M............V....R.......NS.I... 94.geoAus C...........................TD.......V...I...FI..AL.II.........HT..Q......T.....RD.S......IF.YI......T............A....A.....Y..........I.............M............V..SAR.......NS.....
Curated LWS reference sequences from 95 species
>LWS_homSap Homo sapiens (human) -IRAK1 -MECP2 -TEX28 +TKTL1 P530 full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_panTro Pan troglodytes (chimp) odd full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 gPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVSGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIIGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLAIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_gorGor Gorilla gorilla (gorilla) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ponPyg Pongo pygmaeus (orang_sumatran) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVTASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVSGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_nomLeu Nomascus leucogenys (gibbon) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVTASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVSGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIIGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLAIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAFCFCWGPYAFFACFAAANPGYPFHPLMAALPAFFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_macMul Macaca mulatta (rhesus) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVSGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLTIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMFLAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_macFas Macaca fascicularis (macaque) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLTIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* >LWS_papHam Papio hamadras (baboon) full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLTIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMFLAFCFCWGPYAFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_cal539 Callithrix jacchus (marmoset) P539 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFIYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMLFVVVASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQIYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCILPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIAAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_cal553 Callithrix jacchus (marmoset) P553 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFIYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMLFVVVASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFLPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIAAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_cal561 Callithrix jacchus (marmoset) P561 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSVWMLFVVVASVFTNGLVLAATMKFKKLRHPLNWILVNLAIADLAETVIASTISVVNQVHGYFVGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGVAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFLPLGIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIVAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ceb530 Cebus capucinus P530 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFTYTNSNSTR 1 2 gPFEGPNYHIAPRWVYHLTSVWMVFVVFASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLAETVIASTISVVNQIYGYFVLGHPMCVLEGYTVSLC 1 2 gITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGVAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCILPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIVTFCVCWGPYAFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ceb545 Cebus capucinus P545 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFTYTNSNSTR 1 2 gPFEGPNYHIAPRWVYHLTSVWMVFVVVASVFTNGLVLVATMKFKKLRHPLNWILVNLAVALAETVIASTISIVNQIYGYFVLGHPMCVLEGYTVSLC 1 2 gITGLWSLAIISWERWLVVCKPFGNMRFDAKLAIVGVAFSWIWAAVWTAPPIFGWSr 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCILPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMILTFCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ceb560 Cebus capucinus P560 full 0 MAQQWSLQRLAGRHPQDNYEDSTQSSIFTYTNSNSTR 1 2 gPFEGPNYHIAPRWVYHLTSVWMVFVVFASVFTNGLVLAATMKFKKLRHPLNWILVNLAIADLAETVIASTISVVNQVHGYFVLGHPMCVLEGYTVSLC 1 2 gITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGVAFSWIWSAVWTAPPIFGWSr 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFLPLGIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIMAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ate560 Ateles geoffroyi P560 full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSVFTYTNNNATR 1 2 gPFEGPNYHIAPRWVYHLTSVWMVFVVTASVFTNGLVLAATMKFKKLRHPLNWILVNLAVALAETVIASTISVVNQFYGYFVLGHPMCVLEGYTVSLC 1 2 gITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGVAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVKSYMIVLMVTCCFLPLGIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIMAYCVCWGPYTFFACFAAANPGYAFHPLMAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_ate552 Ateles geoffroyi P552 full 0 MAQQWSLQRLAGRHPQDSYEDSTQSSVFTYTNNNATR 1 2 gPFEGPNYHIAPRWVYHLTSVWMLFVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLTETIIASTISVVNQFYGYFVLGHPMCVLEGYTVSLC 1 2 gITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGVAFSWIWSAVWTAPPIFGWSr 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMVTCCILPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMIMAFCICWGPYTFFACFAAAKPGYAFHPLMAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEVSSVSSVSPA* 0 >LWS_tarSyr Tarsius syrichta (tarsier) frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 GPFEGLNYHIAPRWVYHVTSIWMIFVVIASTFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVLEGYTVSLC 1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCITPLTIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAkSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSEVSSASKTEVSSVSSVSPA* 0 >LWS_otoGar Otolemur garnettii (bushbaby) full 0 MAQQWGPQRLTGRQPQDTHEDSTQGSIFTYTNSNTTR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVIASIFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMMVLMVTCCVIPLSIIMLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCVCWGPYAFFACFAASHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSKTEVSSVSSVSPA* 0 >LWS_otoCra Otolemur crassicaudatus full 0 MAQQWGPQRLTGRQPQDTHEDSTQGSIFTYTNSNTTR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVIASIFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMMVLMVTCCVIPLSIIMLCYLQVWLAIRA 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCVCWGPYAFFACFAASHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSKTEVSSVSSVSPA* 0 >LWS_micMur Microcebus murinus (mouse_lemur) full 0 MAQQWGPHRFAGGQPQDSHEDSTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAIADLAETIIASTISVVNQIYGYFVLGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNMRFDAKLAIMGITFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIMLMVTCCIIPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCVCWRPYTFFACFAAAHPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRq 0 0 FRNCILQLFGKKVDDGSELSSTSKTEVSSVSSVSPA* 0 >LWS_tupBel Tupaia belangeri (tree_shrew)full 0 MAQRWGPHKLAGGQPQDSYEDSTQSSIFTYTNTNSTR 1 2 GPFEGPNYHIAPRWVYHITSMWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAIADLAETVIASTISVVNQIFGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVCAYCVCWGPYTFFACFAAAHPGYAFHPLLAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSASRTEASSVSSVSPA* 0 >LWS_musMus Mus musculus (mouse) full 0 MAQzzzzzzLTGEQTLDHYEDSTHASIFTYTNSNSTK 1 2 GPFEGPNYHIAPRWVYHLTSTWMILVVVASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVIEGYIVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLATVGIVFSWVWAAIWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMMVLMVTCCIFPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAYCLCWGPYTFFACFATAHPGYAFHPLVASLPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILHLFGKKVDDSSELSSTSKTEVSSVSSVSPA* 0 >LWS_ratNor Rattus norvegicus (rat) full 0 MAQzzzzzzLTGEQTLDHYEDSTQASIFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTSTWMILVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVIEGYIVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLATVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMMVLMVTCCIFPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAYCLCWGPYTFFACFATAHPGYAFHPLVASLPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSKTEVSSVSSVSPA* 0 >LWS_nanEhr Nannospalax ehrenbergi (mole_rat) full 0 MAQQWAPQRLAGGQTQDSYEDSTQASLFTYTNSNSTR 1 2 GPFEGPNYHIAPRWVYHLTTTWMILVVVASIFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVIEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLATMGITFAWVWAAVWTAPPVFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMVVLMITCCIIPLSIILLCYLQVWLAIRT 2 0 VAKQQKESESTQKAEKEVTHMVVVMVFAYCLCWGPYTFFVCFATAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FQNCILQLFGKKVDDSSELSSTSKTEASSVSSVSPA* 0 >LWS_speTri Spermophilus tridecemlineatus (squirrel) full 0 MAQRWDPQRLAGGQPQDSHEDSTQSSIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHITSIWMIIVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAIADLAETVIASTISVVNQFFGYFVLGHPLCVVEGYTVSVC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFAWTWSAVWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGNSYPGVQSYMMVLMVTCCIIPLSIIILCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYASFACFATANPGYAFHPLVAAVPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDTSELSSASRTEASSVSSVSPA* 0 >LWS_sciCar Sciurus carolinensis (squirrel) frag 0 zAQRWDPQRLAGGQPQDSHEDSTQSSIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHITSTWMIIVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAIADLAETVIASTISVVNQLYGYFVLGHPLCVVEGYTVSVC 1 2 GITGLWSLAIISWERWLVVCKPFGNMRFDAKLAIVGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGMQSYMMVLMVTCCIIPLSIIILCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAYCLCWGPYTFFACFATANPGYAFHPLVAALPAYFAKSATIYNPIzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_dipOrd Dipodomys ordii (pika) frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 GPFEGPNYHIAPRWVYHLTTIWMMFVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQFNGYFVLGHPLCIVEGYTVSLC 1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMMVLMVTCCIIPLSIIVLCYLQVWLAIRA 0 0 VAKLQKESESTQKAEKEVTRMVVVMVFAYCVCWGPYAFFACFATAHPGYAFHPLVAALPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSKTEASSVSSVSPA* 0 >LWS_cavPor Cavia porcellus (guinea_pig) full 0 MAQRWGPHALSGVQAQDAYEDSTQASLFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYHLTSAWMTIVVIASIFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETVIASTISVVNQVYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIVFSWVWSAVWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMMVLMVTCCITPLSIIVLCYLHVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYAFFACFATANPGYSFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVEDSSELSSTSRTEASSVSSVSPA* 0 >LWS_oryCun Oryctolagus cuniculus (rabbit) full 0 MTQPWGPQMLAGGQPPESHEDSTQASIFTYTNSNSTR 1 2 GPFEGPNFHIAPRWVYHLTSAWMILVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETVIASTISVVNQFYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPYGLKTSCGPDVFSGTSYPGVQSYMMVLMVTCCIIPLSVIVLCYLQVWMAIPA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAYCLCWGPYTFFACFATAHPGYSFHPLVAAIPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVEDSSELSSASRTEASSVSSVSPA* 0 >LWS_canFam Canis familiaris (dog) full 0 MTQRWGPQRLAGGQPQAGLEESTQASIFTYTNSNATR 1 2 DPFEGPNYHIAPRWVYHLTSAWMIFVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAIADLAETIIASTISVVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIILCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSASRTEASSVSSVSPA* 0 >LWS_phoVit Phoca vitulina (seal) full 0 MAQTWGPQRLADGRPQPGYEDSTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSTWMIFVVIASVFTNGLVLaATMRFKKLRHPLNWILVNLAVADLAETIIASTISIVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLGVIILCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVFAFCVCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSASKTEASSVSSVSPA* 0 >LWS_lepWed Leptonychotes weddellii (seal) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHFTSIWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISIVNQIHGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLGVIILCYLQVWLAIRA 0 0 VAKQQKESEzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_mirAng Mirounga angustirostris (elephant seal) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GAFEGPNYHIAPRWVYHLTSIWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISIVNQIHGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLGVIILCYLQVWLAIRA 0 0 VAKQQKDzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_eriBar Erignathus barbatus (seal) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSTWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIHGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLGVIILCYLQVWLAIRA 0 0 VAKQQKESEzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_zalCal Zalophus californianus (sea lion) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSIWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIILCYLQVWLAIRA 0 0 VAKQQKESzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_odoRos Odobenus rosmarus (walrus) frag 0 zzzzzzzzzzzzzzzzzzzzzztqASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSTWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIVFSWVWSALWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIILCYLQVWLAIRA 0 0 VAKQQKESESTzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_ursMar Ursus maritimus (polar bear) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETVIASTISVVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIILCYLQVWLAIRA 0 0 VAKQQKESEzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_enhLut Enhydra lutris (sea otter) frag 0 zzzzzzzzzzzzzzzzzzzzzzTQASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSTWMIFVVFASVFTNGLVLAATMRFKkLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIAGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTScGPDVFSGSSYPGVQSYMIVLMITCCIIPLSIIILCYLQVWLAIRA 0 0 VAKQQKESESTzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_felCat Felis catus (cat) full 0 MTQRWGPQRLAGGQPHAGLEDSTRASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHVTSAWMIFVVIASVFTNGLVLAATMKFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCIIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCVCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSELSSASRTEASSVSSVSPA* 0 >LWS_bosTau Bos taurus (cow) full 0 MAHAWGPQRLAGGQPQANFEESTQGSIFTYTNSNSTR 1 2 DPFEGPNYHIAPRWVYHLTSAWMVFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAIADLAETIIASTISVVNQMYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAITGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFIPLSVIILCYLQVWLAIRA 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVSPA* 0 >LWS_odoVir Odocoileus virginianus (deer) frag 0 MAHEWGPQRLAGGQPQANFEESTQGSIFTYTNSNSTR 1 2 DPFEGPNYHIAPRWVYHLTSAWMVFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAIADLVETIIASTISVVNQMYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIAGIAFSWIWAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAFCLCWGPYAFFzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_turTru Tursiops truncatus (dolphin) full 0 MAQQWGPQRFAGGQPQTSFEDSTQGSVFTYTNSNSTR 1 2 DPFDGPNYHIAPRWVYHLTSVWMVFVLIASIFTNGPVLAATMKFKKLRHPLNWMLVNLAVADLAETVIASTISVVNQMYGYFVLGHPLCIVEGFTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMITLMITCCFIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTRKAEKEVTRMVMVMIFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVSPA* 0 >LWS_delDel Delphinus delphis (dolphin) full 0 MAQTWGPQRFAGGQPQTSFEDSTQGSVFTYTNSNSTR 1 2 DPFDGPNYHIAPRWVYHLTSVWMVFVLIASIFTNGPVLAATMKFKKLRHPLNWMLVNLAVADLAETVIASTISVVNQMYGYFVLGHPLCIVEGFTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIAGIAFSWIWAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMITLMITCCFIPLSVTVLCYLQVWLAIRA 0 0 VAKQQKESESTRKVEKEVTRMVMVMIFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVSPA* 0 >LWS_gloMel Globicephala melas (pilot_whale) full 0 MAQQWGPQRFAGGQPQTSFEDSTQGSVFTYTNSNSTR 1 2 DPFDGPNYHIAPRWVYHLTSVWMVFVLIASIFTNGPVLAATMKFKKLRHPLNWMLVNLAVADLAETVIASTISVVNQMYGYFVLGHPLCIVEGFTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIVGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMITLMITCCFIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTRKAEKEVTRMVMVMIFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPSYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVTPA* 0 >LWS_susScr Sus scrofa (pig) full 0 MAQQWGPRRLAGGQPQASFEDSTQGSIFTYTNANSTR 1 2 DPFEGPNYHIAPRWVYHITSAWMVFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAIADLAETIIASTISVVNQMYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCFIPLGVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVSPA* 0 >LWS_equCab Equus caballus (horse) full 0 MAQRWGPQKLAGGQPQAGFEDSTQASIFTYTNNNATR 1 2 DPFEGPNYHIAPRWVYHVTSAWMIFVVIASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVLGHPMCVVEGYTVSLC 1 2 GITGLWSLAIISWERWMVVCKPFGNVRFDAKLAVAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMITCCIIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMVFAFCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSVSKTEASSVSSVSPA* 0 >LWS_myoLuc Myotis lucifugus (brown bat) full 0 MAQRWGPQRLAGGQLQASFEDSTLASIFTYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSAWMVFVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETLIASTISVVNQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNMRFDAKLAIAGITFSWVWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMILAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKRVDDSSELSSTSRTEASSVSSVSPA* 0 >LWS_myoVel Myotis velifer (bat) AM263194 mRNA frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzxxxxxVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGITFSWVWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSVIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMILAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_pteVam Pteropus vampyrus (flying fox) full 0 MAQSWVPQRLAEGQPQASFEESTQASIFIYTNSNVTR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVIASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETIIASTISVANQIYGYFVLGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNMRFDAKLATVGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLSVILFCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSSSKTEASSVSSVSPA* 0 >LWS_pteDas Pteropus dasymallus formosus (bat) AM263195 mRNA frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNMRFDAKLATVGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLSVILFCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_hapFis Haplonycteris fischeri (fruit_bat) mRNA frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDTKLATVGIAFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMVTCCIIPLSVIVFCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCLCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_sorAra Sorex araneus (shrew)full 0 MAQQzzzzzzzzzzzzRGPEDSTQHSIFTFTNSNATR 1 2 GPFEGPNYHIAPRWVYHVTSAWMIFVVFASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETLIASTISVVNQVYGYFVLGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGSMRFDAKLAIVGIAFSWVWSAIWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGDSYPGVQSYMIVLMITCCIIPLGVILFCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMVLAYCVCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSVSKSEATSVSSVSPA* 0 >LWS_eriEur Erinaceus europaeus (hedgehog)frag 0 MAQREGPQRLAGVQPQDGLESSTLGSTFTYSGSNSTI 1 2 GPFEGPNYHITPRWAFHLASAWMVFVVFASVFTNGLVLAATMRFKKLRHPLNWILVNLAVADLAETVIASTISLVNQMFGYFVLGYPVCVLECYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGSLRFDAKMAILGIAFSWIWSAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMMVLMVTCCIIPLSIIVFCYLHVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMVLAYCVCWGPYTFSASFAAAHPGYTFHPLLATLPSYFAKSATIYNPVIYVFMNRQ 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_loxAfr Loxodonta africana (elephant) full 0 MAQQWGPHRLTGARLQDASEDSTQASIFVYTNTNTTR 1 2 GPFEGPNYHIAPRWVYHLTSAWMLFVVLASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETVIASTISVVNQIYGYFILGHPLCVVEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAIWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMIVLMTTCCIIPLSIIVLCYLHVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVVVMILAYCLCWGPYTFFACFATAHPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSASKTEASSVSSVSPA* 0 >LWS_triMan Trichechus manatus (manatee) full 0 MAQQWGTHRLTGGQPQDAYEDSTQASIFVYTNTNTTR 1 2 GPFEGPNHHIAPRWVYHLTSSWMLFVVMASLLTNGLVLAATVRFKKLRHPLNWILVNLAVADLLETVIASTISVVNQIHGYFILGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGIAFSWIWSAIWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVQSYMMVLMITCCIIPLGIIVLCLHVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVLVMILAYCFCWGPYTFFACFAAAHPGYAFHPLVAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSASKTEASSVSSVSPA* 0 >LWS_echTel Echinops telfairi (tenrec) full 0 MAQRWGAHRLTGGQLQDTYEGSTRTSIFVYTNSTSTR 1 2 GPFEGPNYHIAPPWVYHLTSTWMIFVVFASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETIIASTISVVNQIYGYFVFGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIVGITFSWVWSAIWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGVRSYMIVLMITCCIIPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCLCWGPYTIFACFAAAHPGYSFHPLMAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSRTEASSVSSVSPA* 0 >LWS_proCap Procavia capensis (hyrax) full 0 MAEPWGTHRLTGRQLPDASEDSTQASIFVYTNSNATR 1 2 GPFEGPNYHIAPRWVYHLTSAWMIFVVLASVFTNGLVLVATMRFKKLRHPLNWILVNLAVADLAETVIASTISVVNQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWLVVCKPFGNVRFDAKLAIAGIAFSWIWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSYPGAQSYMIVLMITCCIIPLSIIVLCYLHVWLAIRA 0 0 VAKQQKESESTQKAEKEVTRMVMVMIFAYCVCWGPYTFFACFAAAHPGYAFHPLMAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDSSELSSTSKMEASSASSVSPA* 0 >LWS_monDom Monodelphis domesticus (opossum) VNE excised full 0 MTQAWDPAGFLARRRD DDNDETTRSSLFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMVFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETVIASTISVINQIYGYFILGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIIFSWVWAAVWTAPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMATCCIFPLSIILLCYVQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTLFACFAAANPGYSFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >LWS_macEug Macropus eugenii (wallaby) full 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTK 1 2 GPFEGPNYHIAPRWVFNLTSLWMIFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETLIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGNSDPGVQSYMIVLMSTCCILPLSVIFLCYIQVWLAIRS 2 0 VAKQQKESESTQKAEKEVSRMVVVMILAFCFCWGPYAIFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >LWS_smiCra Sminthopsis crassicaudata (dunnart) EU232013 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTK 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMSTCCILPLSIIILCYIQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVMVMILAFCFCWGPYALFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >LWS_setBra Setonix brachyurus (quokka) full 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTK 1 2 GPFEGPNYHIAPRWVFNLTSLWMIFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETMIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGNSDPGVQSYMIVLMSTCCILPLSVILLCYIQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYAIFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVPA* 0 >LWS_cerCon Cercartetus concinnus (pygmy_possum) full 0 MTQAWDPAGFLAWQEDENEETTRASLFVYTNSNNTK 1 2 GPFEGPNYHIAPRWVFNLTSLWMVFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAIADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAIWTSPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGNSDPGIQSYMIVLMSTCCILPLSIILLCYIQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTFFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* >LWS_tarRos Tarsipes rostratus (honey_possum) full 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVFNLTSLWMVFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAIWTSPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGNSDPGIQSYMIVLMSTCCILPLSIILLCYVQVWRAIRA 2 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTLFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >LWS_isoObe Isoodon obesulus (bandicoot) full 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSFWMFVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMTTCCILPLSIILLCYVQVWLAIRA 0 0 VAKQQKDSESTQKAEKEVSRMVVVMIRAYCFCWGPYTLFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSGTSRTEVSSVSSAPA* 0 >LWS_myrFas Myrmecobius fasciatus (numbat) full 0 MTQAWDPAGFLAWRREENEETTRASLFTYTNSNNTK 1 2 GPFEGPNYHIAPRWVYNLTSFWMIFVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMSTCCILPLSVILLCYIQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYAIFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSRTEVSSVSSVAPA* 0 >LWS_ornAna Ornithorhynchus anatinus (platypus) full 0 MTPAWNSGVYAARRRFEDEEDTTRTSVFVYTNSNNTR 1 2 DPFEGPNYHIAPRWAYNVTSLWMIFVVIASVFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETLIASTISVINQIFGYFILGHPMCVLEGYTVSLC 1 2 GITGLWSLSIISWERWIVVCKPFGNVKFDAKLAMVGIVFSWVWAAVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMSTCCILPLSIIVLCYLQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTIFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSELSSTSRTEVSSVSSVSPA* 0 >LWS_tacAcu Tachyglossus aculeatus (echidna) full 0 MTQAWDPAGFLAWRRDENEETTRASLFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVFNLTSLWMVFVVIASIFTNGLVLVATMKFKKLRHPLNWILVNLAVADLGETIIASTISVINQIYGYFILGHPLCVLEGYTVSLC 1 2 GITGLWSLAIISWERWVVVCKPFGNVKFDAKLAMVGIIFSWVWAVWTSPPLFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMIVLMATCCIFPLSIILLCYIQVWLAIRA 0 0 VAKQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTLFACFAAANPGYAFHPLTASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCILQLFGKKVDDGSEVSSTSKTEVSSVSSVAPA* 0 >LWS_galGal Gallus gallus (chicken) missing genome full 0 MAAWEAAFAARRRHEEEDTTRDSVFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVAASVFTNGLVLVATWKFKKLRHPLNWILVNLAVADLGETVIASTISVINQISGYFILGHPMCVVEGYTVSAC 1 2 GITALWSLAIISWERWFVVCKPFGNIKFDGKLAVAGILFSWLWSCAWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMVVLMVTCCFFPLAIIILCYLQVSLAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMIVAYCFCWGPYTFFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSEVSTSRTEVSSVSNSSVSPA* 0 >LWS_taeGut Taeniopygia guttata (finch) GV excised full 0 MAT WDGAVFAARRRHDDEDTTRDSIFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVVASVFTNGLVLVATAKFKKLRHPLNWILVNLAVADLGETVIASTISVVNQIFGYFILGHPMCVIEGYTVSAC 1 2 GITALWSLAIISWERWFVVCKPFGNIKFDGKLAVAGVLFSWIWSCAWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSTDPGVQYMVVLMVTCCFFPLAVIIFCYLQVWLAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTIFACFAAANPGYAFHPLTAALPAFFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSEVSTSRTEVSSVSNSSVSPA* 0 >LWS_colLiv Columba livia (pigeon) P559 full 0 MDGFAAARRRHEDEDTTRDSVFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVVASVFTNGLVLVATWRFKKLRHPLNWILVNLAVADLGETVIASTISVVNQISGYFVLGHPMCVLEGYTVSAC 1 2 GITALWSLAIISWERWFVVCKPFGNIKFDGKLALAGILFSWVWSCAWTAPPVFGWSR 2 0 YWPHGLKTSCGPDVFSGSSDPGVQSYMVVLMVTCCFFPLAIIVLCYLQVWLAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMIVAYCFCWGPYTIFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSEVSTSRTEVSSVSSSSVSPA* 0 >LWS_serCar Serinus canaria (canary) frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVVASVFTNGLVLVATAKFKKLRHPLNWILVKLSVAELGETVIASTISVVNQIFGYFILGHPMCVIEGYTVSAC 1 2 GITALWSLAIISWERWFVVCKPFGNIKFDGKLAVAGVLFSWIWSCAWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGTTDPGVQSYMVVLMVTCCFFPLAVIIFCYLQVWLAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMILAYCFCWGPYTIFASFAAANPYAFHPLTAALPAFFAKSATIYNPIIYVFMNRQ 0 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz* 0 >LWS_anoCar Anolis carolinensis (lizard) GTVT excised full 0 MA EAWDVAVFAARRRNDEDDTTRDSLFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNITSVWMIFVVIASIFTNGLVLVATAKFKKLRHPLNWILVNLAIADLGETVIASTISVINQISGYFILGHPMCVLEGYTVSTC 1 2 GISALWSLAVISWERWVVVCKPFGNVKFDAKLAVAGIVFSWVWSAVWTAPPVFGWSR 2 1 YWPHGLKTSCGPDVFSGSDDPGVLSYMIVLMITCCFIPLAVILLCYLQVWLAIRA 0 0 VAAQQKESESTQKAEKEVSRMVVVMIIAYCFCWGPYTVFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSELSSTSRTEVSSVSNSSVSPA* 0 >LWS_gecGec Gecko gecko (gecko) P521 full 0 MTEAWNVAVFAARRSRDDDDTTRGSVFTYTNTNNTR 1 2 GPFEGPNYHIAPRWVYNLVSFFMIIVVIASCFTNGLVLVATAKFKKLRHPLNWILVNLAFVDLVETLVASTISVFNQIFGYFILGHPLCVIEGYVVSSC 1 2 GITGLWSLAIISWERWFVVCKPFGNIKFDSKLAIIGIVFSWVWAWGWSAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSVELGCQSFMLTLMITCCFLPLFIIIVCYLQVWMAIRA 0 0 VAAQQKESESTQKAEREVSRMVVVMIVAFCICWGPYASFVSFAAANPGYAFHPLAAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSEASTTSRTEVSSVSNSSVAPA* 0 >LWS_pheMad Phelsuma madagascariensis (gecko) full 0 MTEAWNVAVFAARRSRDDDDDTTRGSVFTYTDSNNTK 1 2 GPFDGPNYHIAPRWVYNFTSFWMVIVVIASTFTNGLVLVATVKFKKLRHPLNWILVNLAFVDMVETVIASGISVINQIFGYFILGHPLCVIEGYLVSAC 1 2 GITGLWSLAIISWERWFVVCKPFGNIKFDSKLAILGIAFSWIWAWGWSAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGNNELGCQSYMLALMVSCCFFPLSVIILCYLQVWMAIRA 0 0 VAAQQKESESTQKAEKEVTRMVVVMLLAFCVCWGPYTIFVGFAAAHPGYAFHPLAAALPAYFAKSATIWNPVIYIFMNRQ 0 0 FRNCILQLFGKKVDDASDVSTTSRTEVSSVSNSSVSPA* 0 >LWS_utaSta Uta stansburiana (lizard) frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz1 2 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 GITALWSLAIISWERWVVVCKPFGNVKFDAKLAMAGIVFSWVWSCVWTAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSDDPGVQSYMIVLMITCCFIPLAVIILCYLQVWMAIRA 0 0 VAAQQKESESTQKAEREVSRMVVVMIIAYCFCWGPYTIFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSTSRTEVSSVSNSSVSPA* 0 >LWS_ambTig Ambystoma tigrinum (salamander) full 0 MAHSWNSGAYAARRRYDDEDTTRSSIFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTTLWMIFVVFASVFTNGLVLVATMKFKKLRHPLNWILVNLAIADLGETVIASTISVINQMFGYFILGHPLCVIEGYTVSVC 1 2 GISALWSLTIIAWERWFVVCKPFGNIKFDGKLAAAGIIFSWVWSAGWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMMVLMITCCILPLSIIIICYIQVWWAIRQ 0 0 VAMQQKESESTQKAEREVSRMVVVMIIAYIFCWGPYTFFACFAAANPGYAFHPLAASLPAFFAKSATIYNPIIYVFMNRQ 0 0 FRNCIYQLFGKKVDDGSEMSSASRTEVSSVSNSSVSPA* 0 >LWS_cynPyr Cynops pyrrhogaster (newt) full 0 MAYSWNSGAYAARRRYDDEDTTRSSVFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTTLWMFFVVAASVFTNGLVLVATMKFKKLRHPLNWILVNLAIADIAETLIASTISVINQIFGYFILGHPLCVIEGYTVSVC 1 2 GITGLWSLTIIAWERWFVVCKPFGNIKFDGKLAAAGIIFSWAWAAFWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGNSDPGVQSFMITLMSTCCILPLSIIILCYVQVWWAIRQ 0 0 VAMQQKESESTQKAEREVSRMVVVMIMAYIFCWGPYTFFVCFAAANPGYSFHPLAASLPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIYQLFGKKVDDGSELSSSSRTEVSSVSNSSVSPA* 0 >LWS_xenTro Xenopus tropicalis (frog) full 0 MASHWNEAVFAARRRNDDDDTTRSSVFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNISSLWMIFVVLASVFTNGLVLVATLKFKKLRHPLNWILVNMAIADLGETVIASTISVCNQIFGYFVLGHPMCILEGYTVSVC 1 2 GIAALWSLTVIAWERWFVVCKPFGNIKFDGKLAATGIIFSWVWAAGWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMLVLMITCCIIPLAIIVLCYMHVWLTIRQ 0 0 VAQQQKESESTQKAEREVSRMVVVMIIAYIFCWGPYTFFACFAAFNPGYNFHPLAAAMPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIYQLFGKKVDDGSEVSSTSRTEVSSVSNSSVSPA* 0 >LWS_xenLae Xenopus laevis (frog) full 0 MASQLNEAIFAARRRNDDDDTTRSSVFTYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSIWMIFVVFASVFTNGLVIVATLKFKKLRHPLNWILVNMAIADLGETVIASTISVFNQIFGYFILGHPMCVLEGFTVSTC 1 2 GITALWSLTVIAWERWFVVCKPFGNIKFDEKLAATGIIFSWVWSAGWCAPPMFGWSR 2 1 FWPHGLKTSCGPDVFSGSSDPGVQSYMLVLMITCCIIPLAIIILCYLHVWWTIRQ 0 0 VAQQQKESESTQKAEREVSRMVVVMIVAYIFCWGPYTFFACFAAFSPGYSFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIYQMFGKKVDDGSEVSSTSRTEVSSVSNSSVSPA* 0 >LWS_neoFor Neoceratodus forsteri (lungfish) full 0 MAEPWDAVLAARRRHQDEETTRSTIFVYTNSNNTR 1 2 GPFEGPNYHIAPRWVYNLTSLWMIFVVFASCFTNGLVLMATYKFKKLRHPLNWILVNLAIADLGETLIASTISVTNQIFGYFILGHPMCMLEGFTVATC 1 2 GITGLWSLTIIAWERWVVVCKPFGNIKFDGKWAAGGIIFSWVWSAFWCAMPLFGWSR 2 1 FWPHGLKTSCGPDVFSGEDKYGTRSFMIALMITCCIIPLGVIILCYIQVWWAIRT 0 0 VAKQQKESESTQKAEKEVSRMVVVMIFAYCFCWGPYTFMACFGAAYPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIYQLLGKKVDDGSELSSTSKTEVSSVSNSSVSPA* 0 >LWS_danRer Danio rerio (zebrafish) full 0 MAEHWGDAIYAARRKGDETTREAMFTYTNSNNTK 1 2 DPFEGPNYHIAPRWVYNVATVWMFFVVVASTFTNGLVLVATAKFKKLRHPLNWILVNLAIADLGETLFASTISVINQFFGYFILGHPMCIFEGYTVSVC 1 2 GIAALWSLTVISWERWVVVCKPFGNVKFDAKWASAGIIFSWVWAAAWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMVVLMITCCIIPLAIIILCYIAVYLAIHA 2 0 VAQQQKDSESTQKAEKEVSRMVVVMIFAYCFCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 FRVCIMQLFGKKVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_cypCar Cyprinus carpio (carp) full 0 MAEQWGDAIFAARRRGDETTRETMFVYTNSNNTR 1 2 DPFEGPNYHIAPRWVYNLATVWMFFVVIASTFTNGLVLVATAKFKKLRHPLNWILVNLAIADLAETLLASTISVINQIFGYFILGHPMCIFEGYTVSVC 1 2 GIAGLWSLTVISWERWVVVCKPFGNVKFDAKWASAGIIFSWVWSAFWCAPPIFGWSR 2 1 FWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMITCCIIPLAIIILCYIAVWLAIRA 0 0 VAQQQKDSESTQKAEKEVSRMVVVMIFAYCFCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPIIYVFMNRQ 0 0 FRVCIMQLFGKKVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_salSal Salmo salar (salmon) full 0MAERWGSAAYAARRQNQDTTRESSFTFTNSNNTK 1 2 DPFEGPNYHIAPRWVYNLSTLWMVIVVILSVFTNGLVLVATAKFKKLQHPLNWILVNLAIADIGETLLASTISVCNQFFGYFILGHPMCVFEGYTVSVC 1 2 GIAALWSLAVISWERWVVVCKPFGSVKFDAKWAMGGIIFSWVWAAFWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFGGNEDPGVKSYMITLMITCCFFPLFVIIFCYIFVWLAIRA 0 0 VAAQQKDSESTQKAEKEVSRMVVVMIIAYCVCWGPYTCFACFAAANPGYAFHPLAAAIPAYFAKSATIYNPVIYVFMNRQ 0 0 FRTCIMQLFGKAEDDGTEVSTSKTEVSSxxxxxVAPA* 0 >LWS_pleAlt Plecoglossus altivelis (ayu) full 0 MTDEWGNAVFAARRRNEDTTRESSFTYTNSNNTK 1 2 DPFEGPNYHIAPRWVYNISTMWMIFVVIASVFTNGLVLVATAKFKKLQHPLNWILVNLAIADLGETVLASTISVCNQTFGYFILGHPMCVFEGYTVSTC 1 2 GIAALWSLTVISWERWVVVCKPFGNVKFDAKWATAGIVFSWVWSAAWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSDDPGVKSYMIVLMITCCFLPLAIIILCYIAVWLAIRA 0 0 VAQQQKDSESTQKAEKEVSRMVVVMILAYIVCWGPYTVFACFAAANPGYAFHPLAAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRVCIMQLFGKKVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_tetNig Tetraodon nigroviridis (pufferfish) full 0 MAEEWGKQSFAARRYHEDSTRGSAFVYTNSNHTR 1 2 DPFEGPNYHIAPRWVYNLATLWMFFVVVLSVFTNGLVLVATAKFKKLRHPLNWILVNLAVADLGETLFASTISVCNQFFGYFILGHPMCIFEGYVVSVC 1 2 GIAGLWSLTIISWERWIVVCKPFGNVKFDAKWATAGIVFSWIWPICWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMITCCIIPLAIIVLCYLAVWMAIRA 2 0 VAMQQKESESTQKAEREVSRMVVVMILAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPIIYVFMNRQ 0 0 FRVCIMKLFGKEVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_takRub Takifugu rubripes (pufferfish) full 0 MAEEWGKQSFAARRYHEDTTRGSAFVYTNSNHTR 1 2 DPFEGPNYHIAPRWVYNVATVWMFIVVVLSVFTNGLVLVATAKFKKLRHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGYTVSTC 1 2 GIAALWSLTIISWERWVVVCKPFGNVKFDAKWATGGIVFSWVWAAVWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMITCCIIPLAIIILCYLAVWLAIRS 0 0 VAMQQKESESTQKAEKEVSRMVVVMIVAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 FRVCIMKLFGKEVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_gasAcu Gasterosteus aculeatus (stickleback) full 0 MAEEWGKQAFAARRYNEDTTRGSMFVYTNSNNTK 1 2 DPFEGPNYHIAPRWVYNLSTLWMFIVVALSVFTNGLVLVATAKFKKLQHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGYVVSVC 1 2 GITALWSLTIISWERWIVVCKPFGNVKFDAKWATAGIVFSWIWSAVWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMITCCLIPLAIIILCYLAVWLAIRA 0 0 VAMQQKESESTQKAERDVSRMVVVMIVAYIVCWGPYTTFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 FRSCIMQLFGKEVDDGSEVSTsKTEVSSxxxxxVAPA* 0 >LWS_oryLat Oryzias latipes (medaka) full 0 MAEQWGKQVFAARRQNEDTTRGSAFTYTNSNHTR 1 2 DPFEGPNYHIAPRWVYNLATLWMFFVVVLSVFTNGLVLVATAKFKKLRHPLNWILSNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGYVVSTC 1 2 GIAALWSLTIISWERWVVVCKPFGNVKFDAKWAIGGIVFSWVWSAVWCAPPVFGWSR 2 1 YWPHGLKTSCGPDVFSGSDDPGVQSYMIVLMITCCIIPLAIIILCYLAVWLAIRA 0 0 VAMQQKESESTQKAEREVSRMVVVMIVAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 FRTCIMQLFGKQVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_poeRet Poecilia reticulata (guppy) full 0 MAEEWGKQVFAARRHEDTTRGAAFTYTNSNHTK 1 2 DPFEGPNYHIAPRWVYNVSTLWMCIVVVLSDFTNGLVLVATAKFKKLRHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGYVVSTC 1 2 GIAALWSLTIISWERWIVVCKPFGNVKFDAKWATAGIVFSWAWAAVWCAPPIFGWS 2 1 RYWPHGLKTSCGPDVFSGSDDPGVLSYMIVLMITCCIIPLAIIILCYLAVWLAIRA 0 0 VAMQQKESESTQKAEREVSRMVVVMIIAYCVCWGPYTFFACFPAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 FRTCIMQLFGKQVDDGFEVSTSKTEVSSxxxxxVAPA* 0 >LWS_lucGoo Lucania goodei (killifish) full 0 MAEQWEKQAFAARRYNEETTRGSVFTYTNSNHTR 1 2 DPFEGPNYHIAPRWVYNVSTVWMFFVVVLSVFTNGLVLVATAKFEKLRHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGFIVSTC 1 2 GIAALWSLTIISWERWIVVCKPFGNVKFDAKWATAGIVFSWVWSAVWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMVVLMITCCIIPLAIIILCYLAVWLAIRAVA 0 0 MQQKESESTQKAEREGSRMVVVMILAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPVIYVFMNRQ 0 0 SRTCIMQLFGKQVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_ophVen Ophthalmotilapia ventralis (cichlid) full 0 MAEEWGKQSFAARRYHEDSTRGSAFAYTNSNNTR 1 2 DPFEGPNYHIAPRWIYNVATLWMFVVVVLSVFTNGLVLVATMKFKKLRHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCIFEGYVVSVC 1 2 GIAALWSLTIISWERWIVVCKPFGNVKFDAKWATAGIVFSWVWSAVWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMITCCILPLAVIILCYLAVWLAIRA 0 0 VAMQQKESESTQKAEREVSRMVVVMIVAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCIMQLFGKQVDDGSEVSTSKTEVSSxxxxxVAPA* 0 .>LWS_pseMax Psetta maxima (turbot) full 0 MAEDWGKPAFAARRYHEDTTRGSAFMYTNSNHTK 1 2 DPFEGPNYHIAPRWIYNLATLWMFVVVVASVFTNGLVLVATAKFKKLRHPLNWILVNLAIADLGETVFASTISVCNQFFGYFILGHPMCVFEGYTVSVC 1 2 GIAGLWSLTIISWERWIVVCKPFGNIKFDAKWATGGILFSWIWPIVWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSEDPGVQSYMIVLMVTCCFLPLAIIILCYLAVWWAIHS 0 0 VALQQKESESTQKAEKDVSRMVVVMILAYCVCWGPYTFFACFAAANPGYAFHPLAAAMPAYFAKSATIYNPIIYVFMNRQ 0 0 FRTCIMQLFGKEVDDGSEVSTSKTEVSSxxxxxVAPA* 0 >LWS_calMil Callorhinchus milii (elephantfish) frag 0 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 1 2 dPFEGPNYHIAPRWAYNLTSVWMVGVVVASVFTNGLVLVATVRFKKLRHPLNWILVNMALADLGETVLASTVSVANQFFGYFILGHPLCVFEGFVVSLC 1 2 GITALWSLTIIAWERWVVVCKPFGNVKFDGKWAAFGIIFSWVWSIGWCLPPVFGWSR 2 1 zzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzzz 0 0 zzzzzzzzzzzzzAEKEVSRMVVVMVAAFCLCWGPYACFAMFSALNPGYAFHPLVASIPSYFAKSSTIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSELSSTSKTDVSSVSNSSVSPA* 0 >LWS_petMar Petromyzon marinus (lamprey) full 0 MTASWQGAMFAARRRQDDEDTTMESLFRYTNENNTK 1 2 DPFEGPNYHIAPRWVFNLTSVWMIIVVVLSLFSNGLVLVATVKFKKLRHPLNWIIVNLAIADILETIFASTISVCNQVYGYFILGHPMCVFEGYVVSTC 1 2 GIAGLWSLAIISWERWMVVCKPFGNIKFDGKIATILIVFSWVWPASWCSLPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSTDPGVQSYMVVLMITCCFLPLSIIILCYLQVWLAIHS 0 0 VAQQQKESETTQKAERDVSRMVVVMILAYVFCWGPYTFFACFAAANPGYSFHPIAAALPAYFAKGATIYNPIIYVFMNRQ 0 0 FRNCILQLFGKKVDDGSEVSSSSRTEVSSVSNSSVSPA* 0 >LWS_letJap Lethenteron japonicum (lamprey) full 0 MTASWHGAVFAARRRNDDEDTTKDSIFRYTNENNTR 1 2 DPFEGPNYHIAPRWMFNLTSVWMIIVVVLSLFTNGLVLVATMKFKKLRHPLNWILVNLAIADILETIFASTISVCNQVFGYFILGHPMCVFEGYVVSTC 1 2 GIAGLWSLAIISWERWMVVCKPFGNIKFDGKIAIILIVFSWVWPACWCSLPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSSDPGVQSYMVVLMVTCCFLPLSVIILCYLQVWLAIHS 0 0 VAQQQKESETTQKAERDVSRMVVVMILAYIFCWGPYTFFACYAAANPGYAFHPLTAALPAYFAKSATIYNPVIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSEVSSASRTEVSSVSNSSISPA* >LWS_geoAus Geotria australis (lamprey) full 0 MAQSWERAMFAARRRQDEDTTKGDLFRYTNENNTR 1 2 DPFEGPNYHIAPRWMYNLTSFWMIIVVILSLFTNGLVLVATLKFKKLRHPLNWILVNLAIADIGETIFASTVSVVNQIFGYFILGHPLCVFEGFTVSVC 1 2 GITALWSLAIISFERWMVVCKPFGNLKFDGKVAIVLIIFSWAWSAGWCAPPIFGWSR 2 1 YWPHGLKTSCGPDVFSGSTDPGVQSYMVVLMITCCFIPLALIIICYLQVWLAIHT 0 0 VAQQQKESETTQKAERDVSRMVVVMIFAYIFCWGPYTFFACFAAANPGYAFHPLAAALPAYFAKSATIYNPIIYVFMNRQ 0 0 FRNCIMQLFGKKVDDGSEVSSSARTEVSSVSNSSVSPA* 0